Key Points
-
Stimulator of interferon genes (STING) is activated by binding to cyclic dinucleotides (CDNs), which results in potent cytokine production. CDNs are produced by certain intracellular bacteria and are generated by the cyclic GMP–AMP synthase (cGAS) following binding to cytosolic DNA species, such as viral DNA.
-
STING-inducible innate immune molecules are essential for protection of the host against pathogens and are important for the stimulation of adaptive immunity.
-
Self-DNA, for example from the nucleus or mitochondria, can leak into the cytosolic compartment and stimulate STING activity to cause autoinflammatory disease. Certain mutations in the gene encoding STING can cause the protein to become permanently active and similarly induce autoinflammatory responses.
-
STING can be activated in phagocytes by DNA released from engulfed tumour cells and drive the production of cytokines necessary for generating robust antitumour T cell responses.
-
DNA-damaging agents can cause the release of nuclear DNA into the cytosol that stimulates STING-dependent cytokine production and phagocyte infiltration. This may be essential for eliminating damaged cells and generating antitumour T cell responses, but chronic stimulation may also promote inflammation-aggravated cancer.
-
STING agonists exert potent antitumour activity and may be effective, novel adjuvants in vaccine formulations. In contrast, inhibitors of STING signalling may be beneficial for the treatment of autoinflammatory disease, such as systemic lupus erythematosus (SLE), Aicardi–Goutières syndrome (AGS) and STING-associated vasculopathy with onset in infancy (SAVI).
Abstract
The rapid detection of microbial agents is essential for the effective initiation of host defence mechanisms against infection. Understanding how cells detect cytosolic DNA to trigger innate immune gene transcription has important implications — not only for comprehending the immune response to pathogens but also for elucidating the causes of autoinflammatory disease involving the sensing of self-DNA and the generation of effective antitumour adaptive immunity. The discovery of the STING (stimulator of interferon genes)-controlled innate immune pathway, which mediates cytosolic DNA-induced signalling events, has recently provided important insights into these processes, opening the way for the development of novel immunization regimes, as well as therapies to treat autoinflammatory disease and cancer.
Similar content being viewed by others
Main
Stimulator of interferon genes (STING; also known as MITA and MPYS, and encoded by TMEM173) is a signalling molecule associated with the endoplasmic reticulum (ER) and is essential for controlling the transcription of numerous host defence genes, including type I interferons (IFNs) and pro-inflammatory cytokines, following the recognition of aberrant DNA species or cyclic dinucleotides (CDNs) in the cytosol of the cell1,2,3. The sources of DNA that induce CDNs include the genome of invading pathogens, such as herpes simplex virus 1 (HSV1), whereas certain bacteria can secrete CDNs following infection of the host2,4. Recent reports have indicated that potent activators of the STING pathway may also include self-DNA that has leaked from the nucleus of the host cell, perhaps following cell division or as a consequence of DNA damage5. Such DNA species may be responsible for causing various autoinflammatory diseases, such as systemic lupus erythematosus (SLE) or Aicardi–Goutières syndrome (AGS), and may influence inflammation-associated cancer5. Polymorphisms in TMEM173 have been reported to influence STING-dependent innate immune signalling and might affect susceptibility to pathogen infection and contribute to severe inflammatory disorders (Box 1).
STING signalling has now been shown to be essential for protecting the cell against a variety of pathogens and even against the development of cancer by promoting antitumour immune responses6,7. Here, we review recent insights into the regulation of STING signalling and its role in autoinflammatory diseases and cancer. We also discuss new opportunities for the development of novel anti-inflammatory and antitumour compounds that specifically target this key signalling pathway.
The STING signalling pathway
Activation of STING. STING is a 379 amino acid protein, consisting of several transmembrane regions, that is expressed in various endothelial and epithelial cell types, as well as in haematopoietic cells, such as T cells, macrophages and dendritic cells (DCs), including plasmacytoid dendritic cells (pDCs)1,2. Homologues of STING have been identified in different eukaryotic species as well as in invertebrates (Box 2). Early studies showed that STING stimulates the transcription of numerous innate immune genes in response to invading DNA viruses, certain bacteria or transfected DNA1,2,8,9. Further investigations clarified that STING is robustly activated by CDNs such as cyclic di-AMP (c-di-AMP), which can be secreted by bacteria like Listeria monocytogenes3,10.
Cytosolic DNA species can trigger STING signalling following binding to a 522 amino acid protein termed cyclic GMP–AMP synthase (cGAS) (Fig. 1). In the presence of ATP and GTP, cGAS catalyses the production of a type of CDN referred to as cGAMP (cyclic GMP–AMP), which contains one 2′,5′-phosphodiester linkage and a canonical 3′,5′ linkage (c[G(2′,5′)pA(3′,5′)p])11,12,13. STING is also known to bind double-stranded DNA (dsDNA) directly14, although the physiological relevance of this remains to be clarified.
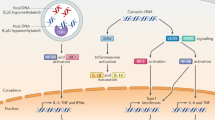
Stimulator of interferon genes (STING) is activated by cyclic dinucleotides (CDNs) produced by certain bacteria or by cyclic GMP–AMP synthase (cGAS), which in the presence of ATP and GTP catalyses the production of a type of CDN referred to as cGAMP (cyclic GMP–AMP) following binding to cytosolic DNA species (from viruses or bacteria, or self -DNA from the nucleus or mitochondria). STING is associated with the endoplasmic reticulum (ER) and, following binding to CDNs, STING forms a complex with TANK-binding kinase 1 (TBK1). This complex traffics to the perinuclear Golgi via pre-autophagosomal-like structures — a process resembling autophagy — to deliver TBK1 to endolysosomal compartments where it phosphorylates the transcription factors interferon regulatory factor 3 (IRF3) and nuclear factor-κB (NF-κB). Stimulation of the IRF3 and NF-κB signalling pathways leads to the induction of cytokines and proteins, such as the type I interferons (IFNs), that exert anti-pathogen activity. c-di-AMP, cyclic di-AMP; dsDNA, double-stranded DNA; ISGF3, interferon-stimulated gene factor 3; JAK, Janus kinase; STAT, signal transducer and activator of transcription; TYK, tyrosine kinase.
cGAS is a member of the nucleotidyltransferase family that includes the human dsRNA sensor oligoadenylate synthetase 1 (OAS1). Sequences of non-specific dsDNA species greater than 30 bp have been reported to stimulate cGAS activity, with a single CDN generated by cGAS binding to two molecules of STING in the ER15. This event probably influences changes in STING conformation, which leads to a striking trafficking event in which STING, complexed with TANK-binding kinase 1 (TBK1), relocates to perinuclear regions of the cell2,16. This process is required to deliver TBK1 to endolysosomal compartments where it phosphorylates the transcription factors interferon regulatory factor 3 (IRF3) and nuclear factor-κB (NF-κB). These transcription factors then translocate into the nucleus to initiate innate immune gene transcription (Fig. 1). STING is then rapidly degraded, an event that may avoid problems associated with sustained cytokine production17.
STING signalling appears to be independent of other DNA sensing pathways such as the Toll-like receptor 9 (TLR9) pathway, which is activated by binding to unmethylated CpG dinucleotides of approximately 21 bases long18. Thus, the TLR9 and STING signalling pathways have evolved to sense different types of DNA species, although both pathways use IRF3 and NF-κB to predominantly control gene induction. Furthermore, TLR9 is mainly expressed in pDCs and B cells, whereas STING is more broadly expressed7,18. The STING pathway is also independent of AIM2 (absent in melanoma 2), which also interacts with non-specific dsDNA species but triggers caspase 1-mediated cleavage of the pro-inflammatory cytokines interleukin-1β (IL-1β) and IL-18 from their precursor proteins19. However, STING signalling is known to potently influence the expression of the precursor proteins (but not their processing), and therefore it is possible that STING may act in concert with the AIM2 pathway20. Finally, although cGAS has been clearly implicated in regulating STING function through the generation of CDNs, other dsDNA-binding proteins have been shown to influence STING activity, such as γ-IFN-inducible protein 16 (IFI16) and DEAD box protein 41 (DDX41), although the underlying mechanisms remain to be clarified7.
