Key Points
-
The kinase mTOR (mammalian target of rapamycin) is part of an evolutionarily conserved pathway that couples cell growth and metabolism to inputs from the environment in all eukaryotes. mTOR exists in two multiprotein complexes in metazoans, mTOR complex 1 (mTORC1) and mTORC2, which have distinct molecular compositions, cellular actions and physiological functions.
-
T cells depend on mTOR signalling to sense and integrate immune signals from dendritic cells (including antigenic signals, co-stimulatory molecules and cytokines), environmental cues derived from growth factors and immunoregulatory factors, and nutrients.
-
Under steady-state conditions, mTOR activity is tightly controlled by multiple inhibitory molecules — in particular the tumour suppressors PTEN (phosphatase and tensin homologue), TSC1 (tuberous sclerosis 1) and LKB1 (liver kinase B1) — to enforce the normal homeostasis of T cells. Loss of these control mechanisms disrupts the development and maintenance of T cells.
-
Antigen recognition triggers robust mTOR activation, which drives the differentiation of naive CD4+ T cells into the T helper 1 (TH1), TH2 and TH17 cell effector lineages, while inhibiting the induction of regulatory T cells and T cell anergy. mTOR also promotes an effector fate over a memory fate for CD8+ T cells in infection and tumour immunity.
-
mTOR serves as a signalling node to activate several downstream effector pathways, including immune receptor signalling, metabolic programmes and migratory activity. The coordination of these pathways by mTOR is important to achieve a productive immune response.
-
Inhibition of mTOR induces immunosuppression in transplant rejections and autoimmune disorders, but also represents a promising strategy for vaccine development to boost immunity against pathogens and tumours. More potent and selective inhibitors of mTOR and mTOR-associated pathways are being developed for clinical and research purposes.
Abstract
The evolutionarily conserved kinase mTOR (mammalian target of rapamycin) couples cell growth and metabolism to environmental inputs in eukaryotes. T cells depend on mTOR signalling to integrate immune signals and metabolic cues for their proper maintenance and activation. Under steady-state conditions, mTOR is actively controlled by multiple inhibitory mechanisms, and this enforces normal T cell homeostasis. Antigen recognition by naive CD4+ and CD8+ T cells triggers mTOR activation, which in turn programmes the differentiation of these cells into functionally distinct lineages. This Review focuses on the signalling mechanisms of mTOR in T cell homeostatic and functional fates, and discusses the therapeutic implications of targeting mTOR in T cells.
Similar content being viewed by others
Main
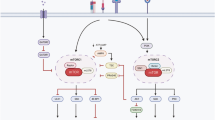
The mammalian target of rapamycin (mTOR; now officially known as the mechanistic target of rapamycin) is a conserved serine/threonine kinase that has a central role in the regulation of cell growth and metabolism1. mTOR senses and integrates diverse environmental signals, including nutrients and growth factors, many of which deliver their inputs to the phosphoinositide 3-kinase (PI3K)–AKT pathway that ultimately activates mTOR. mTOR exists in two multiprotein complexes in metazoans. mTOR complex 1 (mTORC1) contains a scaffolding protein, regulatory associated protein of mTOR (RAPTOR), and is sensitive to the immunosuppressant rapamycin (Fig. 1a). mTORC2 has a distinct scaffolding protein, rapamycin-insensitive companion of mTOR (RICTOR), and is relatively resistant to rapamycin except under prolonged periods of treatment. Aberrantly elevated mTOR activity is frequently associated with human malignancies, and consequently mTOR has been the subject of extensive investigation by cancer biologists1. However, emerging evidence highlights a crucial role of mTOR signalling in both the innate and adaptive immune systems.
a | In T cells, mammalian target of rapamycin (mTOR) can be activated by multiple signals. These include the conventionally defined immune signals 1, 2 and 3 (antigenic stimulation, co-stimulation and cytokines); growth factors and immunomodulatory factors (such as leptin and sphingosine-1-phosphate (S1P)); and nutrients. The tuberous sclerosis 1 (TSC1)–TSC2 complex integrates signals from the phosphoinositide 3-kinase (PI3K)–AKT and liver kinase B1 (LKB1)–AMP-activated protein kinase (AMPK) pathways. These pathways reciprocally regulate TSC2 activity through AKT-dependent phosphorylation of threonine 1462 (Thr1462) and AMPK-dependent phosphorylation of serine 1387 (Ser1387). Following antigenic stimulation, the TSC complex is inactivated by T cell receptor (TCR) signals, resulting in the activation of mTOR complex 1 (mTORC1), whereas TSC function is maintained in naive T cells to keep mTORC1 activation in check. In addition, AKT and AMPK can directly modulate mTORC1 functions independently of TSC and the small GTPase RHEB, and amino acids can activate mTORC1 via the RAG family of small GTPases. mTORC1 is best known for its function in promoting translation initiation and protein synthesis by directly phosphorylating ribosomal protein S6 kinases (S6Ks) and eIF4E-binding proteins (4E-BPs). Additional mTORC1 targets include regulatory proteins with roles in cell signalling, metabolism and autophagy. mTORC2 is important for the full activation of AKT through the phosphorylation of Ser473, and thus AKT can be both upstream of mTORC1 and downstream of mTORC2. Moreover, mTORC2 phosphorylates various protein kinase C (PKC) isoforms — including PKCθ, which activates nuclear factor-κB (NF-κB) in T cells — as well as serum- and glucocorticoid-regulated kinase 1 (SGK1). b | Under steady-state conditions, molecules that inhibit mTOR actively maintain the homeostasis of T cells in the thymus and periphery by preventing the cells from engaging alternative fates. Although phosphatase and tensin homologue (PTEN), TSC1 and LKB1 have a shared capacity to inactivate mTOR, they have distinct effects in enforcing T cell homeostasis. c | Following antigenic stimulation, mTOR signalling promotes the differentiation of T helper 1 (TH1), TH2 and TH17 cells, and inhibits the generation of induced regulatory T (iTReg) cells. AICAR, 5-aminoimidazole-4-carboxamide riboside; APC, antigen-presenting cell; CAMKK2, calcium/calmodulin-dependent protein kinase kinase 2; FOXO, forkhead box O; IL, interleukin; PDK1, 3-phosphoinositide-dependent protein kinase 1; PtdIns(4,5)P2, phosphatidylinositol-4,5-bisphosphate; PtdIns(3,4,5)P3, phosphatidylinositol-3,4,5-trisphosphate; STAT, signal transducer and activator of transcription; TGFβ, transforming growth factor-β.
The outcome of an adaptive immune response depends on the sensing of antigenic and inflammatory signals by T cells. T cells have evolved to perceive these immune stimuli and further coordinate them with diverse environmental and metabolic cues through the evolutionarily conserved mTOR pathway. Thus, mTOR endows T cells with the ability to properly integrate a multitude of signals to determine the outcome of adaptive immunity. The first major function ascribed to mTOR in T cells was the promotion of cell cycle progression, but more recent studies have established mTOR signalling as a fundamental determinant of cell fate decisions both under steady-state conditions and following cognate antigen recognition. It is likely that mTOR affects these diverse processes in T cells by serving as a signalling node to coordinately regulate immune receptor signalling pathways, metabolic programmes and migratory activity. As several excellent reviews have covered the immune functions of mTOR2,3,4, this Review mainly focuses on the most recent genetic studies that have identified new roles for mTOR signalling in T cell fate decisions and on the therapeutic implications of modulating mTOR functions in T cells. Following the hierarchy of signal transduction, I first discuss the extracellular inputs that feed into the mTOR pathway in T cells. Then I describe the mechanisms through which negative and positive components of mTOR signalling impinge on cell fate decisions in T cells, with a special focus on T cell homeostasis and T helper (TH) cell differentiation (Table 1). Finally, I discuss mTOR downstream effector pathways that are involved in T cell metabolism, as well as the implications of targeting mTOR and metabolic pathways for disease therapeutics.
mTOR activating signals
In T cells, mTOR senses three broad categories of instructive signals. The first category comprises the immune activation signals transduced from dendritic cells (DCs) through antigens, co-stimulatory molecules and pro-inflammatory cytokines (known as signals 1, 2 and 3); these signals are essential for directing proper T cell activation and differentiation5. The other two types of instructive signals are mediated by environmental stimuli, such as growth factors and immunomodulatory factors, and by metabolic cues, which are derived mainly from nutrients (Fig. 1a). Whereas the immune activation signals are unique to T cells, the environmental and metabolic stimuli act on all eukaryotic cells. Many of the upstream signals activate mTORC1 through the GTP-loaded form of the small GTPase RHEB (Fig. 1a). RHEB is tightly regulated by the tuberous sclerosis 1 (TSC1)–TSC2 complex, which, through its GTPase-activating protein (GAP) activity towards RHEB, inactivates RHEB and mTORC1. The TSC complex serves as a molecular switch to integrate upstream signals and, in particular, the positive and negative signals transduced from the PI3K–AKT and AMP-activated protein kinase (AMPK) pathways, respectively1. RHEB deficiency in T cells reduces mTORC1 activation in response to T cell receptor (TCR) stimulation6, whereas loss of TSC1 disrupts the entire TSC complex and enhances basal and TCR-induced mTORC1 activity7,8,9. These results highlight a crucial role for the TSC–RHEB axis in T cell responses.
TSC-independent pathways also engage mTORC1, although the importance of these mechanisms in T cells remains to be determined. Activation of mTORC1 in response to amino acids requires the RAG family of GTPases10,11, whereas direct phosphorylation of RAPTOR by AMPK inhibits mTORC1 during energy stress12. Furthermore, the mitogen-activated protein kinase (MAPK) p38β targets different components of mTORC1 to either positively or negatively regulate the activity of this complex under different environmental stresses13,14. Upstream pathways that activate mTORC2 are just beginning to be identified, with recent studies showing a role for ribosomes in linking PI3K to mTORC2 activation15,16. Moreover, another small GTPase, RAC1, binds directly to mTOR to activate mTORC1 and mTORC2, which provides a means to regulate both mTOR complexes simultaneously17. The convergence of multiple signals on mTOR suggests that its basic function is as a signal integrator.
