Key Points
-
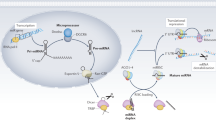
MicroRNAs (miRNAs) are key players in kidney development and physiology
-
Transforming growth factor β1 is a major regulator of kidney fibrosis; its signalling is finely regulated by miRNAs
-
miRNAs contribute to both the induction and progression of chronic kidney disease (CKD)
-
Current translational research on miRNAs in kidney disease is mainly focused on developing reliable biomarkers for diagnosis and prognosis of CKD and renal transplantation
-
miRNAs represent novel therapeutic targets for CKD, but delivery and safety issues must be taken into account before translation into clinical practice
Abstract
MicroRNAs (miRNAs) are small non-coding RNA molecules that regulate gene expression. They have important roles during kidney development, homeostasis and disease. In particular, miRNAs participate in the onset and progression of tubulointerstitial sclerosis and end-stage glomerular lesions that occur in various forms of chronic kidney disease (CKD). Therefore, miRNAs represent potential new therapeutic targets for a debilitating disease that continues to increase in prevalence worldwide and for which fully effective therapies are lacking. Several lines of research aimed at improving common CKD diagnostic tools and avoiding invasive kidney biopsies have also identified circulating miRNAs as possible diagnostic and even prognostic biomarkers of kidney disease. This Review discusses current understanding of the function of miRNAs in CKD, focusing on functions specifically involved in the transforming growth factor β1 pathway, which is activated in CKD. miRNAs that, according to available evidence, seem to be involved in diabetic nephropathy, IgA nephropathy, lupus nephritis, polycystic kidney disease and graft rejection, are also discussed.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
References
Couser, W. G., Remuzzi, G., Mendis, S. & Tonelli, M. The contribution of chronic kidney disease to the global burden of major noncommunicable diseases. Kidney Int. 80, 1258–1270 (2011).
Liu, Y. Cellular and molecular mechanisms of renal fibrosis. Nat. Rev. Nephrol. 7, 684–696 (2011).
López-Hernández, F. J. & López-Novoa, J. M. Role of TGF-β in chronic kidney disease: an integration of tubular, glomerular and vascular effects. Cell Tissue Res. 347, 141–154 (2012).
Ruggenenti, P., Cravedi, P. & Remuzzi, G. Mechanisms and treatment of CKD. J. Am. Soc. Nephrol. 23, 1917–1928 (2012).
Meng, X. M., Chung, A. C. & Lan, H. Y. Role of the TGF-β/BMP-7/Smad pathways in renal diseases. Clin. Sci. (Lond.) 124, 243–254 (2013).
Lorenzen, J. M. & Thum, T. Circulating and urinary microRNAs in kidney disease. Clin. J. Am. Soc. Nephrol. 7, 1528–1533 (2012).
Saikumar, J., Ramachandran, K. & Vaidya, V. S. Noninvasive micromarkers. Clin. Chem. 60, 1158–1173 (2014).
Kozomara, A. & Griffiths-Jones, S. miRBase: annotating high confidence microRNAs using deep sequencing data. Nucleic Acids Res. 42, D68–D73 (2014).
Friedman, L. M. & Avraham, K. B. MicroRNAs and epigenetic regulation in the mammalian inner ear: implications for deafness. Mamm. Genome 20, 581–603 (2009).
Bartel, D. P. MicroRNAs: target recognition and regulatory functions. Cell 136, 215–233 (2009).
Djuranovic, S., Nahvi, A. & Green, R. A parsimonious model for gene regulation by miRNAs. Science 331, 550–553 (2011).
Kriegel, A. J., Liu, Y., Fang, Y., Ding, X. & Liang, M. The miR-29 family: genomics, cell biology, and relevance to renal and cardiovascular injury. Physiol. Genomics 44, 237–244 (2012).
Krol, J., Loedige, I. & Filipowicz, W. The widespread regulation of microRNA biogenesis, function and decay. Nat. Rev. Genet. 11, 597–610 (2010).
Baskerville, S. & Bartel, D. P. Microarray profiling of microRNAs reveals frequent coexpression with neighboring miRNAs and host genes. RNA 11, 241–247 (2005).