Trafficking and regulation of STING. After binding to CDNs, STING (complexed with TBK1) traffics to perinuclear regions via pre-autophagosomal-like structures; a process resembling autophagy (hereafter referred to as STING-related autophagy) in which, generally under starvation conditions, cell organelles are engulfed in a membraned vesicle (that is, an autophagosome) for fusion with endosomes and lysosomes, which degrade and recycle the molecular components2,16. These pre-autophagosomal-like structures probably evolve from the ER, where STING is located1,21.
Studies indicate that STING forms part of the ER-associated translocon-associated protein (TRAP) complex, although it is not clear whether this structure is required for the autophagy-like signalling process1. Proteins intended for N-linked glycosylation and/or secretion are shunted through the TRAP complex into the luminal region of the ER following translation, although it is unknown whether STING is essential for any of these activities17,22. This form of autophagy-assisted type I IFN production has been reported to occur in pDCs that engulf DNA-containing immune complexes, an event noted to require TLR9 (Ref. 23). Thus, both TLR9- and STING-mediated signalling may similarly share an autophagy-related mechanism to induce innate immune gene activation.
Post-translational modifications of STING. Attempts to understand the mechanisms of STING signalling indicated that, in the presence of dsDNA, human STING is rapidly phosphorylated on a number of serine residues located at the carboxy-terminal section. In one study, phosphorylation of S366 was found to inhibit the ability of STING to activate IRF3, which largely prevented the induction of pro-inflammatory cytokines17. However, NF-κB signalling was not affected. This event may ensure that pro-inflammatory protein production is inhibited while ensuring that NF-κB-dependent genes continue to be expressed, perhaps to influence cell survival.
Screening assays designed to identify kinases involved in the regulation of STING by post-translational modification indicated that the autophagy-related serine/threonine protein kinases ULK1 and ULK2 (referred to here as ULK1–2) can specifically phosphorylate S366 and help prevent chronic STING activity17. TBK1 has also been suggested to target this site to control STING24. The function of ULK1–2 can be regulated by AMP-activated protein kinase (AMPK) or by mammalian target of rapamycin (mTOR), which are switched on under conditions of cellular stress25. Cytosolic dsDNA and/or CDNs were found to activate the ULK1–2 axis and to initiate a negative feedback loop that controlled STING overactivity17. Evaluation of ULK1–2-deficient mice may shed further light on the relationship of these kinases and STING function. The significance of STING phosphorylation at additional serine sites is under evaluation, but they are thought to have a role in the control of STING degradation17. Furthermore, several studies have now indicated that STING is ubiquitylated on a number of lysine residues26.
In addition, the function of STING may be controlled by microRNAs (miRNAs); a screen designed to identify miRNAs that influence host defence to viral infection identified miR-576-3p as a suppressor of STING27. Indeed, dsDNA- and CDN-triggered STING signalling was found to initiate miR-576-3p expression to prompt a feedback mechanism that helps to prevent sustained cytokine production.
While the post-translational modifications that control STING function remain to be fully determined, understanding these processes may help to explain mechanisms of innate immune signalling events and perhaps autoinflammatory disease, as well as provide potentially new therapeutic targets for drug intervention.
STING signalling triggered by microorganisms
STING promotes immunity to DNA viruses and retroviruses. Many DNA viruses have been reported to trigger STING-dependent activity, including adenovirus, vaccinia virus and papilloma virus2,28,29,30,31. The importance of the cGAS–STING axis has also been demonstrated for retrovirus infection including HIV32,33,34,35. It is probable that retroviral DNA activates the cGAS–STING axis, although a recent report indicated that retroviral RNA–DNA hybrids can also trigger this pathway suggesting that a wide array of nucleic acid structures may be capable of triggering STING activity32,33,36,37,38. CDNs may also be packaged within viruses to directly stimulate STING antiviral signalling39,40.
Recently, endogenous retroviruses (ERVs) have been reported to have a role in stimulating T cell-independent B cell activity in response to exposure to bacterial capsular polysaccharides or viral capsids41. This B cell response resulted from activation of both retinoic acid-inducible gene I (RIG-I) and STING signalling pathways by reactivated ERVs. It remains unclear how the genomes of DNA viruses, such as HIV and HSV1, trigger STING signalling, since the capsids are thought to deliver the viral genome to the nucleus through the nuclear pore33; however, it is plausible that the viral nucleic acids may be exposed from the protective capsids soon after entry into the cell or at the nuclear pore. It is also possible that some DNA viruses, such as polyomavirus, replicate in the ER where STING resides42, which may help hide the viral genome from cytosolic or nuclear sensors, such as cGAS. It thus remains to be fully determined how STING signalling is triggered through cGAS by pathogen-associated DNA in the cell.
STING suppresses replication of RNA viruses. It became evident soon after its discovery that STING is also required for host protection against a number of RNA-related pathogens, including vesicular stomatitis virus, Sendai virus and possibly dengue virus1,2,43. However, synthetic RNA, such as polyinosinic–polycytidylic acid (poly(I:C)), or RNA viruses do not trigger STING-related autophagy or the innate immune signalling events that usually result in gene induction1,2. Thus, the mechanisms by which STING thwarts gene replication of positive- and negative-sense viruses needs to be clarified. It is possible that STING exerts this effect through its translocon function, conceivably by regulating translation of viral RNA or even autophagy.
The synthase cGAS has also been shown to play a part in host defence against RNA virus infection, although the mechanisms remain to be clarified. For example, ablation of this enzyme rendered RNA virus-infected cells sensitive to viral replication, even though RNA species do not trigger cGAS activity or the generation of CDNs44,45. It is therefore possible that STING may have evolved two disparate host defence-related functions depending on whether the invading pathogen possessed an RNA or DNA genome. DNA viruses may trigger potent STING-activated gene induction, whereas RNA virus replication may be thwarted at the level of translation or post-translation. Alternatively, it is possible that STING signalling may be responsible for constitutively generating low levels of type I IFNs that are required for the robust activation of RNA-sensing pathways following viral infection. Perhaps unsurprisingly, some recent studies have reported that some viruses can suppress STING function. For example, the non-structural protein 2B (NS2B)–NS3 protease complex of dengue virus has been shown to cleave human STING, and severe acute respiratory syndrome (SARS) coronavirus reportedly inhibits the ability of STING to activate IRF3 (Ref. 46). Furthermore, adenovirus and HSV1 as well as other viruses have similarly been described to control STING function1,2,47,48,49.
The STING pathway and bacteria. Gram-negative and Gram-positive bacteria have also been reported to promote STING signalling1,2. For example, L. monocytogenes does this by secreting STING-activating CDNs (that is, c-di-AMP), although the genomes of bacteria have also been shown to be capable of stimulating STING function1,2,3,4,10. Type I IFN production has now been shown to be triggered through the STING pathway by several bacteria including Legionella pneumophila, Francisella tularensis, Chlamydia muridarum, Streptococcus pyogenes, Brucella abortus and Mycobacterium tuberculosis50,51,52,53,54,55.