Immune activation signals. Both mTORC1 and mTORC2 are activated within minutes of TCR stimulation. The magnitude of mTOR activation is directly correlated with the duration of the T cell–DC interaction and the dose of the cognate antigen18,19. The activity of mTOR is further shaped by co-stimulatory signals. CD28-mediated co-stimulation is a classic activating signal for the PI3K–AKT pathway, which in turn upregulates the mTOR activity induced by the TCR to facilitate productive T cell activation20,21,22. Another co-stimulatory receptor, OX40, which is a member of the tumour necrosis factor receptor (TNFR) family, assembles a signalling complex by recruiting PI3K–AKT to augment TCR-dependent AKT signalling23. By contrast, the PDL1 (PD1 ligand 1)–PD1 (programmed cell death protein 1) axis, a negative co-stimulatory pathway in T cells, downregulates mTOR signalling to mediate immune tolerance24.
Compared with the rapid activation of mTOR by the TCR, the homeostatic cytokine interleukin-7 (IL-7) induces delayed yet sustained PI3K–AKT signalling and mTOR activation. This activation depends on the transcriptional activity of signal transducer and activator of transcription 5 (STAT5) and contributes to IL-7-mediated glucose uptake and trophic effects in T cells25,26. However, increased mTORC1 activity as a result of TSC1 deficiency impairs IL-7-dependent survival responses in naive T cells7, suggesting that a defined level of mTOR activity is important to mediate the IL-7 response. In antigen-stimulated CD8+ T cells, IL-12 enhances and prolongs TCR-dependent mTOR activation to programme the functional maturation of effector T cells. Similarly to IL-7-mediated stimulation of naive T cells, IL-12-induced mTOR activation in antigen-stimulated CD8+ T cells is indirect and depends on STAT signalling (in this case, through STAT4)27. In TH2 and TH17 cells, mTOR is activated by IL-4 and IL-1, respectively, to facilitate cell cycle progression28,29. Furthermore, there is extensive interplay between mTOR and STAT signalling in T cells6,30 and other cells31.
The leptin receptor. Leptin — an adipose tissue-derived hormone that regulates energy metabolism — has long been known to directly regulate T cell proliferation and cytokine production, thereby linking nutritional status and pro-inflammatory immune responses32. More recently, regulatory T cells (TReg cells) were shown to express leptin and its receptor and to have greater mTOR activity than conventional T cells33. Neutralization of leptin or deletion of the gene encoding the leptin receptor considerably diminishes mTOR activity in TReg cells, which suggests a link between autocrine secretion of leptin and mTOR activation in TReg cells. The mTOR activity that results from leptin signalling maintains the anergic state of TReg cells, because transient inhibition of mTOR or neutralization of leptin reverses the hyporesponsiveness of TReg cells to TCR stimulation, resulting in their robust proliferation33,34. Leptin also activates mTOR in autoreactive CD4+ T cells to promote their survival and mediate autoimmune neuroinflammation35. As the leptin level is directly correlated with nutrient status, the leptin–mTOR axis has been proposed to bridge metabolism and immunity33.
S1PR1. Sphingosine-1-phosphate receptor 1 (S1PR1) — a G protein-coupled receptor for the bioactive lipid sphingosine-1-phosphate (S1P) — is a crucial regulator of T cell egress from the thymus and secondary lymphoid organs. In TReg cells, S1PR1 activates AKT and mTOR, and this delivers a cell-intrinsic negative signal to restrain TReg cell suppressive activity36. In conventional CD4+ T cells, S1PR1 is dispensable for immediate mTOR activation but is important to sustain mTOR activity during the differentiation of these cells into effector T cells37. S1PR1-dependent activation of mTOR inhibits the generation of TReg cells while driving TH1 cell development in a reciprocal manner. These studies identify an S1PR1–mTOR axis that controls TReg cell function and T cell lineage choices36,37.
TLRs. When activated by their ligands, Toll-like receptors (TLRs) expressed by innate immune cells induce the release of large amounts of pro-inflammatory molecules that are important for immediate immune defence. In addition, both CD4+ and CD8+ T cells express functional TLRs, and recent results suggest a T cell-intrinsic role for TLRs in immune responses. TLR2 signals through the adaptor molecule myeloid differentiation primary-response protein 88 (MYD88) to activate mTOR and promote the expression of T-bet in effector CD8+ T cells38. As mTOR is also downstream of TCR signalling, the activation of mTOR bridges TCR and TLR signals and thereby promotes effector CD8+ T cell function38 and contributes to the generation of T cell memory39. However, MYD88 deficiency in T cells diminishes initial T cell clonal expansion, but not the subsequent generation of the memory population40. Further study is required to clarify how TLR2–MYD88 signalling affects mTOR activation in T cells.
mTOR is also activated by additional extracellular signals in T cells. Insulin induces the activation of mTORC1 and promotes mTOR-dependent expression of T-bet in antigen-activated CD8+ T cells27. Also, Notch 1 activates the AKT–mTOR pathway during early thymocyte development and relies on AKT-dependent metabolic and trophic effects to regulate thymocyte differentiation and survival41. Finally, mTOR is intimately linked to nutrient sensing in all eukaryotic cells1, and the physiological relevance of the metabolic regulation of mTOR is discussed in further detail below.
mTOR in immune homeostasis
T cells develop in the thymus through a stepwise differentiation process. Once mature T cells are released into the periphery, they circulate through the blood and peripheral lymphoid organs in a quiescent state that is characterized by small cell size and low metabolic activity. The survival of naive T cells relies on the engagement of TCRs by self-peptide–MHC complexes and on the availability of IL-7, and is further shaped by growth factors and nutrients for metabolic fitness42,43. Given the central role of mTOR as an environmental sensor, it might be predicted that mTOR activity is required for the homeostasis of thymocytes and peripheral T cells. Surprisingly, deletion of the Mtor gene after initial thymocyte development (using the Cd4-Cre system) had minimal effects on the homeostasis of peripheral T cells under steady-state conditions6, although the requirement for mTOR in early thymic development has yet to be determined. By contrast, several negative regulators of mTOR signalling were found to enforce normal homeostasis of T cells, as manifested by the disrupted development and maintenance in T cells lacking these inhibitory molecules (Fig. 1b).
PTEN suppresses lymphoma and autoimmunity. Phosphatase and tensin homologue (PTEN) mainly functions as a lipid phosphatase and hydrolyses phosphatidylinositol-3,4,5-trisphosphate, thereby mediating the reverse reaction to PI3K (Fig. 1a). Deletion of Pten in T cells causes lymphomas that originate in the thymus and are largely driven by MYC overexpression44,45,46,47,48,49. Pten−/− thymocytes exhibit markedly elevated AKT and mTOR activities even before oncogenic transformation47,50. Importantly, tumour development is blocked by the inhibition of mTOR through rapamycin treatment or by the deletion of 3-phosphoinositide-dependent protein kinase 1 (Pdk1), indicating a crucial role for AKT and mTOR signalling in this process47,48. Somewhat unexpectedly, PTEN deficiency does not overtly disrupt the homeostasis of premalignant thymocytes, except for some minor defects in cell size and in the generation of double-positive thymocytes during ontogeny and of invariant natural killer T cells (iNKT cells)47,50,51.
Pten+/− mice develop a late-onset autoimmune disease that is associated with resistance to apoptosis mediated by CD95 (also known as FAS)52. Consistent with this observation, complete loss of PTEN in T cells impairs central tolerance as well as peripheral tolerance44 and disrupts the induction of forkhead box P3 (FOXP3) expression53. Recent studies demonstrate that the development of autoimmunity in Pten−/− mice can be exclusively mediated by peripheral T cells. Therefore, the two main defects that result from the loss of PTEN function — lymphoma and autoimmunity — are separable and are mediated by T cells at distinct developmental stages49. Because of the pivotal roles for PTEN in immune homeostasis and function, molecular signals that regulate PTEN have received considerable interest. Work from several independent groups has revealed that the microRNA cluster mir-17-92 represses the expression of PTEN to promote AKT–mTOR activity in lymphocytes, and consequently controls PTEN-dependent autoimmune and oncogenic functions54,55,56.
TSC1 maintains the quiescence of naive T cells. TSC1 and TSC2 function as an integral complex to stringently control mTORC1 activity (Fig. 1a). We and others have recently found that TSC1-mediated control of mTOR signalling enforces a programme of quiescence in naive T cells by controlling cell size, cell cycle entry and metabolic machinery7,8,9. The abrogation of quiescence predisposes TSC1-deficient T cells to apoptosis, and this results in the loss of conventional T cells and iNKT cells. The remaining Tsc1−/− T cells exist in a unique 'semi-activated' (CD44+CD122−) state in vivo and exhibit increased levels of activation, cell cycle entry and cytokine expression after acute TCR stimulation compared with activated wild-type T cells. Despite this, TSC1-deficient mice fail to generate effective antibacterial responses, even when the excessive apoptosis is largely blocked, which suggests that the maintenance of naive T cell quiescence is important for a productive immune response7. The precise mechanism by which TSC1 deficiency dampens the immune response is uncertain, but it may involve premature activation of the cell cycle and metabolic machineries and of transcriptional responses in Tsc1−/− naive T cells before they receive proper TCR signals7. These studies identify TSC1 as a bona fide factor in establishing naive T cell quiescence to facilitate immune homeostasis and function.