Kato, M. et al. Post-transcriptional up-regulation of Tsc-22 by Ybx1, a target of miR-216a, mediates TGF-β-induced collagen expression in kidney cells. J. Biol. Chem. 285, 34004–34015 (2010).
Alvarez, M. L., Khosroheidari, M., Eddy, E. & Kiefer, J. Role of microRNA 1207–5P and its host gene, the long non-coding RNA Pvt1, as mediators of extracellular matrix accumulation in the kidney: implications for diabetic nephropathy. PLoS ONE 8, e77468 (2013).
Lytle, J. R., Yario, T. A. & Steitz, J. A. Target mRNAs are repressed as efficiently by microRNA-binding sites in the 5′ UTR as in the 3′ UTR. Proc. Natl Acad. Sci. USA 104, 9667–9672 (2007).
Djuranovic, S., Nahvi, A. & Green, R. miRNA-mediated gene silencing by translational repression followed by mRNA deadenylation and decay. Science 336, 237–240 (2012).
Huntzinger, E. & Izaurralde, E. Gene silencing by microRNAs: contributions of translational repression and mRNA decay. Nat. Rev. Genet. 12, 99–110 (2011).
Ørom, U. A., Nielsen, F. C. & Lund, A. H. MicroRNA-10a binds the 5′UTR of ribosomal protein mRNAs and enhances their translation. Mol. Cell 30, 460–471 (2008).
Place, R. F., Li, L. C., Pookot, D., Noonan, E. J. & Dahiya, R. MicroRNA-373 induces expression of genes with complementary promoter sequences. Proc. Natl Acad. Sci. USA 105, 1608–1613 (2008).
Chau, B. N. et al. MicroRNA-21 promotes fibrosis of the kidney by silencing metabolic pathways. Sci. Transl. Med. 4, 121ra18 (2012).
Denby, L. et al. MicroRNA-214 antagonism protects against renal fibrosis. J. Am. Soc. Nephrol. 25, 65–80 (2014).
Zhang, Z., Qin, Y. W., Brewer, G. & Jing, Q. MicroRNA degradation and turnover: regulating the regulators. Wiley Interdiscip. Rev. RNA 3, 593–600 (2012).
Sun, Y. et al. Development of a micro-array to detect human and mouse microRNAs and characterization of expression in human organs. Nucleic Acids Res. 32, e188 (2004).
Landgraf, P. et al. A mammalian microRNA expression atlas based on small RNA library sequencing. Cell 129, 1401–1414 (2007).
Tang, Y., Liu, D., Zhang, L., Ingvarsson, S. & Chen, H. Quantitative analysis of miRNA expression in seven human foetal and adult organs. PLoS ONE 6, e28730 (2011).
Ho, J. et al. The pro-apoptotic protein Bim is a microRNA target in kidney progenitors. J. Am. Soc. Nephrol. 22, 1053–1063 (2011).
Nagalakshmi, V. K. et al. Dicer regulates the development of nephrogenic and ureteric compartments in the mammalian kidney. Kidney Int. 79, 317–330 (2011).
Marrone, A. K. et al. MicroRNA-17∼92 is required for nephrogenesis and renal function. J. Am. Soc. Nephrol. 25, 1440–1452 (2014).
Pastorelli, L. M. et al. Genetic analyses reveal a requirement for Dicer1 in the mouse urogenital tract. Mamm. Genome 20, 140–151 (2009).
Patel, V. et al. MicroRNAs regulate renal tubule maturation through modulation of Pkd1. J. Am. Soc. Nephrol. 23, 1941–1948 (2012).
Shi, S. et al. Podocyte-selective deletion of Dicer induces proteinuria and glomerulosclerosis. J. Am. Soc. Nephrol. 19, 2159–2169 (2008).
Harvey, S. J. et al. Podocyte-specific deletion of Dicer alters cytoskeletal dynamics and causes glomerular disease. J. Am. Soc. Nephrol. 19, 2150–2158 (2008).
Ho, J. et al. Podocyte-specific loss of functional microRNAs leads to rapid glomerular and tubular injury. J. Am. Soc. Nephrol. 19, 2069–2075 (2008).
Kramann, R. & Moeller, M. J. The next level of complexity: post-transcriptional regulation by microRNAs. Kidney Int. 80, 692–693 (2011).