Although the activation of innate immune genes through STING signalling may help prevent dangerous bacterial infection (Fig. 2), in certain situations STING-mediated innate immune gene induction surprisingly appears to facilitate bacterial survival. For example, L. monocytogenes and M. tuberculosis may activate STING to downregulate cell-mediated immunity and host defence responses55. Patients with severe tuberculosis are known to present with high IFN levels, which correspond to disease severity. Plausibly, type I IFN-induced genes may comprise cytokines that suppress adaptive immune responses, perhaps in an effort to control inflammation. Of note, mice deficient for both the TLR adaptor molecule MYD88 and STING retained the ability to mount adaptive immune responses to L. monocytogenes, suggesting that alternative sensing pathways can recognize bacterial infection and trigger adaptive immunity in the absence of STING55. This could involve activation of the RIG-I-like pathway by bacterial RNAs.
DNA viruses and cyclic dinucleotides (CDNs) produced by certain bacteria stimulate stimulator of interferon genes (STING) activity to trigger the production of primary innate immune genes. The products from these genes can exert direct intracellular anti-pathogen activity as well as trigger the production of type I interferons (IFNs) and other cytokines that can protect uninfected cells and stimulate adaptive immune responses. cGAS, cyclic GMP–AMP synthase.
These studies highlight the delicate equilibrium between an appropriate immune response and inflammation, a balance that has perhaps been exploited by microorganisms. Such findings may have important implications in the development of STING-targeting adjuvants and the design of vaccines intended to induce robust, long-lasting adaptive immune responses (Fig. 2).
STING and autoinflammatory disease
Although transient inflammatory responses are essential for initiating host defence responses to pathogen invasion, sustained or chronic inflammatory signalling is a key factor in the development of autoimmune disease. Over 20% of individuals in the United States alone will suffer some form of inflammatory disease, such as rheumatoid arthritis (RA), SLE or inflammatory bowel disease56. In almost all cases, the aetiology of such diseases remains unknown. What is known is that patients with complex inflammation-related disorders, such as RA and SLE, exhibit high levels of antinuclear antibody (ANA), DNA and even nucleosomes in the blood, as well as high levels of circulating cytokines57. Self-DNA, probably from necrotic or inappropriately apoptosed cells, has therefore been implicated in the pathogenesis of these types of disease, with aberrant innate immune signalling being caused by a failure to distinguish between self and foreign nucleic acid58.
The inability to efficiently eliminate self-DNA that could inadvertently trigger immune responses may also be a key factor in facilitating inflammatory disease. For example, a number of animal models with defects in DNases exhibit lethal inflammatory disorders. In some instances, DNase-related defects have been found in humans with comparable disease57. Defects in DNASE1, for example, have been implicated in autoimmune disorders in humans and mice59. Furthermore, mice with defects in the lysosomal DNA endonuclease Dnase2, which is essential for degrading DNA from apoptotic cells engulfed by phagocytes, die before birth owing to the toxic effects of type I IFN overproduction57. However, Dnase2−/− mice are viable when crossed with mice deficient for STING (Tmem173−/− mice), as the production of apoptotic DNA-driven cytokine production by phagocytes is almost completely eliminated20. Mice defective in three-prime repair exonuclease 1 (Trex1; also known as Dnase3) similarly exhibit significant cytokine overproduction and inflammation and, importantly, TREX1-related mutations with concomitant disease have been reported in humans60. This indicates that STING signalling is responsible for the production of harmful pro-inflammatory cytokines generated by self-DNA derived from necrotic or inappropriately apoptosed cells. Such DNases almost certainly help ensure that apoptotic cells remain 'immunologically silent' following phagocytosis.
STING-dependent signalling was also found to be the cause of inflammation-aggravated lethality in Trex1−/− mice61,62. TREX1 is a 3′ to 5′ DNA exonuclease that degrades single-stranded DNA (ssDNA) and dsDNA substrates. Trex1−/− mice are viable but have a median lifespan of approximately 10 weeks owing, in part, to the development of inflammatory myocarditis63. Patients with severe SLE and AGS have been found to have mutations in TREX1, suggesting that defects in this protein are responsible for these diseases, possibly through the inability of phagocytes to degrade self-DNA60. Such patients typically die at a young age. However, Trex1−/−Tmem173−/− double knockout mice were relatively disease free, exhibiting reduced cytokine levels, reduced ANA levels and no signs of inflammatory myocarditis or early death62. Bone marrow-derived macrophages (BMDMs) or bone marrow- derived dendritic cells (BMDCs) from Trex1−/− mice, but not from Trex1−/−Tmem173−/− mice, were found to produce high levels of inflammatory cytokines in the absence of any notable infection5,62,64. Furthermore, the adoptive transfer of wild-type bone marrow to irradiated Trex1−/− mice rescued mortality62. Thus, STING-dependent inflammatory cytokines produced by immune cells probably have an important role in TREX1-mediated inflammatory disease65 (Fig. 3).
Cyclic GMP–AMP synthase (cGAS) binds to cytosolic self-DNA and generates stimulator of interferon genes (STING)-activating cyclic dinucleotides (CDNs). The activation of STING by self-DNA can induce autoinflammatory disease. Mutations in three-prime repair exonuclease 1 (Trex1), which encodes a DNA exonuclease that degrades single-stranded DNA and double-stranded DNA substrates, can cause STING activation through failure to eliminate self-DNA that has leaked into the cytosol of the cell. Conversely, mutations in STING itself can lead to constitutive activity and autoinflammtory diseases such as STING-associated vasculopathy with onset in infancy.
However, it has been reported that Trex1−/− cells display a defect in the transition from G1 phase to S phase of the cell cycle and also accumulate DNA species in the cytoplasm63. BMDCs from Trex1−/− mice in G2 arrest exhibited increased levels of cytokine expression compared with BMDCs from wild-type mice, suggesting that TREX1 may have a housekeeping role in eliminating aberrant self-DNA species that may accrue in the cytoplasm after genome replication62,63. These nucleic acids may intrinsically activate cytokine production through activation of cGAS and STING. A further report indicated that DNA damaged by ultraviolet light and harbouring oxidative modifications was unable to be degraded by TREX1 (Ref. 66). Such DNA could trigger STING activity and may constitute a new damage-associated molecular pattern (DAMP) that can aggravate STING-dependent autoinflammatory disease.
Collectively, these studies show that the body has devised several mechanisms to prevent self-DNA from activating host defence-related DNA sensors, such as STING. TREX1 probably prevents intrinsic self-DNA-mediated activation of STING signalling, whereas DNase II may prevent STING-dependent activation by extrinsic signals in phagocytes. These DNases constitute indirect negative regulators of STING and, when defective, they can result in constitutive STING activity and the onset of inflammatory disease (Fig. 3). Understanding these processes may enable the design of new drugs for the treatment of inflammatory disease.
Mitochondrial DNA and STING
As discussed, the STING signalling pathway probably arose to detect aberrant cytosolic DNA species from an infectious agent or from leakage of self-DNA from the nucleus of damaged cells7. However, these might not be the only sources of cytosolic DNA. The hundreds, if not thousands, of mitochondria in the cytosol of most cells contain 2–10 copies of mitochondrial DNA (mtDNA), which exist as closed circular dsDNA species of approximately 15,000 base pairs67. Recent evidence indicates that mitochondrial stress, such as that induced by viruses, can cause mtDNA leakage into the cytosol that can activate the STING pathway and the production of cytokines68. Although this particular example may be of benefit to the host, the sensing of mtDNA that has leaked into the cytosol following apoptosis-induced mitochondrial stress may have detrimental effects. However, as up to 50 billion cells undergo apoptosis in the body per day, it is thus inevitable that the body has developed an effective means to enable programmed cell death events to be immunologically silent. Indeed, certain caspases may prevent the induction of inflammatory responses by cytosolic mtDNA69,70.