Tsc1−/− peripheral T cells exhibit increased mTORC1 activity but diminished mTORC2 activity compared with wild-type peripheral T cells7,8,9. Treatment of Tsc1−/− mice with rapamycin partly rectifies the altered T cell homeostasis and survival, whereas loss of mTORC2 alone has no apparent effect. These results indicate that mTORC1 activation makes a crucial contribution to T cell homeostasis7. By contrast, in mature thymocytes lacking TSC1, mTORC1 activity is increased but cell survival is not affected7. Moreover, although PTEN-deficient T cells upregulate mTORC1 activity, they largely maintain their quiescence before tumour development47,50, and this suggests a crucial but cell context-dependent effect of mTORC1 on T cell homeostasis. Collectively, these data illustrate that the active control of mTORC1 by TSC1 is a key checkpoint that enforces the quiescence of naive T cells in the periphery7,8,9.
LKB1 promotes T cell development and survival. Liver kinase B1 (LKB1; also known as STK11) phosphorylates and activates AMPK subfamily members in response to energy depletion. Deletion of Lkb1 in T cells results in profound defects in multiple compartments, including extensive apoptosis of T cells, impaired thymic selection and dysregulated T cell metabolism and proliferation57,58,59,60. Despite the well-documented role of the LKB1–AMPK axis in mTORC1 inhibition61 (Fig. 1a), the signalling mechanisms downstream of LKB1 in T cells are not fully understood. The increased mTORC1 activity in Lkb1−/− T cells contributes to excessive cytokine production60, but whether it leads to the reduced cell survival or other defects is unclear. Also, although AMPK activity is diminished in Lkb1−/− T cells57,58,59,60 and AMPK is activated by TCR stimulation62, loss of AMPKα1 (the predominant AMPK isoform in T cells) causes only a slight disturbance of T cell homeostasis60,63. Therefore, T cell homeostasis probably requires additional AMPK-related kinases that are regulated by LKB1 (Ref. 61).
Together, analyses of T cells deficient in the tumour suppressors PTEN, TSC1 and LKB1 have provided new insights into mechanisms of immune homeostasis. Notably, haematopoietic stem cells (HSCs) that lack these molecules exhibit some analogous defects to the mutant T cells in the control of proliferation, survival and function64,65,66,67,68,69,70, suggesting that HSCs and naive T cells have a common requirement for these pathways for their proper maintenance. The defects in PTEN- or TSC1-deficient HSCs can largely be rescued by rapamycin64,66,67, whereas LKB1 functions independently of mTOR in HSCs68,69,70. Therefore, despite their shared ability to inhibit mTOR signalling, these molecules use distinct pathways for the homeostatic control of HSCs and possibly T cells. Aside from these well-characterized mTOR inhibitory molecules, recent studies have identified additional negative regulators of mTOR with important roles in cell signalling and disease regulation. These regulators include DEPTOR (DEP domain-containing mTOR-interacting protein)71, sestrin 1 and sestrin 2 (Ref. 72), and the mTORC1 component PRAS40 (40 kDa proline-rich AKT substrate)73,74. However, the roles of these molecules in T cell immunity have not been addressed.
Moreover, mTOR may further interact and crosstalk with transcriptional and immune signalling pathways to mediate T cell homeostasis. In particular, forkhead box O1 (FOXO1) — a crucial transcription factor for the survival and trafficking of naive T cells75,76 — is under the stringent control of AKT-mediated phosphorylation and nuclear exclusion and thus could serve as an important downstream target of mTORC2. Consistent with this idea, transcriptional targets of FOXO1 — including IL-7 receptor subunit-α (IL-7Rα) and trafficking molecules such as CD62L — have altered expression levels in T cells that are treated with mTOR or AKT inhibitors, as well as in PTEN-, RICTOR- or PDK1-deficient T cells75,77,78,79. The functional link and physiological relevance of these interactions remain to be fully defined. Finally, diacylglycerol kinases have been shown to inhibit TCR-induced mTOR activity by downregulating diacylglycerol-mediated RAS signalling80, which may contribute to T cell homeostasis and function.
mTOR in T helper cell differentiation
Early studies of mTOR signalling in T cell responses centred on the antiproliferative effect of the inhibitor rapamycin. Analyses of Mtor−/− T cells have confirmed a role for mTOR in cell cycle progression, although the proliferation of these T cells is delayed but not abolished6. Accumulating evidence, however, highlights a central role for mTOR as a fundamental determinant of cell fate in antigen-activated CD4+ and CD8+ T cells. The inhibition of mTOR with rapamycin facilitates the induction of anergic and regulatory CD4+ T cells — two crucial components of peripheral tolerance (Box 1) — as well as the differentiation of memory CD8+ T cells (Box 2). Excellent reviews discuss the role of mTOR signalling in the differentiation of regulatory, effector and memory T cells4,81,82,83, so here I focus mainly on more recent genetic evidence that establishes a crucial role for mTOR in TH cell differentiation.
Decision making between effector T cells and TReg cells. Naive CD4+ T cells respond to antigen stimulation by developing into distinct effector cell lineages (such as TH1, TH2 and TH17 cells), which have specialized properties and effector functions (Fig. 1c), or into TReg cells, which prevent excessive immune reactions84. Powell and colleagues revealed that mTOR promotes the differentiation of effector T cells6. Mtor−/− T cells have normal activation markers and levels of IL-2 production in response to TCR stimulation, but fail to differentiate into TH1, TH2 and TH17 cells. Furthermore, they are unable to activate lineage-selective STAT proteins or express the lineage-selective master transcription factors (these are STAT4 and T-bet for TH1 cells; STAT6 and GATA3 for TH2 cells; and STAT3 and RORγt for TH17 cells). These results identify an indispensable role for mTOR in the differentiation of CD4+ effector T cells6. Conversely, TCR stimulation of Mtor−/− T cells results in the spontaneous induction of FOXP3 expression, even in the absence of exogenous cytokines6. This phenotype is not observed in either RHEB- or RICTOR-deficient T cells, indicating that both mTOR complexes contribute to the inhibition of TReg cell induction, possibly through distinct downstream mechanisms6,30,77 (Fig. 2a).
a | For the differentiation of regulatory T (TReg) cells, the induction of forkhead box P3 (FOXP3) expression depends on the transcription factors SMAD3, SMAD4, forkhead box O1 (FOXO1) and FOXO3. Mammalian target of rapamycin (mTOR) inhibits the generation of induced TReg cells by antagonizing the function of SMAD3 and SMAD4 downstream of transforming growth factor-β receptor (TGFβR) signalling, and by inducing the nuclear exclusion of FOXO1 and FOXO3. These two effects are probably mediated by mTOR complex 1 (mTORC1) and mTORC2, respectively. b | In the differentiation of T helper 1 (TH1) cells, mTORC1 inhibits the induction of suppressor of cytokine signalling 3 (SOCS3) — a crucial negative regulator of signal transducer and activator of transcription 4 (STAT4) — to promote interleukin-12 receptor (IL-12R) signalling and TH1 cell differentiation. In addition, mTORC2 is required for the activation of AKT, which also contributes to interferon-γ (IFNγ) production. c | mTORC2 promotes TH2 cell differentiation via two mechanisms: by preventing the expression of SOCS5, which is a negative regulator of IL-4R and STAT6 signalling; and by activating protein kinase Cθ (PKCθ) and thereby promoting nuclear factor-κB (NF-κB)-mediated transcription. By contrast, mTORC1 negatively regulates STAT6 signalling and TH2 cell differentiation. PDK1, 3-phosphoinositide-dependent protein kinase 1; PI3K, phosphoinositide 3-kinase; PtdIns(4,5)P2, phosphatidylinositol-4,5-bisphosphate; PtdIns(3,4,5)P3, phosphatidylinositol-3,4,5-trisphosphate; PTEN, phosphatase and tensin homologue; TCR, T cell receptor; TGFβR, TGFβ receptor.
TH1 and TH17 cell differentiation. The two mTOR complexes have disparate effects on effector T cell differentiation. Deficiency of RHEB, and consequently loss of mTORC1 activity, largely recapitulates the impaired TH1 and TH17 cell differentiation in Mtor−/− T cells, suggesting that mTORC1 mediates the mTOR-dependent differentiation of TH1 and TH17 cells30. Rheb−/− T cells express increased levels of suppressor of cytokine signalling 3 (SOCS3), which is a negative regulator of STAT signalling, and silencing of SOCS3 expression restores TH1 cell differentiation in these cells (Fig. 2b). Thus, mTORC1 signalling promotes TH1 cell differentiation by modulating cytokine signalling30. As mTORC1 activity can also be regulated by upstream signals other than RHEB12, how such RHEB-independent pathways contribute to T cell differentiation awaits future investigation. In addition, deficiency of RICTOR, and thus loss of mTORC2 activity, reduces TH1 cell differentiation through the downregulation of AKT signalling77 (Fig. 2b), although another independent study shows a dispensable role for mTORC2 in TH1 cell differentiation30. How mTOR affects TH17 cell differentiation remains to be fully established, but the mechanism could involve, in part, the upregulation of hypoxia-inducible factor 1α (HIF1α)85,86 (see below for details).
TH2 cell differentiation. In contrast to RHEB–mTORC1 signalling, mTORC2 signalling is required for TH2 cell differentiation (Fig. 2c). Two groups have independently demonstrated that loss of RICTOR impairs TH2 cell differentiation in vitro and in vivo, without appreciably affecting the development of TH17 cells30,77. In the study by Lee et al., the inability of Rictor−/− T cells to differentiate into TH2 cells was attributed to decreased protein kinase Cθ (PKCθ) activity and nuclear factor-κB (NF-κB)-mediated transcription, as complementation with activated PKCθ restored the TH2 cell defect77. In a separate study, Delgoffe et al. described an increase in SOCS5 expression in Rictor−/− T cells that accounted for the diminished TH2 cell response30. Collectively, these results establish mTORC2 as a crucial regulator of TH2 cell differentiation30,77, and future studies will determine whether PKCθ, NF-κB and SOCS5 act in separate pathways or are components of the same signalling cascade. By contrast, mTORC1 negatively controls TH2 cell differentiation, as indicated by the increased activation of STAT6 and expression of GATA3 in Rheb−/− T cells30 (Fig. 2c).
