Abstract
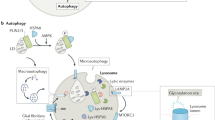
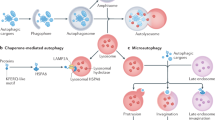
Autophagy is an intracellular degradation process that clears long-lived proteins and organelles from the cytoplasm. It involves the formation of double-membraned structures called autophagosomes that can engulf portions of cytoplasm containing oligomeric protein complexes and organelles, such as mitochondria. Autophagosomes fuse with lysosomes and their contents then are degraded. Failure of autophagy in neurons can result in the accumulation of aggregate-prone proteins and neurodegeneration. Pharmacological induction of autophagy can enhance the clearance of intracytoplasmic aggregate-prone proteins, such as mutant forms of huntingtin, and ameliorate pathology in cell and animal models of neurodegenerative diseases. In this Review, the autophagic machinery and the signaling pathways that regulate the induction of autophagy are described. The ways in which dysfunctions at multiple stages in the autophagic pathways contribute to numerous neurological disorders are highlighted through the use of examples of Mendelian and complex conditions, including Alzheimer disease, Parkinson disease and forms of motor neuron disease. The different ways in which autophagic pathways might be manipulated for the therapeutic benefit of patients with neurodegenerative disorders are also considered.
Key Points
-
Macroautophagy (also known as autophagy) is a process by which cytoplasmic substrates are sequestered within double-membraned vesicles called autophagosomes, which are then delivered to the lysosome for degradation
-
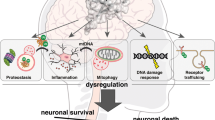
Autophagic dysfunction might contribute to the pathogenesis of numerous neurodegenerative diseases, including forms of Parkinson disease, Alzheimer disease, amyotrophic lateral sclerosis, Huntington disease and Lafora disease
-
Autophagy is regulated by both mTOR-dependent and mTOR-independent pathways, which have multiple components that could be targeted for therapeutic manipulation of this degradative process
-
Patients with conditions that are associated with the accumulation of intracytoplasmic aggregate-prone proteins may benefit from pharmacological upregulation of autophagy
-
An advanced understanding of the pathways that regulate autophagy is required to maximize the potential of targeting autophagy for therapeutic purposes
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
References
Klionsky, D. J. The molecular machinery of autophagy: unanswered questions. J. Cell Sci. 118, 7–18 (2005).
Metcalf, D. J., García-Arencibia, G., Hochfeld, W. E. & Rubinsztein, D. C. Autophagy and misfolded proteins in neurodegeneration. Exp. Neurol. http://dx.doi.org/10.1016/j.exp.neurol.2010.11.003.
Verhoef, L. G., Lindsten, K., Masucci, M. G. & Dantuma, N. D. Aggregate formation inhibits proteasomal degradation of polyglutamine proteins. Hum. Mol. Genet. 11, 2689–2700 (2002).
Yamamoto, A. & Simonsen, A. The elimination of accumulated and aggregated proteins: a role for aggrephagy in neurodegeneration. Neurobiol. Dis. 43, 17–28 (2011).
Knaevelsrud, H. & Simonsen, A. Fighting disease by selective autophagy of aggregate-prone proteins. FEBS Lett. 584, 2635–2645 (2010).
Klionsky, D. J. et al. How shall I eat thee? Autophagy 3, 413–416 (2007).
Wong, E. & Cuervo, A. M. Autophagy gone awry in neurodegenerative diseases. Nat. Neurosci. 13, 805–810 (2010).
Levine, B. & Kroemer, G. Autophagy in the pathogenesis of disease. Cell 132, 27–42 (2008).
Levine, B. & Klionsky, D. J. Development by self-digestion: molecular mechanisms and biological functions of autophagy. Dev. Cell 6, 463–477 (2004).
Ravikumar, B. et al. Regulation of mammalian autophagy in physiology and pathophysiology. Physiol. Rev. 90, 1383–1435 (2010).
Liang, C. et al. Beclin-1 binding to UVRAG targets the class C Vps complex to coordinate autophagosome maturation and endocytic trafficking. Nat. Cell Biol. 10, 776–787 (2008).