The process of apoptosis involves the activation of the pro-death molecules BAK and BAX, which induce mitochondrial outer-membrane permeabilization (MOMP) and causes the release of cytochrome c into the cytosol where it forms a complex with apoptotic protease-activating factor 1 (APAF1) and caspase 9, termed the apoptosome. This in turn activates caspase 3 and caspase 7, which start to target cellular proteins. Despite the fact that certain caspases have an important role in apoptosis, they are not essential for cell death and in vivo clearance of cells, suggesting that they may exert other important functions71. Indeed, recent studies have shown that BAK- and BAX-induced MOMP can also cause the release of mtDNA, the induction of cGAS–STING signalling and type I IFN production69,70. The effector caspases, caspase 3, caspase 7 and caspase 9, were shown to be required to prevent mtDNA-induced STING activation69,70. How this occurs is not yet fully apparent but may involve caspase-mediated targeting of proteins involved in STING-dependent signalling or even indirect targeting of mtDNA. Of course, such events require that STING be present in cells undergoing apoptosis and fortunately, STING is not abundantly expressed in cells such as hepatocytes, which are known to contain thousands of mitochondria7. A further conundrum is that caspase-deficient mice do not necessarily develop autoimmune disease, suggesting that caspase loss may affect other elements of the immune system to prevent disease development71.
It is interesting to note that dysfunctional mitochondria have been reported to occur in autoimmune diseases such as SLE, but whether disease pathogenesis involves release of mtDNA and activation of STING remains to be determined72. It has also been suggested that inhibitors of effector caspases could serve as antiviral therapies because they may upregulate the production of type I IFNs by released mtDNA72. In summary, the immunological silencing of programmed cell death involves an unclarified mechanism that prevents leaked mtDNA from triggering STING-dependent inflammatory processes.
STING signalling in cancer
STING-aggravated tumorigenesis. Chronic inflammatory signalling may not only instigate autoinflammatory disease such as AGS or STING-associated vasculopathy with onset in infancy (SAVI) but can also contribute to the development of cancer, probably through cytokines, chemokines and growth factors that stimulate cellular proliferation and survival, as well as by promoting angiogenesis73.
Inflammation induced by carcinogens, which cause DNA mutations, aggravates tumour development by mechanisms that remained to be determined. In one example, the polyaromatic hydrocarbon 7,12-dimethylbenz[α]anthracene (DMBA) can drive the development of cutaneous skin tumours by promoting pro-inflammatory cytokine production and stimulating phagocytic infiltration69,74,75. Indeed, mice deficient for MYD88 or for tumour necrosis factor (TNF) are relatively resistant to DMBA-induced skin carcinogenesis, indicating the importance of MYD88-dependent signalling and induction of the pro-inflammatory cytokine TNF in this process74,75.
The trigger that controls primary innate immune responses as a consequence of DNA damage, however, has remained largely unknown. Evidence indicates that TLR-independent pathways could have a key role in this process74. Furthermore, it has recently been shown that STING-deficient mice are resistant to DMBA-induced skin polyp formation76. This study showed that DMBA-initiated DNA damage lead to nucleosome leakage into the cytosol and triggered STING-dependent cytokine production by the self-DNA. It is possible that DNA-damaged keratinocytes intrinsically produce pro-inflammatory cytokines that attract phagocytes, which may facilitate the removal of the damaged cells. These infiltrating immune cells may engulf damaged keratinocytes, and the cellular DNA from the dying cells may extrinsically activate STING signalling to further propagate pro-inflammatory cytokine production74 (Fig. 4). This model is supported by the observation that the adoptive transfer of wild-type bone marrow cells to Tmem173−/− mice — which are resistant to DMBA-induced skin tumour development — results in the development of skin tumours similar to those that develop in control mice following treatment with DMBA62,76. Thus, STING signalling in haematopoietic cells, similar to MYD88 signalling, plays an important part in facilitating DMBA-induced tumorigenesis.
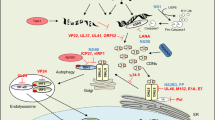
a | Stimulator of interferon genes (STING)-driven inflammation-associated cancer. Carcinogens such as 7,12-dimethylbenz[α]anthracene (DMBA) cause DNA damage, which can result in the leakage of DNA into the cytosol and the intrinsic chronic activation of the STING pathway. This event attracts phagocytes that engulf damaged cells. The self-DNA from engulfed cells can extrinsically activate STING in the phagocytes, which results in the production of more cytokines and growth factors that can activate tumour growth-promoting pathways as well as attract more immune cells to the region. b | STING and wound repair function. Carcinogens and inflammatory agents such as azoxymethane (AOM) and dextran sulfate sodium (DSS) also cause DNA damage that can result in the leakage of DNA into the cytosol and cause activation of the intrinsic STING pathway in intestinal cells. This event produces cytokines such as interleukin-1β (IL-1β) or IL-18 that enable wound repair to commence. Loss of STING prevents wound repair from occurring and may enable the infiltration and growth of genotoxic bacteria that trigger STING-independent inflammation and perhaps cancer (not shown). c | STING-dependent antitumour cytotoxic T lymphocyte (CTL) priming. Dying tumour cells are engulfed by antigen-presenting cells such as CD8α+ dendritic cells (DCs). DNA from the engulfed cell triggers STING-dependent cytokine production in the phagocyte, which facilitates cross-presentation and antitumour CTL responses. Agonists of STING have been shown to exert potent antitumour activity. CDN, cyclic dinucleotide; cGAS, cyclic GMP–AMP synthase; IFN, interferon; TCR, T cell receptor.
STING-dependent dsDNA-induced innate immune gene transcription is unaffected in Myd88−/− cells, which suggests that MYD88 activation occurs downstream of STING induction in response to DNA-damaging agents76. Thus, the current model suggests that DNA damage triggers STING-driven production of cytokines, which in an autocrine or paracrine manner can bind to receptors that signal via the MYD88 adaptor molecule, such as the IL-1 receptor and TLRs. This further drives the production of additional cytokines and growth factors that propagate inflammation and facilitate skin tumorigenesis. It is unclear whether STING has a role in other types of inflammation-aggravated cancer, such as hepatocellular carcinoma. Clearly, understanding the involvement of STING in such events may lead to the design of new anticancer therapeutics.
Antitumour effects of STING. Although MYD88 signalling facilitates carcinogenesis in the skin, MYD88 signalling is protective in models of colitis-associated carcinogenesis (CAC), which can be experimentally induced by carcinogens and inflammatory agents such as azoxymethane (AOM) and dextran sulfate sodium (DSS)74,76. It is possible that cytokines such as IL-1β and IL-18 are secreted by damaged intestinal cells to facilitate wound repair20,72,77,78. In the absence of wound repair, concomitant inflammation alters the microbial composition in the gut to species with enhanced genotoxic capacity, resulting in further inflammation, DNA damage and tumorigenesis79,73.
The phenotype of STING-deficient mice treated with AOM and DSS mimicked that of similarly treated MYD88-deficient mice; the loss of STING rendered mice susceptible to CAC80. Further investigation showed that AOM induced DNA damage and triggered STING-dependent cytokine production, including the production of IL-1β. It is thus possible that cGAS–STING recognizes early DNA damaging events to trigger the production of wound repair-initiating cytokines (Fig. 4b). STING activation in damaged intestinal cells also activates type I IFNs, which exert potent effects on the priming of antitumour T cells (see later). Thus, loss of STING may enable damaged cells to escape tumour immunosurveillance80,81,82. In summary, STING signalling may have an important protective effect against CAC, the second most common cancer in both men and women. Further analysis of the function of STING in controlling colorectal and/or intestinal wound repair, as well as its influence on the composition of the microbiota, will no doubt shed further insight into the importance of STING in influencing tumorigenesis.