In summary, mTOR dictates cell fate decisions in effector and regulatory T cells, with mTORC1 and mTORC2 having distinct effects on immune receptor signalling. These effects are further shaped by metabolic pathways, which are discussed below. Moreover, loss of mTORC1 or mTORC2 activity impairs T cell proliferation, with a stronger effect observed in Rheb−/− T cells30,77. Cell cycle progression is known to be a prerequisite for T cell differentiation, in part because it allows for the epigenetic remodelling of cytokine loci87. It would be informative to examine how mTORC1 and mTORC2 regulate cell cycle progression, and whether this affects epigenetic regulation or lineage differentiation.
Effector pathways and metabolic regulation
To orchestrate T cell homeostasis and differentiation, mTOR regulates several downstream pathways, including those involved in immune receptor signalling, metabolic programming and T cell trafficking (Box 3). Whereas the effects of mTOR on immune receptor signalling molecules, as described above, are easy to appreciate, the relative contribution of mTOR to T cell metabolism and trafficking and how this influences cell fate decisions in vivo remain under debate4,81,83. However, evidence is emerging that mTOR serves as a signalling node to regulate both the metabolism and the migration of T cells and to further link these processes to immune signalling and transcriptional networks. This would ensure that the metabolic programme and migratory activity of a T cell match its cell fate decision. In particular, recent studies have revealed exciting new findings regarding how mTOR-dependent metabolism controls T cell fate, an area with notable therapeutic implications.
Metabolism in T cell activation and differentiation. Naive T cells have a catabolic metabolism through which they generate ATP via the tricarboxylic acid (TCA) cycle and oxidative phosphorylation. T cell activation markedly increases the uptake and consumption of glucose and glutamine, with a concomitant suppression of fatty acid oxidation43,88,89,90 (Fig. 3a). Among the central regulators of this metabolic reprogramming induced by TCR stimulation are PI3K–AKT signalling and MYC, which regulate glucose metabolism and the global metabolic transcriptome, respectively88,91,92. Additional transcription factors such as oestrogen-related receptor-α (ERRα) also contribute93, but HIF1α — which has overlapping functions with MYC in cancer cell metabolism94 — is not required for TCR activation-induced metabolic reprogramming88. The PI3K–AKT pathway signals through mTOR, which engages in extensive crosstalk with MYC in many cellular contexts95. In activated T cells, rapamycin inhibits the expression of MYC and the induction of glycolysis85,88, whereas T cells lacking MYC fail to fully activate mTORC1 in response to TCR signals88. Consistent with a pivotal role of mTOR in T cell metabolism, Tsc2−/− T cells, which have constitutive mTOR activity, are highly glycolytic after TCR stimulation93. Moreover, mTOR orchestrates the metabolic programme of naive T cells under steady-state conditions, as gene expression programmes for the metabolism of glucose, nucleotides and amino acids are abnormally upregulated in Tsc1−/− naive T cells, which have increased mTORC1 activity7. In summary, mTOR has a crucial role in orchestrating the metabolic programmes of both naive and activated T cells.
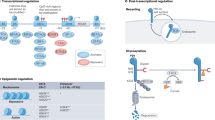
a | This schematic shows the main metabolic programmes that occur in different T cell subsets. b | This scheme shows proposed mechanisms through which metabolic pathways mediated by mammalian target of rapamycin (mTOR) affect T cell differentiation. An important upstream signal for mTOR activation is nutrients, in particular essential amino acids, the levels of which are actively controlled by dendritic cells. Once activated, mTOR serves as a platform to engage several downstream effector pathways, including immune receptor signalling, metabolic programmes and migratory activity. mTOR promotes metabolism by activating a gene expression programme that consists of metabolic gene targets of the transcription factors hypoxia-inducible factor 1α (HIF1α), MYC and sterol regulatory element-binding protein (SREBP). The expressed target proteins in turn affect biosynthetic and bioenergetic pathways. To explain how these metabolic pathways regulate T cell differentiation, three potential downstream mechanisms are proposed: signal crosstalk through metabolite-sensitive signalling targets; feedback control; and selective expansion and survival. Among metabolites with signalling activities, reactive oxygen species (ROS) oxidize the catalytic cysteine residues of phosphatases to cause the inactivation of these enzymes, whereas NAD+ is required for the activity of sirtuin family deacetylases. α-KG, α-ketoglutarate; ETC, electron-transport chain; FAO, fatty acid oxidation; ROS, reactive oxygen species; TCA, tricarboxylic acid cycle.
Recent studies demonstrate that a metabolic switch to AMPK-dependent fatty acid oxidation is required for the differentiation of CD8+ memory T cells and CD4+ TReg cells96,97, both of which are less anabolic than their effector cell counterparts4. The defective memory formation in CD8+ T cells lacking TNFR-associated factor 6 (TRAF6) is associated with a failure to upregulate fatty acid oxidation, and enhancement of fatty acid oxidation by treatment with metformin (which activates AMPK) or rapamycin restores memory formation96. Notably, AKT-independent metabolic responses have been identified in CD8+ T cells79,83, although such findings do not necessarily exclude a role for mTOR in these processes because AKT inhibition has only a weak effect on mTOR activity in CD8+ T cells79. In CD4+ T cells, elevated levels of fatty acid oxidation and AMPK activity are associated with differentiation into TReg cells but not effector TH cells, and inhibition of fatty acid oxidation prevents rapamycin-induced TReg cell generation97. Therefore, a common component for the differentiation of memory T cells and TReg cells is the selective requirement for fatty acid oxidation, in a process reciprocally regulated by mTOR and AMPK.
In contrast to the differentiation of TReg cells and memory CD8+ T cells, effector T cell differentiation is accompanied by strong upregulation of glycolysis. Pharmacological blocking of mTOR or glycolysis reduces the differentiation of TH1, TH2 and TH17 cells85,97. HIF1α is selectively expressed in T cells undergoing TH17 cell differentiation, and its induction requires signalling through mTOR85,86. HIF1α is required for mediating glycolysis during TH17 cell differentiation and contributes to lineage choices between TH17 cells and induced TReg cells85. Therefore, the mTOR-dependent induction of the transcription factors MYC and HIF1α orchestrates a metabolic checkpoint in TCR-activated and TH17-polarized cells, respectively85,88. HIF1α has also been shown to have direct effects in promoting the degradation of FOXP3 and the transcriptional activity of RORγt86. Interestingly, similarly to the induction of TReg cells, TH17 cell differentiation seems to be particularly sensitive to metabolic perturbations. Impairments in TH17 cell differentiation result not only from the blockade of glycolysis85, but also from the depletion of specific amino acids98 and from increases in lipid metabolism99. As mTORC1 is a key regulator of amino acid and lipid metabolism100, it will be interesting to examine whether these additional metabolic processes in TH17 cells are regulated by mTOR.
Sensing and propagating metabolic cues. As a central environmental sensor, mTOR links growth factor signalling with the availability of nutrients, especially amino acids1,101. The ancient pathways of amino acid metabolism and mTOR-mediated metabolic control are exploited by the immune system as important mechanisms for immune regulation102. For example, in response to TReg cell-mediated suppression, DCs upregulate enzymes that consume multiple essential amino acids that are present in the tissue microenvironment. Consequently, T cells respond to nutrient starvation by downregulating the activity of mTORC1 and inducing FOXP3 expression103. Although other nutrient sensors, such as the kinase GCN2 (also known as EIF2AK4), have been identified in T cells104, mTORC1 seems to have a dominant role in sensing amino acid availability103,105. Consistent with this notion, the inhibition of mTOR-dependent amino acid or glucose metabolism attenuates T cell activation and instead induces T cell anergy21,22 (Box 1).
Despite these advances, the extent to which mTOR-dependent metabolic pathways regulate T cell differentiation remains a contentious issue4,81,83. As T cell fate is ultimately manifested as lineage-specific gene expression programmes, how do the metabolic intermediates downstream of mTOR activation dictate signalling and transcriptional events? Here, I discuss three potential mechanisms (Fig. 3b). First, direct signalling activities have been identified for certain metabolic intermediates, such as reactive oxygen species (ROS) and nicotinamide adenine dinucleotide (NAD+), the production of which during the metabolic flux depends on mTOR. These intermediates signal by serving as substrates or modifiers of enzymes and other regulators, and thereby function as metabolic checkpoints106. Given the recently identified roles of ROS and the NAD+-dependent deacetylase sirtuin 1 in T cell function and differentiation107,108, mTOR-dependent metabolic flux may directly engage signal transduction and crosstalk.
Second, nutrient or energetic signals can mediate positive and negative feedback on mTOR activity itself, which may affect not only cell metabolism but also other mTOR-dependent events, such as immune receptor signalling. In particular, the activation of mTORC1 by amino acids and the crosstalk of mTOR with MYC upregulate the expression of the amino acid transporter CD98, which establishes a positive loop that amplifies mTOR signalling in T cells88,101. Considering the classic role of mTOR in protein translation, which consumes amino acids, it is perhaps appropriate that mTOR activity is regulated by such a positive loop. Consistent with this notion, inhibition of translation by cycloheximide considerably increases mTORC1 activity, presumably by elevating the levels of intracellular amino acids109. However, excessive mTORC1 activation may deplete cellular ATP and cause energetic stress, which activates AMPK to negatively control mTOR activity110.