Fimia, G. et al. Ambra1 regulates autophagy and development of the nervous system. Nature 447, 1121–1125 (2007).
Ravikumar, B., Moreau, K., Jahreiss, L., Puri, C. & Rubinsztein, D. C. Plasma membrane contributes to the formation of pre-autophagosomal structures. Nat. Cell Biol. 12, 747–757 (2010).
Hayashi-Nishino, M. et al. A subdomain of the endoplasmic reticulum forms a cradle for autophagosome formation. Nat. Cell Biol. 11, 1433–1437 (2009).
Ylä-Anttila, P., Vihinen, H., Jokitalo, E. & Eskelinen, E. 3D tomography reveals connections between the phagophore and endoplasmic reticulum. Autophagy 5, 1180–1185 (2009).
van der Vaart, A., Griffith, J. & Reggiori, F. Exit from the Golgi is required for the expansion of the autophagosomal phagophore in yeast Saccharomyces cerevisiae. Mol. Biol. Cell 21, 2270–2284 (2010).
Yen, W.-L. et al. The conserved oligomeric Golgi complex is involved in double-membrane vesicle formation during autophagy. J. Cell Biol. 188, 101–114 (2010).
Geng, J., Nair, U., Yasumura-Yorimitsu, K. & Klionsky, D. J. Post-Golgi Sec proteins are required for autophagy in Saccharomyces cerevisiae. Mol. Biol. Cell 21, 2257–2269 (2010).
Hailey, D. et al. Mitochondria supply membranes for autophagosome biogenesis during starvation. Cell 141, 656–667 (2010).
Moreau, K., Ravikumar, B., Renna, M., Puri, C. & Rubinsztein, D. C. Autophagosome precursor maturation requires homotypic fusion. Cell 146, 303–317 (2011).
Mizushima, N., Sugita, H., Yoshimori, T. & Ohsumi, Y. A new protein conjugation system in human. The counterpart of the yeast Apg12p conjugation system essential for autophagy. J. Biol. Chem. 273, 33889–33892 (1998).
Mizushima, N. et al. Mouse Apg16L, a novel WD-repeat protein, targets to the autophagic isolation membrane with the Apg12–Apg5 conjugate. J. Cell Sci. 116, 1679–1688 (2003).
Luzio, J., Pryor, P. & Bright, N. Lysosomes: fusion and function. Nat. Rev. Mol. Cell Biol. 8, 622–632 (2007).
Furuta, N., Fujita, N., Noda, T., Yoshimori, T. & Amano, A. Combinational soluble N-ethylmaleimide-sensitive factor attachment protein receptor proteins VAMP8 and Vti1b mediate fusion of antimicrobial and canonical autophagosomes with lysosomes. Mol. Biol. Cell 21, 1001–1010 (2010).
Sarbassov, D. D., Ali, S. M. & Sabatini, D. M. Growing roles for the mTOR pathway. Curr. Opin. Cell Biol. 17, 596–603 (2005).
Wei, Y., Pattingre, S., Sinha, S., Bassik, M. & Levine, B. JNK1-mediated phosphorylation of Bcl-2 regulates starvation-induced autophagy. Mol. Cell 30, 678–688 (2008).
Cao, Y. & Klionsky, D. J. Physiological functions of Atg6/Beclin 1: a unique autophagy-related protein. Cell Res. 17, 839–849 (2007).
Williams, A. et al. Novel targets for Huntington's disease in an mTOR-independent autophagy pathway. Nat. Chem. Biol. 4, 295–305 (2008).
Cárdenas, C. et al. Essential regulation of cell bioenergetics by constitutive InsP3 receptor Ca2+ transfer to mitochondria. Cell 142, 270–283 (2010).
Vicencio, J. M. et al. The inositol 1,4,5-trisphosphate receptor regulates autophagy through its interaction with Beclin 1. Cell Death Differ. 16, 1006–1017 (2009).
Yousefi, S. et al. Calpain-mediated cleavage of Atg5 switches autophagy to apoptosis. Nat. Cell Biol. 8, 1124–1132 (2006).