STING signalling and adaptive antitumour immunity. Adaptive T cell responses are important for the control and eradication of tumour cells82. Numerous immunotherapeutic strategies involving stimulating the adaptive immune response against cancers through checkpoint blockade are presently under evaluation in clinical trials. However, how adaptive immune responses are generated by DCs against tumour cells remains unknown. What is apparent is that type I IFN production by CD8α+ DCs is involved in tumour antigen-specific T cell activation through the cross-presentation of antigen and T cell priming83. The signalling pathway responsible for type I IFN induction is thought to be independent of the TLR or RIG-I–mitochondrial antiviral signalling (MAVS) protein pathways.
Recent studies have shown that the STING pathway is essential for radiation-induced and spontaneous natural antitumour T cell responses83,84. STING-deficient mice are unable to generate efficient antitumour T cell responses and prevent melanoma tumour growth. Furthermore, the ability of checkpoint inhibitors to stimulate T cell responses was also abrogated in STING-deficient mice, indicating a role for STING in the efficacy of checkpoint inhibitors79. One hypothesis for the underlying mechanism is that CD8α+ DCs engulf necrotic tumour cells, and the tumour cell-derived DNA triggers STING signalling in the DC20,83,84,85. The resultant type I IFNs, functioning in a paracrine or autocrine manner, may induce the production of additional proteins in the DC that facilitates cross-presentation and T cell activation (Fig. 4c).
It is known that apoptotic cells do not potently activate innate immune signalling. But, engulfment of necrotic cells may trigger cytokine production, albeit at low levels, in a STING-dependent manner20. Usually, DNase II in phagocytes efficiently digests engulfed cellular DNA in the lysosomal compartment, and it is not yet clear why some host DNA from necrotic cells escape this process. It is possible that STING is activated by DNA that is present in autophagosomes or in engulfed exosomes83,84. Collectively, further understanding of the role of STING in facilitating adaptive immune responses may have notable consequences in the design of future anticancer immunotherapies.
Insight into the role of STING in facilitating antitumour T cell responses has stimulated interest in evaluating whether STING agonists could be useful therapies to treat cancer. The STING activator 5,6-dimethylxanthenone-4-acetic acid (DMXAA) was already known to exert antitumour activity but was only specific for mouse STING and failed to work in human patients86. However, several reports indicate that CDNs that bind human STING can exert antitumour activity in animal studies87. In some instances, STING agonists were shown to be effective against tumours that were resistant to programmed cell death protein 1 (PD1) blockade88. Such agonists have also been shown to be experimentally useful as adjuvants in anticancer vaccine studies89,90. Thus, the development of novel STING activators may lead to the generation and rapid expansion of immunotherapy trials to combat cancer.
Finally, it is noteworthy that anticancer drugs such as cisplatin and etoposide are also DNA adduct-forming agents that trigger cell death by instigating DNA damage91. These drugs were found to cause nuclear DNA leakage into the cytosol and to trigger intrinsic STING-dependent cytokine activity62. It is tempting to speculate that such drugs may exert their anticancer effects, in part, by stimulating the STING pathway and activating the immune system. Conversely, defects in STING signalling may contribute to chemoresistance in certain types of cancer.
The importance of STING in generating cytokines in response to DNA damage has been further emphasized in cells with defects in their DNA repair machinery92,93. Dysfunction in ataxia-telangiectasia mutated (ATM), a kinase that facilitates DNA repair, or in the co-binding protein meiotic recombination 11 homologue (MRE11) lead to the production of cytokines, which may contribute to the inflammatory phenotypes noted in patients with Ataxia telangiectasia93. Loss of DNA repair mechanisms may enable self-DNA to leak out of the cytoplasm to activate STING, or STING may be targeted by DNA repair machinery directly or indirectly to induce signalling76,93. Collectively, it is becoming apparent that STING has a key role in facilitating antitumour immune responses. Furthermore, stimulating STING activity within the tumour microenvironment may comprise a new immunotherapeutic strategy to help treat malignant disease.
Concluding remarks
Growing evidence indicates that the intrinsic STING pathway is crucial for recognizing DNA pathogens or damaged DNA in the cytosol. The innate immune response that ensues can attract immune cells that phagocytose the infected or damaged cells. The DNA from engulfed cells can stimulate extrinsic STING signalling in the phagocytes, thereby promoting further inflammatory signals.
The engulfment of apoptotic cells is immunologically silent, perhaps owing to the expression of caspases and DNases that prevent intrinsic and extrinsic STING activation. However, cancer cells are able to stimulate modest extrinsic STING activity in antigen-presenting cells such as CD8α+ DCs, and this appears to be a sufficient and essential process for the efficient priming of antitumour T cells. Understanding the underlying mechanisms of this cross-priming may lead to the development of potent antitumour vaccines and therapies94,95.
It remains to be seen what other types of inflammatory or other diseases may be caused by defects in the STING pathway. But it is possible that suppressing STING activity may help to avoid autoinflammatory disease and certain types of cancer83,96. Thus, designing drugs that trigger or repress cGAS or STING activation and/or signalling could be of interest to the anticancer, anti-pathogen, anti-inflammatory and vaccine research fields.
References
Ishikawa, H. & Barber, G. N. STING is an endoplasmic reticulum adaptor that facilitates innate immune signalling. Nature 455, 674–678 (2008). This is the first report on the discovery of STING as an essential innate immune regulator.
Ishikawa, H., Ma, Z. & Barber, G. N. STING regulates intracellular DNA-mediated, type I interferon-dependent innate immunity. Nature 461, 788–792 (2009).
Burdette, D. L. et al. STING is a direct innate immune sensor of cyclic di-GMP. Nature 478, 515–518 (2011). This report demonstrates that CDNs bind to and activate STING.
Sauer, J. D. et al. The N-ethyl-N-nitrosourea-induced Goldenticket mouse mutant reveals an essential function of Sting in the in vivo interferon response to Listeria monocytogenes and cyclic dinucleotides. Infect. Immun. 79, 688–694 (2011).
Ahn, J. & Barber, G. N. Self-DNA, STING-dependent signaling and the origins of autoinflammatory disease. Curr. Opin. Immunol. 31, 121–126 (2014).
Woo, S. R., Corrales, L. & Gajewski, T. F. The STING pathway and the T cell-inflamed tumor microenvironment. Trends Immunol. 36, 250–256 (2015).
Barber, G. N. STING-dependent cytosolic DNA sensing pathways. Trends Immunol. 35, 88–93 (2014).
Zhong, B. et al. The adaptor protein MITA links virus-sensing receptors to IRF3 transcription factor activation. Immunity 29, 538–550 (2008).
Jin, L. et al. MPYS, a novel membrane tetraspanner, is associated with major histocompatibility complex class II and mediates transduction of apoptotic signals. Mol. Cell. Biol. 28, 5014–5026 (2008).
Woodward, J. J., Iavarone, A. T. & Portnoy, D. A. c-di-AMP secreted by intracellular Listeria monocytogenes activates a host type I interferon response. Science 328, 1703–1705 (2010). This paper reports that CDNs from bacteria activate innate immune signalling.
Sun, L., Wu, J., Du, F., Chen, X. & Chen, Z. J. Cyclic GMP-AMP synthase is a cytosolic DNA sensor that activates the type I interferon pathway. Science 339, 786–791 (2013). This study shows that cGAS is a synthase that binds dsDNA to generate STING-activating CDNs.