Third, metabolism is intricately linked to the cell cycle and apoptotic machineries111,112. T cells with suitable metabolism are more likely to survive and/or proliferate and, consequently, on a population level, are selected to develop into a particular lineage. In support of this notion, rapamycin differentially affects the proliferation and survival of effector T cells and TReg cells113,114,115,116. Future efforts to identify the molecular components that orchestrate these processes should provide exciting insights into the interface between metabolism and immunity.
Therapeutic targeting of mTOR
The immunosuppressive effect of rapamycin was recognized in the 1970s, before the identification of its molecular target. Recent studies suggest that blocking mTOR not only mediates immunosuppression to reduce transplant rejection and autoimmune disorders, but also boosts immunity under selective conditions and affects other aspects of T cell homeostasis and function.
mTOR inhibition for immunosuppression. In the late 1990s, the US Food and Drug Administration approved the use of rapamycin to prevent rejection in kidney transplantation. A major mechanism of action for rapamycin is the induction and expansion of TReg cell populations and the inhibition of effector T cell differentiation, as described above. This mechanism is distinct from that of other immunosuppressants (such as cyclosporine A and FK506), which mainly block Ca2+ signalling and calcineurin activation downstream of TCR stimulation, and thus allows for the design of combinatorial therapies with rapamycin and other immunosuppressive agents. This is important because rapamycin monotherapy has a rather weak effect in preventing graft rejection, despite its multiple immunomodulatory functions.
The immunosuppressive effect of rapamycin is also apparent in patients with and experimental models of systemic autoimmune diseases, such as systemic lupus erythematosus and rheumatoid arthritis, as well as organ-specific autoimmune disorders117,118. Rapamycin probably establishes long-term immune tolerance in these models by expanding TReg cell populations and inhibiting the differentiation and function of effector T cells. Notably, rapamycin also has potent effects on DCs and other immune components2, and the effects of rapamycin in vivo might be attributed to these cells, in addition to T cells.
Rapamycin for vaccine development to boost immunity. Despite the well-established immunosuppressive effects of mTOR inhibition, rapamycin and metformin promote the generation of protective T cell memory in models of infection with lymphocytic choriomeningitis virus and Listeria monocytogenes. Mechanistically, this has been associated with the induction of a metabolic switch from glycolysis to fatty acid oxidation in the presence of rapamycin and metformin96,119. The immunostimulatory effect of rapamycin for CD8+ T cell responses has been extended to additional infectious models120,121 and to antitumour immune responses122,123. Therefore, mTOR inhibitors may serve as novel adjuvants for the development of vaccines against pathogens and tumours. In particular, this strategy of tumour immunotherapy is promising owing to its synergy with the direct suppression of tumour growth by mTOR inhibitors. Notably, the effect of mTOR inhibition depends on the dose range and kinetics of the treatment. Administration of a very high dose of rapamycin prevents the expansion of CD8+ T cell populations, whereas the duration and timing of rapamycin treatment affect the quantity and quality of memory T cell responses82,119. To explore the basis for rapamycin-mediated immunosuppressive and immunostimulatory effects on CD8+ T cell responses, Ferrer et al. compared the effects of rapamycin on immune responses induced by L. monocytogenes and by a skin transplant against the same antigen120. Treatment with rapamycin augmented antigen-specific T cell responses to the pathogen but not to the transplant. Thus, the environment in which an antigen is presented influences the effects of rapamycin on T cell responses120.
T cell malignancy and metabolic dysregulation. A common haematological malignancy is T cell acute lymphoblastic leukaemia (T-ALL), which frequently harbours activating mutations of NOTCH1 and/or loss-of-function mutations of PTEN. Interestingly, both types of mutation activate mTOR signalling, suggesting a pivotal role for mTOR in the development of T-ALL. Consistent with this notion, in a PTEN-deficient mouse model of T-ALL, rapamycin was effective in blocking T-ALL initiation and development. However, following rapamycin withdrawal, most of the PTEN-deficient mice rapidly became ill and died. These results indicate that leukaemia stem cells and leukaemia cell blasts have differential responses to rapamycin, which could contribute to the limited clinical efficacy of rapamycin in cancer48.
Chronic inflammation in adipose tissues has been established as a major mechanism that causes insulin resistance and the subsequent development of type 2 diabetes. Although earlier studies mostly implicated macrophages as the main inflammatory cell infiltrates124, recent reports have identified the recruitment of effector CD8+ T cells and FOXP3+ TReg cells to adipose tissues, where they function to exacerbate and ameliorate inflammation, respectively125,126,127. Given the prominent immunomodulatory role of mTOR in T cells, it will be interesting to explore whether mTOR signalling in T cells contributes to the pathogenesis of metabolic diseases. This raises the notion that therapeutic targeting of mTOR in metabolic diseases may have a dual effect, through the regulation of T cell metabolism and effector function as well as through the direct modulation of the function of metabolic tissues (such as adipose tissues, skeletal muscles and liver), in which mTOR signalling has a central role.
Targeting mTOR: beyond rapamycin. Rapamycin inhibits mTOR through an unusual allosteric mechanism that involves the interaction with its intracellular partner FKBP12. Whereas rapamycin strongly inhibits the functions of ribosomal protein S6 kinases (S6Ks) downstream of mTORC1, it has a surprisingly weak effect on the phosphorylation of the other major mTORC1 targets, eIF4E-binding proteins (4E-BPs)128. Moreover, rapamycin treatment frequently abrogates mTORC1-mediated feedback inhibition of PI3K–AKT signalling, leading to a paradoxical enhancement of AKT activity129. The incomplete inhibition of mTORC1 and the reversal of the feedback loop by rapamycin attenuate its therapeutic effects, as reflected by the disappointing therapeutic outcomes of rapamycin and other first-generation mTOR inhibitors in patients with cancer130. Therefore, selective ATP-competitive mTOR inhibitors (including Torin1 and PP242) were recently developed, and these molecules completely inhibit mTORC1 activity, including the rapamycin-resistant phosphorylation of 4E-BPs131,132,133. Compared with rapamycin, these second-generation inhibitors exert stronger inhibition of mTORC2 and thus are less likely to activate the feedback loop131,132,133. Indeed, these new inhibitors have greater antitumour effects than rapamycin134,135, and some of them have entered clinical trials as new cancer therapeutics130. Their immunomodulatory function is expected to be extensively investigated in the near future.
Because of the pleiotropic effects of mTOR in T cells, targeting the upstream and downstream components of mTOR signalling, rather than mTOR itself, offers an alternative strategy for added specificity. In particular, modulation of mTOR-dependent metabolic pathways is efficacious for modulating T cell-mediated diseases. A classic example is provided by the AMPK activators metformin and AICAR (5-aminoimidazole-4-carboxamide riboside), which block energy-mediated activation of mTORC1 without interfering with PI3K–AKT-mediated growth factor signalling. Aside from modulating T cell memory, as described above96, these AMPK activators have both prophylactic and therapeutic effects on experimental autoimmune encephalomyelitis by attenuating the severity of disease136,137. Similarly, blocking glycolysis protects mice from autoimmune neuroinflammation mediated by TH17 cells85. Moreover, blocking upstream activators of mTOR in T cells, such as S1PR1 and the leptin receptor, has similar effects to rapamycin in modulating T cell responses in selective models33,37. Further studies of mTOR-dependent signalling axes will provide more opportunities to specifically target mTOR-associated pathways.
Concluding remarks
Cell fate decisions mediated by mTOR are of particular importance to T cells because of the unique features of these cells. Such features include: their constant exposure to unpredictable pathogen threats; their extensive proliferation following antigenic stimulation; and their continuous migration in a variety of tissues. Recent advances have established mTOR as a fundamental determinant of T cell homeostatic and functional fates, but many questions remain. Further studies are needed to elucidate the upstream signals and downstream effectors that mediate T cell homeostasis and antigen-specific immune responses. Also, we have yet to define the emerging roles of mTOR in certain cell biological processes such as autophagy, a process that is crucial for T cell homeostasis and function138. Much of our current knowledge of the mTOR-controlled signalling networks is derived from the use of in vitro systems and cell lines. However, the identification of T cell-specific and context-dependent signals promises to provide more insight into the regulation of adaptive immunity. From this perspective, the use of sophisticated genetic systems offers the ultimate molecular and cellular specificities to dissect the underlying processes. Moreover, although research in this area has benefited from the use of rapamycin over the past two decades, the development and application of more potent and selective inhibitors of mTOR is essential to advance both research and clinical applications. The continued expansion of our understanding of mTOR signalling will provide legitimate therapeutic opportunities for T cell-mediated disorders.
References
Zoncu, R., Efeyan, A. & Sabatini, D. M. mTOR: from growth signal integration to cancer, diabetes and ageing. Nature Rev. Mol. Cell Biol. 12, 21–35 (2011).
Thomson, A. W., Turnquist, H. R. & Raimondi, G. Immunoregulatory functions of mTOR inhibition. Nature Rev. Immunol. 9, 324–337 (2009).
Araki, K., Ellebedy, A. H. & Ahmed, R. TOR in the immune system. Curr. Opin. Cell Biol. 23, 707–715 (2011).
Powell, J. D. & Delgoffe, G. M. The mammalian target of rapamycin: linking T cell differentiation, function, and metabolism. Immunity 33, 301–311 (2010).
Iwasaki, A. & Medzhitov, R. Regulation of adaptive immunity by the innate immune system. Science 327, 291–295 (2010).
Delgoffe, G. M. et al. The mTOR kinase differentially regulates effector and regulatory T cell lineage commitment. Immunity 30, 832–844 (2009). This paper provides the first genetic evidence that mTOR promotes CD4+ effector T cell differentiation while inhibiting the induction of T Reg cells.