Ross, C. A. & Poirier, M. A. What is the role of aggregation in neurodegeneration? Nat. Rev. Mol. Cell Biol. 6, 891–898 (2005).
Komatsu, M. et al. Loss of autophagy in the central nervous system causes neurodegeneration in mice. Nature 441, 880–884 (2006).
Hara, T. et al. Suppression of basal autophagy in neural cells causes neurodegenerative disease in mice. Nature 441, 885–880 (2006).
Boya, P. et al. Inhibition of macroautophagy triggers apoptosis. Mol. Cell Biol. 25, 1025–1040 (2005).
Ravikumar, B., Berger, Z., Vacher, C., O'Kane, C. J. & Rubinsztein, D. C. Rapamycin pre-treatment protects against apoptosis. Hum. Mol. Genet. 15, 1209–1216 (2006).
Aguado, C. et al. Laforin, the most common protein mutated in Lafora disease, regulates autophagy. Hum. Mol. Genet. 19, 2867–2876 (2010).
Kirkin, V., McEwan, D. G., Novak, I. & Dikic, I. A role for ubiquitin in selective autophagy. Mol. Cell. 34, 259–269 (2009).
Kirkin, V. et al. A role for NBR1 in autophagosomal degradation of ubiquitinated substrates. Mol. Cell. 33, 505–516 (2009).
Pankiv, S. et al. p62/SQSTM1 binds directly to Atg8/LC3 to facilitate degradation of ubiquitinated protein aggregates by autophagy. J. Biol. Chem. 282, 24131–24145 (2007).
Imarisio, S. et al. Huntington's disease: from pathology and genetics to potential therapies. Biochem. J. 412, 191–209 (2008).
Martinez-Vicente, M. et al. Cargo recognition failure is responsible for inefficient autophagy in Huntington's disease. Nat. Neurosci. 13, 567–576 (2010).
Puls, I. et al. Mutant dynactin in motor neuron disease. Nat. Genet. 33, 455–456 (2003).
Münch, C. et al. Point mutations of the p150 subunit of dynactin (DCTN1) gene in ALS. Neurology 63, 724–726 (2004).
Webb, J. L., Ravikumar, B. & Rubinsztein, D. C. Microtubule disruption inhibits autophagosome-lysosome fusion: implications for studying the roles of aggresomes in polyglutamine diseases. Int. J. Biochem. Cell Biol. 36, 2541–2550 (2004).
Ravikumar, B. et al. Dynein mutations impair autophagic clearance of aggregate-prone proteins. Nat. Genet. 37, 771–776 (2005).
Filimonenko, M. et al. Functional multivesicular bodies are required for autophagic clearance of protein aggregates associated with neurodegenerative disease. J. Cell Biol. 179, 485–500 (2007).
Rusten, E. et al. ESCRTing autophagic clearance of aggregating proteins. Autophagy 4, 233–236 (2008).
Skibinski, G. et al. Mutations in the endosomal ESCRTIII-complex subunit CHMP2B in frontotemporal dementia. Nat. Genet. 37, 806–808 (2005).
Parkinson, N. et al. ALS phenotypes with mutations in CHMP2B (charged multivesicular body protein 2B). Neurology 67, 1074–1077 (2006).
Lee, J.-A., Beigneux, A., Ahmad, S. T., Young, S. G. & Gao. F.-B. ESCRT-III dysfunction causes autophagosome accumulation and neurodegeneration. Curr. Biol. 17, 1561–1567 (2007).
Raben, N., Shea, L., Hill, V. & Plotz, P. Monitoring autophagy in lysosomal storage disorders. Methods Enzymol. 453, 417–449 (2009).
Yue, Z., Friedman, L., Komatsu, M. & Tanaka, K. The cellular pathways of neuronal autophagy and their implication in neurodegenerative diseases. Biochim. Biophys. Acta 1793, 1496–1507 (2009).
Settembre, C. et al. A block of autophagy in lysosomal storage disorders. Hum. Mol. Genet. 17, 119–129 (2008).