Zhdanova, O. et al. The inducible deletion of Drosha and microRNAs in mature podocytes results in a collapsing glomerulopathy. Kidney Int. 80, 719–730 (2011).
Wei, Q. et al. Targeted deletion of Dicer from proximal tubules protects against renal ischemia-reperfusion injury. J. Am. Soc. Nephrol. 21, 756–761 (2010).
Lorenzen, J. M., Haller, H. & Thum, T. MicroRNAs as mediators and therapeutic targets in chronic kidney disease. Nat. Rev. Nephrol. 7, 286–294 (2011).
Sequeira-Lopez, M. L. et al. The microRNA-processing enzyme Dicer maintains juxtaglomerular cells. J. Am. Soc. Nephrol. 21, 460–467 (2010).
Mladinov, D., Liu, Y., Mattson, D. L. & Liang, M. MicroRNAs contribute to the maintenance of cell-type-specific physiological characteristics: miR-192 targets Na+/K+-ATPase β1. Nucleic Acids Res. 41, 1273–1283 (2013).
Elvira-Matelot, E. et al. Regulation of WNK1 expression by miR-192 and aldosterone. J. Am. Soc. Nephrol. 21, 1724–1731 (2010).
Edinger, R. S. et al. Aldosterone regulates microRNAs in the cortical collecting duct to alter sodium transport. J. Am. Soc. Nephrol. http://dx.doi.org/10.1681/ASN.2013090931.
Huang, W. et al. Tonicity-responsive microRNAs contribute to the maximal induction of osmoregulatory transcription factor OREBP in response to high-NaCl hypertonicity. Nucleic Acids Res. 39, 475–485 (2011).
Lin, D. H., Yue, P., Pan, C., Sun, P. & Wang, W. H. MicroRNA 802 stimulates ROMK channels by suppressing caveolin-1. J. Am. Soc. Nephrol. 22, 1087–1098 (2011).
Lin, D. H., Yue, P., Zhang, C. & Wang, W. H. MicroRNA-194 (miR-194) regulates ROMK channel activity by targeting intersectin 1. Am. J. Physiol. Renal Physiol. 306, F53–F60 (2014).
Gong, Y. et al. Claudin-14 regulates renal Ca++ transport in response to CaSR signalling via a novel microRNA pathway. EMBO J. 31, 1999–2012 (2012).
Gong, Y. & Hou, J. Claudin-14 underlies Ca++-sensing receptor-mediated Ca++ metabolism via NFAT-microRNA-based mechanisms. J. Am. Soc. Nephrol. 25, 745–760 (2014).
Wu, L. & Derynck, R. Essential role of TGF-β signaling in glucose-induced cell hypertrophy. Dev. Cell 17, 35–48 (2009).
Rüster, C. & Wolf, G. Angiotensin II as a morphogenic cytokine stimulating renal fibrogenesis. J. Am. Soc. Nephrol. 22, 1189–1199 (2011).
Jiang, F., Liu, G. S., Dusting, G. J. & Chan, E. C. NADPH oxidase-dependent redox signaling in TGF-β-mediated fibrotic responses. Redox Biol. 2, 267–272 (2014).
Boor, P. & Floege, J. Chronic kidney disease growth factors in renal fibrosis. Clin. Exp. Pharmacol. Physiol. 38, 441–450 (2011).
Zeisberg, M. et al. BMP-7 counteracts TGF-β1-induced epithelial-to-mesenchymal transition and reverses chronic renal injury. Nat. Med. 9, 964–968 (2003).
Eddy, A. A. The origin of scar-forming kidney myofibroblasts. Nat. Med. 19, 964–966 (2013).
LeBleu, V. S. et al. Origin and function of myofibroblasts in kidney fibrosis. Nat. Med. 19, 1047–1053 (2013).
Ha, H., Oh, E. Y. & Lee, H. B. The role of plasminogen activator inhibitor 1 in renal and cardiovascular diseases. Nat. Rev. Nephrol. 5, 203–211 (2009).
Samarakoon, R., Overstreet, J. M., Higgins, S. P. & Higgins, P. J. TGF-β1 → SMAD/p53/USF2 → PAI-1 transcriptional axis in ureteral obstruction-induced renal fibrosis. Cell Tissue Res. 347, 117–128 (2012).