Diner, E. J. et al. The innate immune DNA sensor cGAS produces a noncanonical cyclic dinucleotide that activates human STING. Cell Rep. 3, 1355–1361 (2013).
Ablasser, A. et al. cGAS produces a 2′-5′-linked cyclic dinucleotide second messenger that activates STING. Nature 498, 380–384 (2013).
Abe, T. et al. STING recognition of cytoplasmic DNA instigates cellular defense. Mol. Cell 50, 5–15 (2013).
Cai, X., Chiu, Y. H. & Chen, Z. J. The cGAS-cGAMP-STING pathway of cytosolic DNA sensing and signaling. Mol. Cell 54, 289–296 (2014).
Saitoh, T. et al. Atg9a controls dsDNA-driven dynamic translocation of STING and the innate immune response. Proc. Natl Acad. Sci. USA 106, 20842–20846 (2009).
Konno, H., Konno, K. & Barber, G. N. Cyclic dinucleotides trigger ULK1 (ATG1) phosphorylation of STING to prevent sustained innate immune signaling. Cell 155, 688–698 (2013).
Kumar, H., Kawai, T. & Akira, S. Pathogen recognition by the innate immune system. Int. Rev. Immunol. 30, 16–34 (2011).
Franchi, L., Munoz-Planillo, R. & Nunez, G. Sensing and reacting to microbes through the inflammasomes. Nat. Immunol. 13, 325–332 (2012).
Ahn, J., Gutman, D., Saijo, S. & Barber, G. N. STING manifests self DNA-dependent inflammatory disease. Proc. Natl Acad. Sci. USA 109, 19386–19391 (2012). This report demonstrates that STING activation in phagocytes following engulfment of dead cells is responsible for self-DNA-induced inflammatory disease.
Gomes, L. C. & Dikic, I. Autophagy in antimicrobial immunity. Mol. Cell 54, 224–233 (2014).
Nyathi, Y., Wilkinson, B. M. & Pool, M. R. Co-translational targeting and translocation of proteins to the endoplasmic reticulum. Biochim. Biophys. Acta 1833, 2392–2402 (2013).
Henault, J. et al. Noncanonical autophagy is required for type I interferon secretion in response to DNA-immune complexes. Immunity 37, 986–997 (2012).
Liu, S. et al. Phosphorylation of innate immune adaptor proteins MAVS, STING, and TRIF induces IRF3 activation. Science 347, aaa2630 (2015).
Alers, S., Loffler, A. S., Wesselborg, S. & Stork, B. Role of AMPK-mTOR-Ulk1/2 in the regulation of autophagy: cross talk, shortcuts, and feedbacks. Mol. Cell. Biol. 32, 2–11 (2012).
Liu, X., Wang, Q., Pan, Y. & Wang, C. Sensing and responding to cytosolic viruses invasions: An orchestra of kaleidoscopic ubiquitinations. Cytokine Growth Factor Rev. 26, 379–387 (2015).
Yarbrough, M. L. et al. Primate-specific miR-576-3p sets host defense signalling threshold. Nat. Commun. 5, 4963 (2014).
Lam, E., Stein, S. & Falck-Pedersen, E. Adenovirus detection by the cGAS/STING/TBK1 DNA sensing cascade. J. Virol. 88, 974–981 (2014).
Sunthamala, N. et al. E2 proteins of high risk human papillomaviruses down-modulate STING and IFN-κ transcription in keratinocytes. PLoS ONE 9, e91473 (2014).
Ablasser, A. et al. Cell intrinsic immunity spreads to bystander cells via the intercellular transfer of cGAMP. Nature 503, 530–534 (2013).
Lau, L., Gray, E. E., Brunette, R. L. & Stetson, D. B. DNA tumor virus oncogenes antagonize the cGAS-STING DNA sensing pathway. Science 350, 568–571 (2015).
Gao, D. et al. Cyclic GMP-AMP synthase is an innate immune sensor of HIV and other retroviruses. Science 341, 903–906 (2013).
Lahaye, X. et al. The capsids of HIV-1 and HIV-2 determine immune detection of the viral cDNA by the innate sensor cGAS in dendritic cells. Immunity 39, 1132–1142 (2013).
Herzner, A. M. et al. Sequence-specific activation of the DNA sensor cGAS by Y-form DNA structures as found in primary HIV-1 cDNA. Nat. Immunol. 16, 1025–1033 (2015).
Yoh, S. M. et al. PQBP1 is a proximal sensor of the cGAS-dependent innate response to HIV-1. Cell 161, 1293–1305 (2015).
Mankan, A. K. et al. Cytosolic RNA:DNA hybrids activate the cGAS-STING axis. EMBO J. 33, 2937–2946 (2014).
Yan, N., Regalado-Magdos, A. D., Stiggelbout, B., Lee-Kirsch, M. A. & Lieberman, J. The cytosolic exonuclease TREX1 inhibits the innate immune response to human immunodeficiency virus type 1. Nat. Immunol. 11, 1005–1013 (2010).
Booiman, T., Setiawan, L. C. & Kootstra, N. A. Genetic variation in Trex1 affects HIV-1 disease progression. AIDS 28, 2517–2521 (2014).
Bridgeman, A. et al. Viruses transfer the antiviral second messenger cGAMP between cells. Science 349, 1228–1232 (2015).
Gentili, M. et al. Transmission of innate immune signaling by packaging of cGAMP in viral particles. Science 349, 1232–1236 (2015).
Zeng, M. et al. MAVS, cGAS, and endogenous retroviruses in T-independent B cell responses. Science 346, 1486–1492 (2014).
Inoue, T. & Tsai, B. How viruses use the endoplasmic reticulum for entry, replication, and assembly. Cold Spring Harb. Perspect. Biol. 5, a013250 (2013).
Aguirre, S. et al. DENV inhibits type I IFN production in infected cells by cleaving human STING. PLoS Pathog. 8, e1002934 (2012).
Li, X. D. et al. Pivotal roles of cGAS-cGAMP signaling in antiviral defense and immune adjuvant effects. Science 341, 1390–1394 (2013).
Schoggins, J. W. et al. Pan-viral specificity of IFN-induced genes reveals new roles for cGAS in innate immunity. Nature 505, 691–695 (2014).
Maringer, K. & Fernandez-Sesma, A. Message in a bottle: lessons learned from antagonism of STING signalling during RNA virus infection. Cytokine Growth Factor Rev. 25, 669–679 (2014).
Kalamvoki, M. & Roizman, B. HSV-1 degrades, stabilizes, requires, or is stung by STING depending on ICP0, the US3 protein kinase, and cell derivation. Proc. Natl Acad. Sci. USA 111, E611–E617 (2014).
Zhou, Q. et al. The ER-associated protein ZDHHC1 is a positive regulator of DNA virus-triggered, MITA/STING-dependent innate immune signaling. Cell Host Microbe 16, 450–461 (2014).
Luecke, S. & Paludan, S. R. Innate recognition of alphaherpesvirus DNA. Adv. Virus Res. 92, 63–100 (2015).
Mitzel, D. N., Lowry, V., Shirali, A. C., Liu, Y. & Stout-Delgado, H. W. Age-enhanced endoplasmic reticulum stress contributes to increased Atg9A inhibition of STING-mediated IFN-β production during Streptococcus pneumoniae infection. J. Immunol. 192, 4273–4283 (2014).
Prantner, D., Darville, T. & Nagarajan, U. M. Stimulator of IFN gene is critical for induction of IFN-β during Chlamydia muridarum infection. J. Immunol. 184, 2551–2560 (2010).