Price, D. L., Tanzi, R. E., Borchelt, D. R. & Sisodia, S. S. Alzheimer's disease: genetic studies and transgenic models. Annu. Rev. Genet. 32, 461–493 (1998).
Sherrington, R. et al. Cloning of a gene bearing missense mutations in early-onset familial Alzheimer's disease. Nature 375, 754–760 (1995).
Pimplikar, S., Nixon, R. A., Robakis, N. K., Shien, J. & Tsai, J. H. Amyloid-independent mechanisms in Alzheimer's disease pathogenesis. J. Neurosci. 30, 14946–14954 (2010).
Nixon, R. A. & Yang, D.-S. Autophagy failure in Alzheimer's disease—locating the primary defect. Neurobiol. Dis. 43, 38–45 (2011).
Nixon, R. A. et al. Extensive involvement of autophagy in Alzheimer disease: an immuno-electron microscopy study. J. Neuropathol. Exp. Neurol. 64, 113–122 (2005).
Boland, B. et al. Autophagy induction and autophagosome clearance in neurons: relationship to autophagic pathology in Alzheimer's disease. J. Neurosci. 28, 6926–6937 (2008).
Lee, J. H. et al. Lysosomal proteolysis and autophagy require presenilin 1 and are disrupted by Alzheimer-related PS1 mutations. Cell 141, 1146–1158 (2010).
Yang, D. S. et al. Reversal of autophagy dysfunction in the TgCRND8 mouse model of Alzheimer's disease ameliorates amyloid pathologies and memory deficits. Brain 134, 258–277 (2011).
Mueller-Steiner, S. et al. Antiamyloidogenic and neuroprotective function of cathepsins B: implications for Alzheimer's disease. Neuron 51, 703–714 (2006).
Sun, B. et al. Cystatin C–cathepsin B axis regulates amyloid β levels and associated neuronal deficits in an animal model of Alzheimer's disease. Neuron 60, 247–257 (2008).
Pickford, F. et al. The autophagy related protein beclin 1 shows reduced expression in early Alzheimer's disease and regulates amyloid β accumulation in mice. J. Clin. Invest. 118, 2190–2199 (2008).
Rohn, T. et al. Depletion of Beclin-1 due to proteolytic cleavage by caspases in the Alzheimer's disease brain. Neurobiol. Dis. 43, 68–78 (2010).
Spillantini, M. G. et al. α-Synuclein in Lewy bodies. Nature 388, 839–840 (1997).
Rubinsztein, D. C. The roles of intracellular protein-degradation pathways in neurodegeneration. Nature 443, 780–786 (2006).
Lesage, S. & Brice, A. Parkinson's disease: from monogenic forms to genetic susceptibility factors. Hum. Mol. Genet. 18, R48–R59 (2009).
Youle, R. J. & Narendra, D. P. Mechanisms of mitophagy. Nat. Rev. Mol. Cell Biol. 12, 9–14 (2011).
Narendra, D., Tanaka, A., Suen, D. F. & Youle, R. J. Parkin is recruited selectively to impaired mitochondria and promotes their autophagy. J. Cell Biol. 183, 795–803 (2008).
Narendra, D. et al. PINK1 is selectively stabilized on impaired mitochondria to activate Parkin. PLoS Biol. 8, e1000298 (2010).
Vives-Bauza, C. et al. PINK1-dependent recruitment of Parkin to mitochondria in mitophagy. Proc. Natl Acad. Sci. USA 107, 378–383 (2010).
Geisler, S. et al. PINK1/Parkin-mediated mitophagy is dependent on VDAC1 and p62/SQSTM1. Nat. Cell Biol. 12, 119–131 (2010).
Matsuda, N. et al. PINK1 stabilized by mitochondrial depolarization recruits Parkin to damaged mitochondria and activates latent Parkin for mitophagy. J. Cell Biol. 189, 211–221 (2010).
Kim, P. K., Hailey, D. W., Mullen, R. T. & Lippincott-Schwartz, J. Ubiquitin signals autophagic degradation of cytosolic proteins and peroxisomes. Proc. Natl Acad. Sci. USA 105, 20567–20574 (2008).