Moustakas, A., Souchelnytskyi, S. & Heldin, C. H. Smad regulation in TGF-β signal transduction. J. Cell Sci. 114, 4359–4369 (2001).
Samarakoon, R. et al. Induction of renal fibrotic genes by TGF-β1 requires EGFR activation, p53 and reactive oxygen species. Cell. Signal. 25, 2198–2209 (2013).
Derynck, R. & Zhang, Y. E. Smad-dependent and Smad-independent pathways in TGF-β family signalling. Nature 425, 577–584 (2003).
Inman, G. J., Nicolás, F. J. & Hill, C. S. Nucleocytoplasmic shuttling of Smads 2, 3, and 4 permits sensing of TGF-β receptor activity. Mol. Cell 10, 283–294 (2002).
Butz, H., Rácz, K., Hunyady, L. & Patócs, A. Crosstalk between TGF-β signaling and the microRNA machinery. Trends Pharmacol. Sci. 33, 382–393 (2012).
Davis, B. N., Hilyard, A. C., Nguyen, P. H., Lagna, G. & Hata, A. Smad proteins bind a conserved RNA sequence to promote microRNA maturation by Drosha. Mol. Cell 39, 373–384 (2010).
Davis, B. N., Hilyard, A. C., Lagna, G. & Hata, A. SMAD proteins control DROSHA-mediated microRNA maturation. Nature 454, 56–61 (2008).
Zhong, X., Chung, A. C., Chen, H. Y., Meng, X. M. & Lan, H. Y. Smad3-mediated upregulation of miR-21 promotes renal fibrosis. J. Am. Soc. Nephrol. 22, 1668–1681 (2011).
Qin, W. et al. TGF-β/Smad3 signaling promotes renal fibrosis by inhibiting miR-29. J. Am. Soc. Nephrol. 22, 1462–1474 (2011).
Chung, A. C., Huang, X. R., Meng, X. & Lan, H. Y. miR-192 mediates TGF-β/Smad3-driven renal fibrosis. J. Am. Soc. Nephrol. 21, 1317–1325 (2010).
Kato, M. et al. MicroRNA-192 in diabetic kidney glomeruli and its function in TGF-β-induced collagen expression via inhibition of E-box repressors. Proc. Natl Acad. Sci. USA 104, 3432–3437 (2007).
Putta, S. et al. Inhibiting microRNA-192 ameliorates renal fibrosis in diabetic nephropathy. J. Am. Soc. Nephrol. 23, 458–469 (2012).
Wang, B. et al. E-cadherin expression is regulated by miR-192/215 by a mechanism that is independent of the profibrotic effects of transforming growth factor-β. Diabetes 59, 1794–1802 (2010).
Sun, L. et al. Low-dose paclitaxel ameliorates fibrosis in the remnant kidney model by down-regulating miR-192. J. Pathol. 225, 364–377 (2011).
Kato, M. et al. TGF-β induces acetylation of chromatin and of Ets-1 to alleviate repression of miR-192 in diabetic nephropathy. Sci. Signal. 6, ra43 (2013).
Jenkins, R. H., Martin, J., Phillips, A. O., Bowen, T. & Fraser, D. J. Transforming growth factor β1 represses proximal tubular cell microRNA-192 expression through decreased hepatocyte nuclear factor DNA binding. Biochem. J. 443, 407–416 (2012).
Igarashi, P., Shao, X., McNally, B. T. & Hiesberger, T. Roles of HNF-1β in kidney development and congenital cystic diseases. Kidney Int. 68, 1944–1947 (2005).
Kato, M. et al. A microRNA circuit mediates transforming growth factor-β1 autoregulation in renal glomerular mesangial cells. Kidney Int. 80, 358–368 (2011).
Deshpande, S. D. et al. Transforming growth factor-β-induced cross talk between p53 and a microRNA in the pathogenesis of diabetic nephropathy. Diabetes 62, 3151–3162 (2013).
Du, B. et al. High glucose down-regulates miR-29a to increase collagen IV production in HK-2 cells. FEBS Lett. 584, 811–816 (2010).
Wang, B. et al. Suppression of microRNA-29 expression by TGF-β1 promotes collagen expression and renal fibrosis. J. Am. Soc. Nephrol. 23, 252–265 (2012).