Storek, K. M., Gertsvolf, N. A., Ohlson, M. B. & Monack, D. M. cGAS and Ifi204 cooperate to produce type I IFNs in response to Francisella infection. J. Immunol. 194, 3236–3245 (2015).
Watson, R. O., Manzanillo, P. S. & Cox, J. S. Extracellular M. tuberculosis DNA targets bacteria for autophagy by activating the host DNA-sensing pathway. Cell 150, 803–815 (2012).
Dey, B. et al. A bacterial cyclic dinucleotide activates the cytosolic surveillance pathway and mediates innate resistance to tuberculosis. Nat. Med. 21, 401–406 (2015).
Archer, K. A., Durack, J. & Portnoy, D. A. STING-dependent type I IFN production inhibits cell-mediated immunity to Listeria monocytogenes. PLoS Pathog. 10, e1003861 (2014).
Centers for Disease Control and Prevention. Prevalence of doctor-diagnosed arthritis and arthritis-attributable activity limitation—United States, 2010–2012. MMWR Morb. Mortal. Wkly Rep. 62, 869–873 (2013).
Nagata, S. & Kawane, K. Autoinflammation by endogenous DNA. Adv. Immunol. 110, 139–161 (2011).
Janko, C. et al. Inflammatory clearance of apoptotic remnants in systemic lupus erythematosus (SLE). Autoimmun. Rev. 8, 9–12 (2008).
Martinez Valle, F., Balada, E., Ordi-Ros, J. & Vilardell-Tarres, M. DNase 1 and systemic lupus erythematosus. Autoimmun Rev. 7, 359–363 (2008).
Rice, G. I., Rodero, M. P. & Crow, Y. J. Human disease phenotypes associated with mutations in TREX1. J. Clin. Immunol. 35, 235–243 (2015).
Gall, A. et al. Autoimmunity initiates in nonhematopoietic cells and progresses via lymphocytes in an interferon-dependent autoimmune disease. Immunity 36, 120–131 (2012). This study shows that STING activation is responsible for TREX1-mediated autoinflamatory disease and plausibly AGS.
Ahn, J., Ruiz, P. & Barber, G. N. Intrinsic self-DNA triggers inflammatory disease dependent on STING. J. Immunol. 193, 4634–4642 (2014). This report demonstrates that, in the absence of TREX1, self-DNA in macrophages can activate STING and cause inflammatory diseases, such as AGS.
Yang, Y. G., Lindahl, T. & Barnes, D. E. Trex1 exonuclease degrades ssDNA to prevent chronic checkpoint activation and autoimmune disease. Cell 131, 873–886 (2007).
Pereira-Lopes, S. et al. The exonuclease Trex1 restrains macrophage proinflammatory activation. J. Immunol. 191, 6128–6135 (2013).
Ablasser, A. et al. TREX1 deficiency triggers cell-autonomous immunity in a cGAS-dependent manner. J. Immunol. 192, 5993–5997 (2014). This study reports that cGAS is involved in STING-dependent autoinflammatory disease that is triggered by TREX1 deficiency.
Gehrke, N. et al. Oxidative damage of DNA confers resistance to cytosolic nuclease TREX1 degradation and potentiates STING-dependent immune sensing. Immunity 39, 482–495 (2013).
Kukat, C. & Larsson, N. G. mtDNA makes a U-turn for the mitochondrial nucleoid. Trends Cell Biol. 23, 457–463 (2013).
West, A. P. et al. Mitochondrial DNA stress primes the antiviral innate immune response. Nature 520, 553–557 (2015).
White, M. J. et al. Apoptotic caspases suppress mtDNA-induced STING-mediated type I IFN production. Cell 159, 1549–1562 (2014).
Rongvaux, A. et al. Apoptotic caspases prevent the induction of type I interferons by mitochondrial DNA. Cell 159, 1563–1577 (2014). References 69 and 70 show that mitochondrial DNA that has leaked into the cytosol may activate STING-dependent inflammatory responses.
Creagh, E. M. Caspase crosstalk: integration of apoptotic and innate immune signalling pathways. Trends Immunol. 35, 631–640 (2014).
Perl, A., Hanczko, R. & Doherty, E. Assessment of mitochondrial dysfunction in lymphocytes of patients with systemic lupus erythematosus. Methods Mol. Biol. 900, 61–89 (2012).
Nowarski, R., Gagliani, N., Huber, S. & Flavell, R. A. Innate immune cells in inflammation and cancer. Cancer Immunol. Res. 1, 77–84 (2013).
Salcedo, R., Cataisson, C., Hasan, U., Yuspa, S. H. & Trinchieri, G. MyD88 and its divergent toll in carcinogenesis. Trends Immunol. 34, 379–389 (2013).
Swann, J. B. et al. Demonstration of inflammation-induced cancer and cancer immunoediting during primary tumorigenesis. Proc. Natl Acad. Sci. USA 105, 652–656 (2008).
Ahn, J. et al. Inflammation-driven carcinogenesis is mediated through STING. Nat. Commun. 5, 5166 (2014). This study shows that DNA-damaging agents can cause DNA leakage in the cytosol to activate STING-mediated inflammatory events and drive skin cancer.
Salcedo, R. et al. MyD88-mediated signaling prevents development of adenocarcinomas of the colon: role of interleukin 18. J. Exp. Med. 207, 1625–1636 (2010).
Huber, S. et al. IL-22BP is regulated by the inflammasome and modulates tumorigenesis in the intestine. Nature 491, 259–263 (2012).
Irrazabal, T., Belcheva, A., Girardin, S. E., Martin, A. & Philpott, D. J. The multifaceted role of the intestinal microbiota in colon cancer. Mol. Cell 54, 309–320 (2014).
Ahn, J., Konno, H. & Barber, G. N. Diverse roles of STING-dependent signaling on the development of cancer. Oncogene 34, 5302–5308 (2015).
Zhu, Q. et al. Cutting edge: STING mediates protection against colorectal tumorigenesis by governing the magnitude of intestinal inflammation. J. Immunol. 193, 4779–4782 (2014). References 80 and 81 show a protective role for STING in recognizing DNA damage and facilitating wound repair in the colon.
Gajewski, T. F., Schreiber, H. & Fu, Y. X. Innate and adaptive immune cells in the tumor microenvironment. Nat. Immunol. 14, 1014–1022 (2013).
Woo, S. R. et al. STING-dependent cytosolic DNA sensing mediates innate immune recognition of immunogenic tumors. Immunity 41, 830–842 (2014). This study shows that antitumour T cell responses require the activation of STING in phagocytes that have engulfed tumour cells.
Deng, L. et al. STING-dependent cytosolic DNA sensing promotes radiation-induced type I interferon-dependent antitumor immunity in immunogenic tumors. Immunity 41, 843–852 (2014). This study shows that radiation-induced tumour cell death stimulates STING-dependent antitumour T cell responses.
Klarquist, J. et al. STING-mediated DNA sensing promotes antitumor and autoimmune responses to dying cells. J. Immunol. 193, 6124–6134 (2014).
Roberts, Z. J. et al. The chemotherapeutic agent DMXAA potently and specifically activates the TBK1-IRF-3 signaling axis. J. Exp. Med. 204, 1559–1569 (2007).
Corrales, L. et al. Direct activation of STING in the tumor microenvironment leads to potent and systemic tumor regression and immunity. Cell Rep. 11, 1018–1030 (2015). This study demonstrates that the use of STING agonists can exert potent antitumour therapeutic effects.
Fu, J. et al. STING agonist formulated cancer vaccines can cure established tumors resistant to PD-1 blockade. Sci. Transl Med. 7, 283ra52 (2015).