Kawajiri, S. et al. PINK1 is recruited to mitochondria with parkin and associates with LC3 in mitophagy. FEBS Lett. 584, 1073–1079 (2010).
Singleton, A. et al. α-Synuclein locus triplication causes Parkinson's disease. Science 302, 5646–5841 (2003).
Winslow, A. et al. α-Synuclein impairs macroautophagy: implications for Parkinson's disease. J. Cell Biol. 190, 1023–1037 (2010).
Cuervo, A. M., Stefanis, L., Fredenburg, R., Lansbury, P. T. & Sulzer, D. Impaired degradation of mutant α-synuclein by chaperone-mediated autophagy. Science 305, 1292–1295 (2004).
Martinez-Vicente, M. et al. Dopamine-modified α-synuclein blocks chaperone-mediated autophagy. J. Clin. Invest. 118, 777–788 (2008).
Fleming, A., Takeshi, N., Tamotsu, Y. & Rubinsztein, D. C. Chemical modulators of autophagy as biological probes and potential therapeutics. Nat. Chem. Biol. 7, 9–17 (2010).
Ravikumar, B. et al. Inhibition of mTOR induces autophagy and reduces toxicity of polyglutamine expansions in fly and mouse models of Huntington disease. Nat. Genet. 36, 585–595 (2004).
Wu, Y. et al. Autophagy plays a protective role during zVAD-induced necrotic cell death. Autophagy 4, 457–466 (2008).
Ravikumar, B., Duden, R. & Rubinsztein, D. C. Aggregate-prone proteins with polyglutamine and polyalanine expansions are degraded by autophagy. Hum. Mol. Genet. 11, 1107–1117 (2002).
Berger, Z. et al. Rapamycin alleviates toxicity of different aggregate-prone proteins. Hum. Mol. Genet. 15, 433–442 (2006).
Menzies, F. et al. Autophagy induction reduces mutant ataxin-3 levels and toxicity in a mouse model of spinocerebellar ataxia type 3. Brain 133, 93–104 (2010).
Menzies, F. & Rubinsztein, D. C. Broadening the therapeutic scope for rapamycin treatment. Autophagy 6, 286–287 (2010).
Sarkar, S. et al. Lithium induces autophagy by inhibiting inositol monophosphatase. J. Cell Biol. 170, 1101–1111 (2005).
Zhang, L. et al. Small molecule regulators of autophagy identified by an image-based high-throughput screen. Proc. Natl Acad. Sci. USA 104, 19023–19028 (2007).
Rose, C. et al. Rilmenidine attenuates toxicity of polyglutamine expansions in a mouse model of Huntington's disease. Hum. Mol. Genet. 19, 2144–2153 (2010).
Sarkar, S., Davies, J. E., Huang, Z., Tunnacliffe, A. & Rubinsztein, D. C. Trehalose, a novel mTOR-independent autophagy enhancer, accelerates the clearance of mutant huntingtin and α-synuclein. J. Biol. Chem. 282, 5641–5652 (2007).
Tanaka, M. et al. Trehalose alleviates polyglutamine-mediated pathology in a mouse model of Huntington disease. Nat. Med. 10, 148–154 (2004).
Sarkar, S. et al. A rational mechanism for combination treatment of Huntington's disease using lithium and rapamycin. Hum. Mol. Genet. 17, 170–178 (2008).
Sarkar, S. et al. Small molecules enhance autophagy and reduce toxicity in Huntington's disease models. Nat. Chem. Biol. 6, 331–338 (2007).
Massey, A. C., Zhang, C. & Cuervo, A. M. Chaperone-mediated autophagy in aging and disease. Curr. Top. Dev. Biol. 73, 205–235 (2006).
Yang, Z. & Klionsky, D. J. Mammalian autophagy: core molecular machinery and signaling regulation. Curr. Opin. Cell Biol. 22, 124–131 (2010).
Nakagawa, I. et al. Autophagy defends cells against invading group A Streptococcus. Science 306, 1037–1040 (2004).
Gutierrez, M. et al. Autophagy is a defense mechanism inhibiting BCG and Mycobacterium tuberculosis survival in infected macrophages. Cell 119, 753–766 (2004).
