Abstract
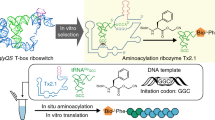
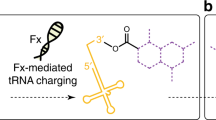
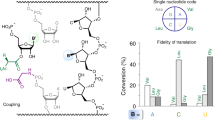
The RNA world hypothesis implies that coded protein synthesis evolved from a set of ribozyme catalyzed acyl-transfer reactions, including those of aminoacyl-tRNA synthetase ribozymes. We report here that a bifunctional ribozyme generated by directed in vitro evolution can specifically recognize an activated glutaminyl ester and aminoacylate a targeted tRNA, via a covalent aminoacyl-ribozyme intermediate. The ribozyme consists of two distinct catalytic domains; one domain recognizes the glutamine substrate and self-aminoacylates its own 5'-hydroxyl group, and the other recognizes the tRNA and transfers the aminoacyl group to the 3'-end. The interaction of these domains results in a unique pseudoknotted structure, and the ribozyme requires a change in conformation to perform the sequential aminoacylation reactions. Our result supports the idea that aminoacyl-tRNA synthetase ribozymes could have played a key role in the evolution of the genetic code and RNA-directed translation.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Schimmel, P., Giegé, R., Moras, D. & Yokoyama, S. Proc. Natl. Acad. Sci. USA 90, 8763– 8768 (1993).
Yarus, M. Science 240, 1751–1758 ( 1988).
Hager, A.J., Pollard, J.D. & Szostak, J.W. Chem. Biol. 3, 717– 725 (1996).
Piccirilli, J.A., McConnell, T.S., Zaug, A.J., Noller, H.F. & Cech, T.R. Science 256, 1420–1424 (1992).
Noller, H.F., Hoffarth, V. & Zimniak, L. Science 256, 1416– 1419 (1992).
Schimmel, P. Biochemistry 28, 2747–2759 (1989).
Giegé, R., Sissler, M. & Florentz, C. Nucleic Acids Res. 26, 5017– 5035 (1998).
McClain, W.H. J. Mol. Biol. 234, 257–280 (1993).
Nureki, O.,et al. 280, 578–582 ( 1998).
Famulok, M. J. Am. Chem. Soc. 116, 1698–1706 (1994).
Majerfeld, I. & Yarus, M. Nature Struct. Biol. 1 , 287–292 (1994).
Yang, Y., Kochoyan, M., Burgstaller, P., Westhof, E. & Famulok, M. Science 272, 1343–1347 (1996).
Illangasekare, M., Sanchez, G., Nickles, T. & Yarus, M. Science 267, 643–647 (1995).
Illangasekare, M., Kovalchuke, O. & Yarus, M. J. Mol. Biol. 274, 519– 529 (1997).
Jenne, A. & Famulok, M. Chem. Biol. 5, 23–34 (1998).
Lohse, P.A. & Szostak, J.W. Nature 381, 442–444 (1996).
Wiegand, T.W., Janssen, R.C. & Eaton, B.E. Chem. Biol. 4, 675– 683 (1997).
Zhang, B. & Cech, T.R. Nature 390, 96–100 (1997).
Suga, H., Lohse, P.A. & Szostak, J.W. J. Am. Chem. Soc. 120, 1151 –1156 (1998).
Suga, H., Cowan, J.A. & Szostak, J.W. Biochemistry 37, 10118– 10125 (1998).
Mathews, D.H., Sabina, J., Zuker, M. & Turner, D.H. J. Mol. Biol. 288, 911–940 ( 1999).
Burke, J.M.,et al. Cell 45, 167–176 (1986).
Golden, B.L. & Cech, T.R. Biochemistry 35, 3754–3763 (1996).
Chanfreau, G. & Jacquier, A. EMBO J. 15, 3466–3476 (1996).
Costa, M., Deme, E., Jacquier, A. & Michel, F. J. Mol. Biol. 267, 520–536 (1997).
Chu, V.T., Liu, Q., Podar, M., Perlman, P.S. & Pyle, A.M. RNA 4, 1186– 1202 (1998).
Lodmell, J.S. & Dahlberg, A.E. Science 277, 1262–1267 (1997).
Agrawal, R.K. & Frank, J. Curr. Opin. Struct. Biol. 9, 215–221 (1999).
Konarska, M.M. & Sharp, P.A. Cell 49, 763–774 (1987).
Fortner, D.M., Troy, R.G. & Brow, D.A. Genes Dev. 8, 221– 233 (1994).
Kambach, C., Walke, S. & Nagai, K. Curr. Opin. Struct. Biol. 9, 222 –230 (1999).
Noren, C.J., Anthony-Cahill, S.J., Griffith, M.C. & Schultz, P.G. Science 244, 182–188 ( 1989).
Arslan, T., Mamaev, S.V., Mamaev, N.V. & Hecht, S.M. J. Am. Chem. Soc. 119, 10877–10887 (1997).
Acknowledgements
We thank the NMR facility in the Department of Chemistry, the CAMBI nucleic acid facility, and the Phosphorimager facility in the Department of Biological Sciences. We thank M. Hollingsworth and P. Gollnick for critical reading the manuscript. Y.B. is supported by a Research Scientist Abroad Fellowship sponsored by the Japanese Ministry of Education. This work was supported by the State University of New York at Buffalo Start-up Fund and a PRF-ACS grant (H.S.) and partly by an NIH grant (J.W.S.).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Lee, N., Bessho, Y., Wei, K. et al. Ribozyme-catalyzed tRNA aminoacylation. Nat Struct Mol Biol 7, 28–33 (2000). https://doi.org/10.1038/71225
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/71225
This article is cited by
-
Ribosome-mediated polymerization of long chain carbon and cyclic amino acids into peptides in vitro
Nature Communications (2020)
-
An aminoacylation ribozyme evolved from a natural tRNA-sensing T-box riboswitch
Nature Chemical Biology (2020)
-
Expanding the limits of the second genetic code with ribozymes
Nature Communications (2019)
-
The chemistry and applications of RNA 2′-OH acylation
Nature Reviews Chemistry (2019)
-
The Chemical Likelihood of Ribonucleotide-α-Amino acid Copolymers as Players for Early Stages of Evolution
Journal of Molecular Evolution (2019)