Yang, K., Neale, G., Green, D. R., He, W. & Chi, H. The tumor suppressor Tsc1 enforces quiescence of naive T cells to promote immune homeostasis and function. Nature Immunol. 12, 888–897 (2011).
Wu, Q. et al. The tuberous sclerosis complex–mammalian target of rapamycin pathway maintains the quiescence and survival of naive T cells. J. Immunol. 187, 1106–1112 (2011). References 7 and 8 show that the maintenance of T cell quiescence requires TSC1.
O'Brien, T. F. et al. Regulation of T-cell survival and mitochondrial homeostasis by TSC1. Eur. J. Immunol. 41, 3361–3370 (2011).
Sancak, Y. et al. The Rag GTPases bind raptor and mediate amino acid signaling to mTORC1. Science 320, 1496–1501 (2008).
Sancak, Y. et al. Ragulator–Rag complex targets mTORC1 to the lysosomal surface and is necessary for its activation by amino acids. Cell 141, 290–303 (2010).
Gwinn, D. M. et al. AMPK phosphorylation of raptor mediates a metabolic checkpoint. Mol. Cell 30, 214–226 (2008).
Zheng, M. et al. Inactivation of Rheb by PRAK-mediated phosphorylation is essential for energy-depletion-induced suppression of mTORC1. Nature Cell Biol. 13, 263–272 (2011).
Wu, X. N. et al. Phosphorylation of Raptor by p38β participates in arsenite-induced mammalian target of rapamycin complex 1 (mTORC1) activation. J. Biol. Chem. 286, 31501–31511 (2011).
Zinzalla, V., Stracka, D., Oppliger, W. & Hall, M. N. Activation of mTORC2 by association with the ribosome. Cell 144, 757–768 (2011).
Oh, W. J. et al. mTORC2 can associate with ribosomes to promote cotranslational phosphorylation and stability of nascent Akt polypeptide. EMBO J. 29, 3939–3951 (2010).
Saci, A., Cantley, L. C. & Carpenter, C. L. Rac1 regulates the activity of mTORC1 and mTORC2 and controls cellular size. Mol. Cell 42, 50–61 (2011).
Turner, M. S., Kane, L. P. & Morel, P. A. Dominant role of antigen dose in CD4+Foxp3+ regulatory T cell induction and expansion. J. Immunol. 183, 4895–4903 (2009).
Katzman, S. D. et al. Duration of antigen receptor signaling determines T-cell tolerance or activation. Proc. Natl Acad. Sci. USA 107, 18085–18090 (2010).
Colombetti, S., Basso, V., Mueller, D. L. & Mondino, A. Prolonged TCR/CD28 engagement drives IL-2-independent T cell clonal expansion through signaling mediated by the mammalian target of rapamycin. J. Immunol. 176, 2730–2738 (2006).
Zheng, Y. et al. A role for mammalian target of rapamycin in regulating T cell activation versus anergy. J. Immunol. 178, 2163–2170 (2007).
Zheng, Y., Delgoffe, G. M., Meyer, C. F., Chan, W. & Powell, J. D. Anergic T cells are metabolically anergic. J. Immunol. 183, 6095–6101 (2009).
So, T., Choi, H. & Croft, M. OX40 complexes with phosphoinositide 3-kinase and protein kinase B (PKB) to augment TCR-dependent PKB signaling. J. Immunol. 186, 3547–3555 (2011).
Francisco, L. M. et al. PD-L1 regulates the development, maintenance, and function of induced regulatory T cells. J. Exp. Med. 206, 3015–3029 (2009).
Wofford, J. A., Wieman, H. L., Jacobs, S. R., Zhao, Y. & Rathmell, J. C. IL-7 promotes Glut1 trafficking and glucose uptake via STAT5-mediated activation of Akt to support T-cell survival. Blood 111, 2101–2111 (2008).
Rathmell, J. C., Farkash, E. A., Gao, W. & Thompson, C. B. IL-7 enhances the survival and maintains the size of naive T cells. J. Immunol. 167, 6869–6876 (2001).
Rao, R. R., Li, Q., Odunsi, K. & Shrikant, P. A. The mTOR kinase determines effector versus memory CD8+ T cell fate by regulating the expression of transcription factors T-bet and Eomesodermin. Immunity 32, 67–78 (2010). This study describes mTOR-dependent regulation of T-bet and EOMES as important mechanisms for CD8+ effector and memory T cell differentiation.
Stephenson, L. M., Park, D. S., Mora, A. L., Goenka, S. & Boothby, M. Sequence motifs in IL-4Rα mediating cell-cycle progression of primary lymphocytes. J. Immunol. 175, 5178–5185 (2005).
Gulen, M. F. et al. The receptor SIGIRR suppresses Th17 cell proliferation via inhibition of the interleukin-1 receptor pathway and mTOR kinase activation. Immunity 32, 54–66 (2010).
Delgoffe, G. M. et al. The kinase mTOR regulates the differentiation of helper T cells through the selective activation of signaling by mTORC1 and mTORC2. Nature Immunol. 12, 295–303 (2011). This paper describes the differential requirements for mTORC1 and mTORC2 in T H 1, T H 2 and T H 17 cell differentiation.
Zhou, J. et al. Activation of the PTEN/mTOR/STAT3 pathway in breast cancer stem-like cells is required for viability and maintenance. Proc. Natl Acad. Sci. USA 104, 16158–16163 (2007).
Lord, G. M. et al. Leptin modulates the T-cell immune response and reverses starvation-induced immunosuppression. Nature 394, 897–901 (1998).
Procaccini, C. et al. An oscillatory switch in mTOR kinase activity sets regulatory T cell responsiveness. Immunity 33, 929–941 (2010). Here, the authors describe a leptin–mTOR axis that modulates the responsiveness of T Reg cells to TCR signals.
De Rosa, V. et al. A key role of leptin in the control of regulatory T cell proliferation. Immunity 26, 241–255 (2007).
Galgani, M. et al. Leptin modulates the survival of autoreactive CD4+ T cells through the nutrient/energy-sensing mammalian target of rapamycin signaling pathway. J. Immunol. 185, 7474–7479 (2010).
Liu, G. et al. The receptor S1P1 overrides regulatory T cell-mediated immune suppression through Akt–mTOR. Nature Immunol. 10, 769–777 (2009).
Liu, G., Yang, K., Burns, S., Shrestha, S. & Chi, H. The S1P1–mTOR axis directs the reciprocal differentiation of TH1 and Treg cells. Nature Immunol. 11, 1047–1056 (2010). This study identifies an important role for S1PR1 in the activation of mTOR and the modulation of T H 1 and T Reg cell differentiation.
Geng, D. et al. When Toll-like receptor and T-cell receptor signals collide: a mechanism for enhanced CD8 T-cell effector function. Blood 116, 3494–3504 (2010).
Quigley, M., Martinez, J., Huang, X. & Yang, Y. A critical role for direct TLR2-MyD88 signaling in CD8 T-cell clonal expansion and memory formation following vaccinia viral infection. Blood 113, 2256–2264 (2009).
Rahman, A. H. et al. Antiviral memory CD8 T-cell differentiation, maintenance, and secondary expansion occur independently of MyD88. Blood 117, 3123–3130 (2011).
Ciofani, M. & Zuniga-Pflucker, J. C. Notch promotes survival of pre-T cells at the β-selection checkpoint by regulating cellular metabolism. Nature Immunol. 6, 881–888 (2005).
Sprent, J. & Surh, C. D. Normal T cell homeostasis: the conversion of naive cells into memory-phenotype cells. Nature Immunol. 12, 478–484 (2011).
Michalek, R. D. & Rathmell, J. C. The metabolic life and times of a T-cell. Immunol. Rev. 236, 190–202 (2010).
Suzuki, A. et al. T cell-specific loss of Pten leads to defects in central and peripheral tolerance. Immunity 14, 523–534 (2001).
Xue, L., Nolla, H., Suzuki, A., Mak, T. W. & Winoto, A. Normal development is an integral part of tumorigenesis in T cell-specific PTEN-deficient mice. Proc. Natl Acad. Sci. USA 105, 2022–2027 (2008).
Hagenbeek, T. J. & Spits, H. T-cell lymphomas in T-cell-specific Pten-deficient mice originate in the thymus. Leukemia 22, 608–619 (2008).
Finlay, D. K. et al. Phosphoinositide-dependent kinase 1 controls migration and malignant transformation but not cell growth and proliferation in PTEN-null lymphocytes. J. Exp. Med. 206, 2441–2454 (2009).
Guo, W. et al. Suppression of leukemia development caused by PTEN loss. Proc. Natl Acad. Sci. USA 108, 1409–1414 (2011).
Liu, X. et al. Distinct roles for PTEN in prevention of T cell lymphoma and autoimmunity in mice. J. Clin. Invest. 120, 2497–2507 (2010).
Hagenbeek, T. J. et al. The loss of PTEN allows TCRαβ lineage thymocytes to bypass IL-7 and pre-TCR-mediated signaling. J. Exp. Med. 200, 883–894 (2004).
Kishimoto, H. et al. The Pten/PI3K pathway governs the homeostasis of Vα14 iNKT cells. Blood 109, 3316–3324 (2007).
Di Cristofano, A. et al. Impaired Fas response and autoimmunity in Pten+/− mice. Science 285, 2122–2125 (1999).
Sauer, S. et al. T cell receptor signaling controls Foxp3 expression via PI3K, Akt, and mTOR. Proc. Natl Acad. Sci. USA 105, 7797–7802 (2008).
Xiao, C. et al. Lymphoproliferative disease and autoimmunity in mice with increased miR-17-92 expression in lymphocytes. Nature Immunol. 9, 405–414 (2008).