Ramdas, V., McBride, M., Denby, L. & Baker, A. H. Canonical transforming growth factor-β signaling regulates disintegrin metalloprotease expression in experimental renal fibrosis via miR-29. Am. J. Pathol. 183, 1885–1896 (2013).
Chen, H. Y. et al. MicroRNA-29b inhibits diabetic nephropathy in db/db mice. Mol. Ther. 22, 842–853 (2014).
Macconi, D. et al. MicroRNA-324-3p promotes renal fibrosis and is a target of ACE inhibition. J. Am. Soc. Nephrol. 23, 1496–1505 (2012).
Wang, Q. et al. MicroRNA-377 is up-regulated and can lead to increased fibronectin production in diabetic nephropathy. FASEB J. 22, 4126–4135 (2008).
Tang, O., Chen, X. M., Shen, S., Hahn, M. & Pollock, C. A. MiRNA-200b represses transforming growth factor-β1-induced EMT and fibronectin expression in kidney proximal tubular cells. Am. J. Physiol. Renal Physiol. 304, F1266–F1273 (2013).
Xiong, M. et al. The miR-200 family regulates TGF-β1-induced renal tubular epithelial to mesenchymal transition through Smad pathway by targeting ZEB1 and ZEB2 expression. Am. J. Physiol. Renal Physiol. 302, F369–F379 (2012).
Krupa, A. et al. Loss of MicroRNA-192 promotes fibrogenesis in diabetic nephropathy. J. Am. Soc. Nephrol. 21, 438–447 (2010).
Jiang, L. et al. A microRNA-30e/mitochondrial uncoupling protein 2 axis mediates TGF-β1-induced tubular epithelial cell extracellular matrix production and kidney fibrosis. Kidney Int. 84, 285–296 (2013).
Denby, L. et al. miR-21 and miR-214 are consistently modulated during renal injury in rodent models. Am. J. Pathol. 179, 661–672 (2011).
Zhang, Z. et al. MicroRNA-21 protects from mesangial cell proliferation induced by diabetic nephropathy in db/db mice. FEBS Lett. 583, 2009–2014 (2009).
Kato, M. et al. TGF-β activates Akt kinase through a microRNA-dependent amplifying circuit targeting PTEN. Nat. Cell Biol. 11, 881–889 (2009).
Mahimainathan, L., Das, F., Venkatesan, B. & Choudhury, G. G. Mesangial cell hypertrophy by high glucose is mediated by downregulation of the tumor suppressor PTEN. Diabetes 55, 2115–2125 (2006).
Kantharidis, P., Wang, B., Carew, R. M. & Lan, H. Y. Diabetes complications: the microRNA perspective. Diabetes 60, 1832–1837 (2011).
Zhong, X. et al. miR-21 is a key therapeutic target for renal injury in a mouse model of type 2 diabetes. Diabetologia 56, 663–674 (2013).
Park, J. T. et al. FOG2 protein down-regulation by transforming growth factor-β1-induced microRNA-200b/c leads to Akt kinase activation and glomerular mesangial hypertrophy related to diabetic nephropathy. J. Biol. Chem. 288, 22469–22480 (2013).
Li, R. et al. The microRNA miR-433 promotes renal fibrosis by amplifying the TGF-β/Smad3-Azin1 pathway. Kidney Int. 84, 1129–1144 (2013).
Wang, B. et al. Transforming growth factor-β1-mediated renal fibrosis is dependent on the regulation of transforming growth factor receptor 1 expression by let-7b. Kidney Int. 85, 352–361 (2014).
Brennan, E. P. et al. Lipoxins attenuate renal fibrosis by inducing let-7c and suppressing TGFβR1. J. Am. Soc. Nephrol. 24, 627–637 (2013).
Wang, J. Y. et al. Tongxinluo ameliorates renal structure and function by regulating miR-21-induced epithelial-mesenchymal transition in diabetic nephropathy. Am. J. Physiol. Renal Physiol. 306, F486–F495 (2014).
Lin, C. L. et al. MicroRNA-29a promotion of nephrin acetylation ameliorates hyperglycemia-induced podocyte dysfunction. J. Am. Soc. Nephrol. 25, 1698–1709 (2014).