Wang, Z. & Celis, E. STING activator c-di-GMP enhances the anti-tumor effects of peptide vaccines in melanoma-bearing mice. Cancer Immunol. Immunother. 64, 1057–1066 (2015).
Ohkuri, T., Ghosh, A., Kosaka, A., Sarkar, S. N. & Okada, H. Protective role of STING against gliomagenesis: rational use of STING agonist in anti-glioma immunotherapy. Oncoimmunology 4, e999523 (2015).
Zitvogel, L., Galluzzi, L., Smyth, M. J. & Kroemer, G. Mechanism of action of conventional and targeted anticancer therapies: reinstating immunosurveillance. Immunity 39, 74–88 (2013).
Hartlova, A. et al. DNA damage primes the type I interferon system via the cytosolic DNA sensor STING to promote anti-microbial innate immunity. Immunity 42, 332–343 (2015).
Kondo, T. et al. DNA damage sensor MRE11 recognizes cytosolic double-stranded DNA and induces type I interferon by regulating STING trafficking. Proc. Natl Acad. Sci. USA 110, 2969–2974 (2013).
Lemos, H., Huang, L., McGaha, T. & Mellor, A. L. STING, nanoparticles, autoimmune disease and cancer: a novel paradigm for immunotherapy? Expert Rev. Clin. Immunol. 11, 155–165 (2015).
Zhang, H. et al. Cell-free tumor microparticle vaccines stimulate dendritic cells via cGAS/STING signaling. Cancer Immunol. Res. 3, 196–205 (2015).
Dubensky, T. W. Jr., Kanne, D. B. & Leong, M. L. Rationale, progress and development of vaccines utilizing STING-activating cyclic dinucleotide adjuvants. Ther. Adv. Vaccines 1, 131–143 (2013).
Gao, P. et al. Structure-function analysis of STING activation by c[G(2′,5′)pA(3′,5′)p] and targeting by antiviral DMXAA. Cell 154, 748–762 (2013).
Jin, L. et al. Identification and characterization of a loss-of-function human MPYS variant. Genes Immun. 12, 263–269 (2011).
Prantner, D. et al. 5,6-Dimethylxanthenone-4-acetic acid (DMXAA) activates stimulator of interferon gene (STING)-dependent innate immune pathways and is regulated by mitochondrial membrane potential. J. Biol. Chem. 287, 39776–39788 (2012).
Liu, Y. et al. Activated STING in a vascular and pulmonary syndrome. N. Engl. J. Med. 371, 507–518 (2014). This paper reports that mutations in TMEM173 that cause constitutive activity of STING can induce inflammatory disease.
Jeremiah, N. et al. Inherited STING-activating mutation underlies a familial inflammatory syndrome with lupus-like manifestations. J. Clin. Invest. 124, 5516–5520 (2014).
Lemos, H. et al. Activation of the STING adaptor attenuates experimental autoimmune encephalitis. J. Immunol. 192, 5571–5578 (2014).
Wu, X. et al. Molecular evolutionary and structural analysis of the cytosolic DNA sensor cGAS and STING. Nucleic Acids Res. 42, 8243–8257 (2014).
Mozzi, A. et al. OASes and STING: adaptive evolution in concert. Genome Biol. Evol. 7, 1016–1032 (2015).
Yi, G. et al. Single nucleotide polymorphisms of human STING can affect innate immune response to cyclic dinucleotides. PLoS ONE 8, e77846 (2013).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The author declares no competing financial interests.
Glossary
- Stimulator of interferon genes
-
(STING). An endoplasmic reticulum-associated protein that binds to cyclic dinucleotides or cytosolic DNA to trigger cytokine production.
- Cyclic dinucleotides
-
(CDNs). Secondary messengers secreted by certain bacteria, such as Listeria monocytogenes, or generated by cyclic GMP–AMP synthase (cGAS) following binding to cytosolic DNA.
- Systemic lupus erythematosus
-
(SLE). An autoimmune disease in which autoantibodies specific for DNA, RNA or proteins associated with nucleic acids form immune complexes that damage small blood vessels, especially in the kidneys. Patients with SLE generally have abnormal B and T cell function. SLE can be associated with hyperproduction of type I interferons.
- Aicardi–Goutières syndrome
-
(AGS). A neurodegenerative disorder that can be caused by STING-dependent cytokine hyperproduction owing to mutations in genes such as TREX1 (three-prime repair exonuclease 1).
- Cyclic GMP–AMP synthase
-
(cGAS). A cellular protein that binds to cytosolic DNA species and generates stimulator of interferon genes (STING)-activating cyclic dinucleotides.
- Oligoadenylate synthetase 1
-
(OAS1). A double-stranded RNA sensor that, in the presence of ATP, generates 2′,5′-linked oligoadenylates, which can activate RNase L-mediated viral RNA degradation.
- Translocon-associated protein
-
(TRAP). A complex of proteins that shunt proteins destined for N-linked glycosylation and secretion into the endoplasmic reticulum after translation.
- Immune complexes
-
Complexes of antigen (including DNA) bound to antibody and, sometimes, components of the complement system. The levels of immune complexes are increased in many autoimmune disorders, in which they become deposited in tissues and cause tissue damage.
- MicroRNAs
-
(miRNAs). Small RNA molecules that regulate the expression of genes by binding to the 3′ untranslated regions (3′ UTR) of specific mRNAs.
- Three-prime repair exonuclease 1
-
(TREX1). A DNase that degrades self-DNA species that can otherwise chronically activate stimulator of interferon genes (STING) signalling.
- Damage-associated molecule pattern
-
(DAMP). A cellular molecule, such as uric acid, ATP and heat shock proteins, that can activate the inflammatory response in the absence of infection.
- Mitochondrial outer-membrane permeabilization
-
(MOMP). Permeablization induced by virus infection and/or apoptosis to release mitochondrial DNA that could conceivably activate stimulator of interferon genes (STING).
- Angiogenesis
-
The development of new blood vessels from existing blood vessels. It is frequently associated with tumour development and inflammation.
- Checkpoint blockade
-
The targeting of T cell co-inhibitory receptors, such as cytotoxic T lymphocyte antigen 4 (CTLA4) or programmed cell death protein 1 (PD1), by antibodies that have clinical activity in multiple cancer types.
- Cross-presentation
-
The initiation of a CD8+ T cell response to an antigen that is not present within antigen-presenting cells (APCs). This exogenous antigen must be taken up by APCs and then re-routed to the MHC class I pathway of antigen presentation.
- Ataxia telangiectasia
-
A neurodegenerative disease caused by mutations in the ATM (ataxia-telangiectasia mutated) gene, the product of which is essential for the recognition and repair of DNA breaks.
Rights and permissions
About this article
Cite this article
Barber, G. STING: infection, inflammation and cancer. Nat Rev Immunol 15, 760–770 (2015). https://doi.org/10.1038/nri3921
Published:
Issue Date:
DOI: https://doi.org/10.1038/nri3921
This article is cited by
-
cGAS-STING, inflammasomes and pyroptosis: an overview of crosstalk mechanism of activation and regulation
Cell Communication and Signaling (2024)
-
Cystoid Macular Oedema in a Patient Treated with STING Agonist and Ezabenlimab for Disseminated Melanoma
Ophthalmology and Therapy (2024)
-
Regulatory cells and the effect of cancer immunotherapy
Molecular Cancer (2023)
-
Targeting cGAS/STING signaling-mediated myeloid immune cell dysfunction in TIME
Journal of Biomedical Science (2023)
-
A systems biology approach to pathogenesis of gastric cancer: gene network modeling and pathway analysis
BMC Gastroenterology (2023)