Ogawa, M. et al. Escape of intracellular Shigella from autophagy. Science 307, 727–731 (2005).
Birmingham, C., Smith, A., Bakowski, M., Yoshimori, T. & Brummel, J. Autophagy controls Salmonella infection in response to damage to the Salmonella-containing vacuole. J. Biol. Chem. 281, 11374–11383 (2006).
Py, B., Lipinski, M. & Yuan, J. Autophagy limits Listeria monocytogenes intracellular growth in the early phase of primary infection. Autophagy 3, 117–125 (2007).
Andrade, R., Wessendarp, M., Gubbels, M., Striepen, B. & Subauste, C. CD40 induces macrophage anti-Toxoplasma gondii activity by triggering autophagy-dependent fusion of pathogen-containing vacuoles and lysosomes. J. Clin. Invest. 116, 2366–2377 (2006).
Ling, Y. et al. Vacuolar and plasma membrane stripping and autophagic elimination of Toxoplasma gondii in primed effector macrophages. J. Exp. Med. 203, 2063–2071 (2006).
Rioux, J. et al. Genome-wide association study identifies new susceptibility loci for Crohn disease and implicates autophagy in disease pathogenesis. Nat. Genet. 39, 596–604 (2007).
Parkes, M. et al. Sequence variants in the autophagy gene IRGM and multiple other replicating loci contribute to Crohn's disease susceptibility. Nat. Genet. 39, 830–832 (2007).
Terman, A. & Brunk, B. Autophagy in cardiac myocyte homeostasis, aging, and pathology. Cardiovasc. Res. 68, 355–365 (2005).
Liang, X. et al. Induction of autophagy and inhibition of tumorigenesis by beclin 1. Nature 402, 672–676 (1999).
Marino, G. et al. Tissue-specific autophagy alterations and increased tumorigenesis in mice deficient in Atg4C/autophagin-3. J. Biol. Chem. 282, 18573–18583 (2007).
Degenhardt, K. et al. Autophagy promotes tumor cell survival and restricts necrosis, inflammation, and tumorigenesis. Cancer Cell 10, 51–64 (2006).
Acknowledgements
The authors are grateful to the Wellcome Trust for providing funding for the studentship of H. Harris and a Senior Fellowship for D. C. Rubinsztein, to the Medical Research Council for providing a program grant and a Medical Research Council/Wellcome Trust Strategic Award to D. C. Rubinsztein, and to the National Institute for Health Research Biomedical Research Center at Addenbrooke's Hospital. We also thank M. Renna for helpful comments on the manuscript.
Author information
Authors and Affiliations
Contributions
Both authors researched data for the article, discussed the content, wrote the text, and reviewed and edited the manuscript before submission.
Corresponding author
Ethics declarations
Competing interests
D. C. Rubinsztein has received grant support from Lilly and Wyeth. H. Harris declares no competing interests.
Rights and permissions
About this article
Cite this article
Harris, H., Rubinsztein, D. Control of autophagy as a therapy for neurodegenerative disease. Nat Rev Neurol 8, 108–117 (2012). https://doi.org/10.1038/nrneurol.2011.200
Published:
Issue Date:
DOI: https://doi.org/10.1038/nrneurol.2011.200
This article is cited by
-
The protective effect of inhibiting mitochondrial fission on the juvenile rat brain following PTZ kindling through inhibiting the BCL2L13/LC3 mitophagy pathway
Metabolic Brain Disease (2023)
-
Metformin and Trehalose-Modulated Autophagy Exerts a Neurotherapeutic Effect on Parkinsonʼs Disease
Molecular Neurobiology (2023)
-
Transcription factor EB-mediated mesenchymal stem cell therapy induces autophagy and alleviates spinocerebellar ataxia type 3 defects in neuronal cells model
Cell Death & Disease (2022)
-
Different Roles of Beclin1 in the Interaction Between Glia and Neurons after Exposure to Morphine and the HIV- Trans-Activator of Transcription (Tat) Protein
Journal of Neuroimmune Pharmacology (2022)
-
Role of lysosomes in physiological activities, diseases, and therapy
Journal of Hematology & Oncology (2021)