Long, J., Wang, Y., Wang, W., Chang, B. H. & Danesh, F. R. MicroRNA-29c is a signature microRNA under high glucose conditions that targets Sprouty homolog 1, and its in vivo knockdown prevents progression of diabetic nephropathy. J. Biol. Chem. 286, 11837–11848 (2011).
Fu, Y. et al. Regulation of NADPH oxidase activity is associated with miRNA-25-mediated NOX4 expression in experimental diabetic nephropathy. Am. J. Nephrol. 32, 581–589 (2010).
Long, J., Wang, Y., Wang, W., Chang, B. H. & Danesh, F. R. Identification of microRNA-93 as a novel regulator of vascular endothelial growth factor in hyperglycemic conditions. J. Biol. Chem. 285, 23457–23465 (2010).
Argyropoulos, C. et al. Urinary microRNA profiling in the nephropathy of type 1 diabetes. PLoS ONE 8, e54662 (2013).
Barutta, F. et al. Urinary exosomal microRNAs in incipient diabetic nephropathy. PLoS ONE 8, e73798 (2013).
Szeto, C. C. et al. Micro-RNA expression in the urinary sediment of patients with chronic kidney diseases. Dis. Markers 33, 137–144 (2012).
Serino, G., Sallustio, F., Cox, S. N., Pesce, F. & Schena, F. P. Abnormal miR-148b expression promotes aberrant glycosylation of IgA1 in IgA nephropathy. J. Am. Soc. Nephrol. 23, 814–824 (2012).
Wang, G. et al. Intrarenal expression of microRNAs in patients with IgA nephropathy. Lab. Invest. 90, 98–103 (2010).
Wang, G. et al. Elevated levels of miR-146a and miR-155 in kidney biopsy and urine from patients with IgA nephropathy. Dis. Markers 30, 171–179 (2011).
Bao, H. et al. MiR-223 downregulation promotes glomerular endothelial cell activation by upregulating importin α4 and α5 in IgA nephropathy. Kidney Int. 85, 624–635 (2014).
Wang, G. et al. Urinary miR-21, miR-29, and miR-93: novel biomarkers of fibrosis. Am. J. Nephrol. 36, 412–418 (2012).
Dai, Y. et al. Comprehensive analysis of microRNA expression patterns in renal biopsies of lupus nephritis patients. Rheumatol. Int. 29, 749–754 (2009).
Zhou, H. et al. miR-150 promotes renal fibrosis in lupus nephritis by downregulating SOCS1. J. Am. Soc. Nephrol. 24, 1073–1087 (2013).
Tomasoni, S. & Benigni, A. Post-transcriptional gene regulation makes things clearer in renal fibrosis. J. Am. Soc. Nephrol. 24, 1026–1028 (2013).
Lu, J. et al. Glomerular and tubulointerstitial miR-638, miR-198 and miR-146a expression in lupus nephritis. Nephrology (Carlton) 17, 346–351 (2012).
Carlsen, A. L. et al. Circulating microRNA expression profiles associated with systemic lupus erythematosus. Arthritis Rheum. 65, 1324–1334 (2013).
Guan, J. et al. Urinary sediment ICAM-1 level in lupus nephritis. Lupus 21, 1190–1195 (2012).
Te, J. L. et al. Identification of unique microRNA signature associated with lupus nephritis. PLoS ONE 5, e10344 (2010).
Li, Y., Fang, X. & Li, Q. Z. Biomarker profiling for lupus nephritis. Genomics Proteomics Bioinformatics 11, 158–165 (2013).
Patel, V. et al. miR-17∼92 miRNA cluster promotes kidney cyst growth in polycystic kidney disease. Proc. Natl Acad. Sci. USA 110, 10765–10770 (2013).
Sun, H. et al. MicroRNA-17 post-transcriptionally regulates polycystic kidney disease-2 gene and promotes cell proliferation. Mol. Biol. Rep. 37, 2951–2958 (2010).
Hiesberger, T. et al. Mutation of hepatocyte nuclear factor-1β inhibits Pkhd1 gene expression and produces renal cysts in mice. J. Clin. Invest. 113, 814–825 (2004).
Tran, U. et al. The RNA-binding protein bicaudal C regulates polycystin 2 in the kidney by antagonizing miR-17 activity. Development 137, 1107–1116 (2010).