Olive, V. et al. miR-19 is a key oncogenic component of mir-17-92. Genes Dev. 23, 2839–2849 (2009).
Jiang, S. et al. Molecular dissection of the miR-17-92 cluster's critical dual roles in promoting Th1 responses and preventing inducible TReg differentiation. Blood 118, 5487–5497 (2011).
Tamas, P. et al. LKB1 is essential for the proliferation of T-cell progenitors and mature peripheral T cells. Eur. J. Immunol. 40, 242–253 (2010).
Cao, Y. et al. The serine/threonine kinase LKB1 controls thymocyte survival through regulation of AMPK activation and Bcl-XL expression. Cell Res. 20, 99–108 (2010).
Cao, Y. et al. LKB1 regulates TCR-mediated PLCγ1 activation and thymocyte positive selection. EMBO J. 30, 2083–2093 (2011).
Maciver, N. J. et al. The liver kinase B1 is a central regulator of T cell development, activation, and metabolism. J. Immunol. 187, 4187–4198 (2011).
Mihaylova, M. M. & Shaw, R. J. The AMPK signalling pathway coordinates cell growth, autophagy and metabolism. Nature Cell Biol. 13, 1016–1023 (2011).
Tamas, P. et al. Regulation of the energy sensor AMP-activated protein kinase by antigen receptor and Ca2+ in T lymphocytes. J. Exp. Med. 203, 1665–1670 (2006).
Mayer, A., Denanglaire, S., Viollet, B., Leo, O. & Andris, F. AMP-activated protein kinase regulates lymphocyte responses to metabolic stress but is largely dispensable for immune cell development and function. Eur. J. Immunol. 38, 948–956 (2008).
Yilmaz, O. H. et al. Pten dependence distinguishes haematopoietic stem cells from leukaemia-initiating cells. Nature 441, 475–482 (2006).
Zhang, J. et al. PTEN maintains haematopoietic stem cells and acts in lineage choice and leukaemia prevention. Nature 441, 518–522 (2006).
Chen, C. et al. TSC–mTOR maintains quiescence and function of hematopoietic stem cells by repressing mitochondrial biogenesis and reactive oxygen species. J. Exp. Med. 205, 2397–2408 (2008).
Gan, B. et al. mTORC1-dependent and -independent regulation of stem cell renewal, differentiation, and mobilization. Proc. Natl Acad. Sci. USA 105, 19384–19389 (2008).
Gan, B. et al. Lkb1 regulates quiescence and metabolic homeostasis of haematopoietic stem cells. Nature 468, 701–704 (2010).
Gurumurthy, S. et al. The Lkb1 metabolic sensor maintains haematopoietic stem cell survival. Nature 468, 659–663 (2010).
Nakada, D., Saunders, T. L. & Morrison, S. J. Lkb1 regulates cell cycle and energy metabolism in haematopoietic stem cells. Nature 468, 653–658 (2010).
Peterson, T. R. et al. DEPTOR is an mTOR inhibitor frequently overexpressed in multiple myeloma cells and required for their survival. Cell 137, 873–886 (2009).
Budanov, A. V. & Karin, M. p53 target genes Sestrin1 and Sestrin2 connect genotoxic stress and mTOR signaling. Cell 134, 451–460 (2008).
Sancak, Y. et al. PRAS40 is an insulin-regulated inhibitor of the mTORC1 protein kinase. Mol. Cell 25, 903–915 (2007).
Vander Haar, E., Lee, S. I., Bandhakavi, S., Griffin, T. J. & Kim, D. H. Insulin signalling to mTOR mediated by the Akt/PKB substrate PRAS40. Nature Cell Biol. 9, 316–323 (2007).
Kerdiles, Y. M. et al. Foxo1 links homing and survival of naive T cells by regulating L-selectin, CCR7 and interleukin 7 receptor. Nature Immunol. 10, 176–184 (2009).
Ouyang, W., Beckett, O., Flavell, R. A. & Li, M. O. An essential role of the Forkhead-box transcription factor Foxo1 in control of T cell homeostasis and tolerance. Immunity 30, 358–371 (2009).
Lee, K. et al. Mammalian target of rapamycin protein complex 2 regulates differentiation of Th1 and Th2 cell subsets via distinct signaling pathways. Immunity 32, 743–753 (2010). This paper describes the roles of mTORC2 in T H 1 and T H 2 cell differentiation through its effects on AKT and PKCθ signalling.
Sinclair, L. V. et al. Phosphatidylinositol-3-OH kinase and nutrient-sensing mTOR pathways control T lymphocyte trafficking. Nature Immunol. 9, 513–521 (2008). This study provided the first evidence for the regulatory role of mTOR in T cell trafficking.
Macintyre, A. N. et al. Protein kinase B controls transcriptional programs that direct cytotoxic T cell fate but is dispensable for T cell metabolism. Immunity 34, 224–236 (2011).
Gorentla, B. K., Wan, C. K. & Zhong, X. P. Negative regulation of mTOR activation by diacylglycerol kinases. Blood 117, 4022–4031 (2010).
Peter, C., Waldmann, H. & Cobbold, S. P. mTOR signalling and metabolic regulation of T cell differentiation. Curr. Opin. Immunol. 22, 655–661 (2010).
Araki, K., Youngblood, B. & Ahmed, R. The role of mTOR in memory CD8 T-cell differentiation. Immunol. Rev. 235, 234–243 (2010).
Finlay, D. & Cantrell, D. A. Metabolism, migration and memory in cytotoxic T cells. Nature Rev. Immunol. 11, 109–117 (2011).
Zhu, J., Yamane, H. & Paul, W. E. Differentiation of effector CD4 T cell populations. Annu. Rev. Immunol. 28, 445–489 (2010).
Shi, L. Z. et al. HIF1α-dependent glycolytic pathway orchestrates a metabolic checkpoint for the differentiation of TH17 and Treg cells. J. Exp. Med. 208, 1367–1376 (2011).
Dang, E. V. et al. Control of TH17/Treg balance by hypoxia-inducible factor 1. Cell 146, 772–784 (2011). References 85 and 86 describe an important role for HIF1α and metabolic pathways in T H 17 and T Reg cell differentiation.
Bird, J. J. et al. Helper T cell differentiation is controlled by the cell cycle. Immunity 9, 229–237 (1998).
Wang, R. et al. The transcription factor Myc controls metabolic reprogramming upon T lymphocyte activation. Immunity 35, 871–882 (2011).
Jones, R. G. & Thompson, C. B. Revving the engine: signal transduction fuels T cell activation. Immunity 27, 173–178 (2007).
Pearce, E. L. Metabolism in T cell activation and differentiation. Curr. Opin. Immunol. 22, 314–320 (2010).
Frauwirth, K. A. et al. The CD28 signaling pathway regulates glucose metabolism. Immunity 16, 769–777 (2002).
Rathmell, J. C. T cell Myc-tabolism. Immunity 35, 845–846 (2011).
Michalek, R. D. et al. Estrogen-related receptor-α is a metabolic regulator of effector T-cell activation and differentiation. Proc. Natl Acad. Sci. USA 108, 18348–18353 (2011).
Gordan, J. D., Thompson, C. B. & Simon, M. C. HIF and c-Myc: sibling rivals for control of cancer cell metabolism and proliferation. Cancer Cell 12, 108–113 (2007).
DeBerardinis, R. J., Lum, J. J., Hatzivassiliou, G. & Thompson, C. B. The biology of cancer: metabolic reprogramming fuels cell growth and proliferation. Cell Metab. 7, 11–20 (2008).
Pearce, E. L. et al. Enhancing CD8 T-cell memory by modulating fatty acid metabolism. Nature 460, 103–107 (2009).
Michalek, R. D. et al. Cutting edge: distinct glycolytic and lipid oxidative metabolic programs are essential for effector and regulatory CD4+ T cell subsets. J. Immunol. 186, 3299–3303 (2011).
Sundrud, M. S. et al. Halofuginone inhibits TH17 cell differentiation by activating the amino acid starvation response. Science 324, 1334–1338 (2009).
Cui, G. et al. Liver X receptor (LXR) mediates negative regulation of mouse and human Th17 differentiation. J. Clin. Invest. 121, 658–670 (2011).
Duvel, K. et al. Activation of a metabolic gene regulatory network downstream of mTOR complex 1. Mol. Cell 39, 171–183 (2010).
Nicklin, P. et al. Bidirectional transport of amino acids regulates mTOR and autophagy. Cell 136, 521–534 (2009).
Grohmann, U. & Bronte, V. Control of immune response by amino acid metabolism. Immunol. Rev. 236, 243–264 (2010).
Cobbold, S. P. et al. Infectious tolerance via the consumption of essential amino acids and mTOR signaling. Proc. Natl Acad. Sci. USA 106, 12055–12060 (2009).
Munn, D. H. et al. GCN2 kinase in T cells mediates proliferative arrest and anergy induction in response to indoleamine 2,3-dioxygenase. Immunity 22, 633–642 (2005).
Cobbold, S. P., Adams, E., Nolan, K. F., Regateiro, F. S. & Waldmann, H. Connecting the mechanisms of T-cell regulation: dendritic cells as the missing link. Immunol. Rev. 236, 203–218 (2010).
Locasale, J. W. & Cantley, L. C. Metabolic flux and the regulation of mammalian cell growth. Cell Metab. 14, 443–451 (2011).
Lee, K., Won, H. Y., Bae, M. A., Hong, J. H. & Hwang, E. S. Spontaneous and aging-dependent development of arthritis in NADPH oxidase 2 deficiency through altered differentiation of CD11b+ and Th/TReg cells. Proc. Natl Acad. Sci. USA 108, 9548–9553 (2011).