Piazzon, N., Maisonneuve, C., Guilleret, I., Rotman, S. & Constam, D. B. Bicc1 links the regulation of cAMP signaling in polycystic kidneys to microRNA-induced gene silencing. J. Mol. Cell Biol. 4, 398–408 (2012).
Pandey, P. et al. Microarray-based approach identifies microRNAs and their target functional patterns in polycystic kidney disease. BMC Genomics 9, 624 (2008).
Pandey, P., Qin, S., Ho, J., Zhou, J. & Kreidberg, J. A. Systems biology approach to identify transcriptome reprogramming and candidate microRNA targets during the progression of polycystic kidney disease. BMC Syst. Biol. 5, 56 (2011).
Dweep, H., Sticht, C., Kharkar, A., Pandey, P. & Gretz, N. Parallel analysis of mRNA and microRNA microarray profiles to explore functional regulatory patterns in polycystic kidney disease: using PKD/Mhm rat model. PLoS ONE 8, e53780 (2013).
Ben-Dov, I. Z. et al. Urine microRNA as potential biomarkers of autosomal dominant polycystic kidney disease progression: description of miRNA profiles at baseline. PLoS ONE 9, e86856 (2014).
Sui, W. et al. Microarray analysis of MicroRNA expression in acute rejection after renal transplantation. Transpl. Immunol. 19, 81–85 (2008).
Anglicheau, D. et al. MicroRNA expression profiles predictive of human renal allograft status. Proc. Natl Acad. Sci. USA 106, 5330–5335 (2009).
Wilflingseder, J. et al. miRNA profiling discriminates types of rejection and injury in human renal allografts. Transplantation 95, 835–841 (2013).
Lorenzen, J. M. et al. Urinary miR-210 as a mediator of acute T-cell mediated rejection in renal allograft recipients. Am. J. Transplant. 11, 2221–2227 (2011).
Danger, R. et al. Upregulation of miR-142-3p in peripheral blood mononuclear cells of operationally tolerant patients with a renal transplant. J. Am. Soc. Nephrol. 23, 597–606 (2012).
Scian, M. J. et al. MicroRNA profiles in allograft tissues and paired urines associate with chronic allograft dysfunction with IF/TA. Am. J. Transplant. 11, 2110–2122 (2011).
Ben-Dov, I. Z. et al. MicroRNA sequence profiles of human kidney allografts with or without tubulointerstitial fibrosis. Transplantation 94, 1086–1094 (2012).
Maluf, D. G. et al. The urine microRNA profile may help monitor post-transplant renal graft function. Kidney Int. 85, 439–449 (2014).
Janssen, H. L. et al. Treatment of HCV infection by targeting microRNA. N. Engl. J. Med. 368, 1685–1694 (2013).
Acknowledgements
We would like to thank S. Tomasoni and D. Macconi for helpful discussions regarding the manuscript.
Author information
Authors and Affiliations
Contributions
P.T. researched data for the article. P.T. and A.B. contributed substantially to discussion of the article's content and wrote the article. A.B. and G.R. reviewed and edited the manuscript before submission.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Rights and permissions
About this article
Cite this article
Trionfini, P., Benigni, A. & Remuzzi, G. MicroRNAs in kidney physiology and disease. Nat Rev Nephrol 11, 23–33 (2015). https://doi.org/10.1038/nrneph.2014.202
Published:
Issue Date:
DOI: https://doi.org/10.1038/nrneph.2014.202
This article is cited by
-
Mechanisms of norcantharidin against renal tubulointerstitial fibrosis
Pharmacological Reports (2024)
-
Identification and Functional Mechanism Verification of Novel MicroRNAs Associated with the Fibrosis Progression in Chronic Kidney Disease
Biochemical Genetics (2024)
-
Ultra-sensitive electrochemiluminescent biosensor for miRNA based on CRISPR/Cas13a trans-cleavage-triggered hybridization chain reaction and magnetic-assisted enrichment
Microchimica Acta (2023)
-
Emerging role of tumor suppressor p53 in acute and chronic kidney diseases
Cellular and Molecular Life Sciences (2022)
-
Identification of miRNA-mRNA network and immune-related gene signatures in IgA nephropathy by integrated bioinformatics analysis
BMC Nephrology (2021)