Zhang, J. et al. The type III histone deacetylase Sirt1 is essential for maintenance of T cell tolerance in mice. J. Clin. Invest. 119, 3048–3058 (2009).
Beugnet, A., Tee, A. R., Taylor, P. M. & Proud, C. G. Regulation of targets of mTOR (mammalian target of rapamycin) signalling by intracellular amino acid availability. Biochem. J. 372, 555–566 (2003).
Choo, A. Y. et al. Glucose addiction of TSC null cells is caused by failed mTORC1-dependent balancing of metabolic demand with supply. Mol. Cell 38, 487–499 (2010).
Buchakjian, M. R. & Kornbluth, S. The engine driving the ship: metabolic steering of cell proliferation and death. Nature Rev. Mol. Cell Biol. 11, 715–727 (2010).
Mason, E. F. & Rathmell, J. C. Cell metabolism: An essential link between cell growth and apoptosis. Biochim. Biophys. Acta 1813, 645–654 (2011).
Battaglia, M., Stabilini, A. & Roncarolo, M. G. Rapamycin selectively expands CD4+CD25+FoxP3+ regulatory T cells. Blood 105, 4743–4748 (2005).
Strauss, L. et al. Selective survival of naturally occurring human CD4+CD25+Foxp3+ regulatory T cells cultured with rapamycin. J. Immunol. 178, 320–329 (2007).
Zeiser, R. et al. Differential impact of mammalian target of rapamycin inhibition on CD4+CD25+Foxp3+ regulatory T cells compared with conventional CD4+ T cells. Blood 111, 453–462 (2008).
Strauss, L., Czystowska, M., Szajnik, M., Mandapathil, M. & Whiteside, T. L. Differential responses of human regulatory T cells (TReg) and effector T cells to rapamycin. PLoS ONE 4, e5994 (2009).
Young, D. A. & Nickerson-Nutter, C. L. mTOR — beyond transplantation. Curr. Opin. Pharmacol. 5, 418–423 (2005).
Fernandez, D., Bonilla, E., Mirza, N., Niland, B. & Perl, A. Rapamycin reduces disease activity and normalizes T cell activation-induced calcium fluxing in patients with systemic lupus erythematosus. Arthritis Rheum. 54, 2983–2988 (2006).
Araki, K. et al. mTOR regulates memory CD8 T-cell differentiation. Nature 460, 108–112 (2009). References 96 and 119 are the first reports to implicate mTOR and metabolic pathways in CD8+ memory T cell formation.
Ferrer, I. R. et al. Cutting edge: rapamycin augments pathogen-specific but not graft-reactive CD8+ T cell responses. J. Immunol. 185, 2004–2008 (2010).
Turner, A. P. et al. Sirolimus enhances the magnitude and quality of viral-specific CD8+ T-cell responses to vaccinia virus vaccination in rhesus macaques. Am. J. Transplant. 11, 613–618 (2011).
Li, Q. et al. A central role for mTOR kinase in homeostatic proliferation induced CD8+ T cell memory and tumor immunity. Immunity 34, 541–553 (2011).
Wang, Y., Wang, X. Y., Subjeck, J. R., Shrikant, P. A. & Kim, H. L. Temsirolimus, an mTOR inhibitor, enhances anti-tumour effects of heat shock protein cancer vaccines. Br. J. Cancer. 104, 643–652 (2011).
Chawla, A., Nguyen, K. D. & Goh, Y. P. Macrophage-mediated inflammation in metabolic disease. Nature Rev. Immunol. 11, 738–749 (2011).
Winer, S. et al. Normalization of obesity-associated insulin resistance through immunotherapy. Nature Med. 15, 921–929 (2009).
Nishimura, S. et al. CD8+ effector T cells contribute to macrophage recruitment and adipose tissue inflammation in obesity. Nature Med. 15, 914–920 (2009).
Feuerer, M. et al. Lean, but not obese, fat is enriched for a unique population of regulatory T cells that affect metabolic parameters. Nature Med. 15, 930–939 (2009).
Choo, A. Y., Yoon, S. O., Kim, S. G., Roux, P. P. & Blenis, J. Rapamycin differentially inhibits S6Ks and 4E-BP1 to mediate cell-type-specific repression of mRNA translation. Proc. Natl Acad. Sci. USA 105, 17414–17419 (2008).
O'Reilly, K. E. et al. mTOR inhibition induces upstream receptor tyrosine kinase signaling and activates Akt. Cancer Res. 66, 1500–1508 (2006).
Benjamin, D., Colombi, M., Moroni, C. & Hall, M. N. Rapamycin passes the torch: a new generation of mTOR inhibitors. Nature Rev. Drug Discov. 10, 868–880 (2011).
Apsel, B. et al. Targeted polypharmacology: discovery of dual inhibitors of tyrosine and phosphoinositide kinases. Nature Chem. Biol. 4, 691–699 (2008).
Feldman, M. E. et al. Active-site inhibitors of mTOR target rapamycin-resistant outputs of mTORC1 and mTORC2. PLoS Biol. 7, e38 (2009).
Thoreen, C. C. et al. An ATP-competitive mammalian target of rapamycin inhibitor reveals rapamycin-resistant functions of mTORC1. J. Biol. Chem. 284, 8023–8032 (2009).
Hsieh, A. C. et al. Genetic dissection of the oncogenic mTOR pathway reveals druggable addiction to translational control via 4EBP–eIF4E. Cancer Cell 17, 249–261 (2010).
Janes, M. R. et al. Effective and selective targeting of leukemia cells using a TORC1/2 kinase inhibitor. Nature Med. 16, 205–213 (2010).
Nath, N. et al. 5-aminoimidazole-4-carboxamide ribonucleoside: a novel immunomodulator with therapeutic efficacy in experimental autoimmune encephalomyelitis. J. Immunol. 175, 566–574 (2005).
Nath, N. et al. Metformin attenuated the autoimmune disease of the central nervous system in animal models of multiple sclerosis. J. Immunol. 182, 8005–8014 (2009).
Dunkle, A. & He, Y. W. Apoptosis and autophagy in the regulation of T lymphocyte function. Immunol. Res. 49, 70–86 (2011).
Haribhai, D. et al. A requisite role for induced regulatory T cells in tolerance based on expanding antigen receptor diversity. Immunity 35, 109–122 (2011).
Kang, J., Huddleston, S. J., Fraser, J. M. & Khoruts, A. De novo induction of antigen-specific CD4+CD25+Foxp3+ regulatory T cells in vivo following systemic antigen administration accompanied by blockade of mTOR. J. Leukoc. Biol. 83, 1230–1239 (2008).
Gao, W. et al. Contrasting effects of cyclosporine and rapamycin in de novo generation of alloantigen-specific regulatory T cells. Am. J. Transplant. 7, 1722–1732 (2007).
Haxhinasto, S., Mathis, D. & Benoist, C. The AKT–mTOR axis regulates de novo differentiation of CD4+Foxp3+ cells. J. Exp. Med. 205, 565–574 (2008).
Song, K., Wang, H., Krebs, T. L. & Danielpour, D. Novel roles of Akt and mTOR in suppressing TGF-β/ALK5-mediated Smad3 activation. EMBO J. 25, 58–69 (2006).
Ouyang, W. et al. Foxo proteins cooperatively control the differentiation of Foxp3+ regulatory T cells. Nature Immunol. 11, 618–627 (2010).
Harada, Y. et al. Transcription factors Foxo3a and Foxo1 couple the E3 ligase Cbl-b to the induction of Foxp3 expression in induced regulatory T cells. J. Exp. Med. 207, 1381–1391 (2010).
Kerdiles, Y. M. et al. Foxo transcription factors control regulatory T cell development and function. Immunity 33, 890–904 (2010).
Merkenschlager, M. & von Boehmer, H. PI3 kinase signalling blocks Foxp3 expression by sequestering Foxo factors. J. Exp. Med. 207, 1347–1350 (2010).
Seoane, J., Le, H. V., Shen, L., Anderson, S. A. & Massague, J. Integration of Smad and forkhead pathways in the control of neuroepithelial and glioblastoma cell proliferation. Cell 117, 211–223 (2004).
Zhang, N. & Bevan, M. J. CD8+ T cells: foot soldiers of the immune system. Immunity 35, 161–168 (2011).
van der Windt, G. J. et al. Mitochondrial respiratory capacity is a critical regulator of CD8+ T cell memory development. Immunity 36, 68–78 (2012).
Hand, T. W. et al. Differential effects of STAT5 and PI3K/AKT signaling on effector and memory CD8 T-cell survival. Proc. Natl Acad. Sci. USA 107, 16601–16606 (2010).
Finlay, D. & Cantrell, D. Phosphoinositide 3-kinase and the mammalian target of rapamycin pathways control T cell migration. Ann. NY Acad. Sci. 1183, 149–157 (2010).
Koch, M. A. et al. The transcription factor T-bet controls regulatory T cell homeostasis and function during type 1 inflammation. Nature Immunol. 10, 595–602 (2009).
Taqueti, V. R. et al. T-bet controls pathogenicity of CTLs in the heart by separable effects on migration and effector activity. J. Immunol. 177, 5890–5901 (2006).
Acknowledgements
I acknowledge the large number of researchers who have contributed to this field whose work was not cited owing to space limitations. I thank R. Wang and Y. Wang for critical comments on the manuscript, and members of my laboratory for helpful discussions. The author's research is supported by the US National Institutes of Health (K01 AR053573 and R01 NS064599), the National Multiple Sclerosis Society (RG4180-A-1), the Lupus Research Institute, the Cancer Research Institute and the American Lebanese Syrian Associated Charities.
Author information
Authors and Affiliations
Ethics declarations
Competing interests
The author declares no competing financial interests.
Related links
Glossary
- Metabolism
-
Intracellular chemical reactions that convert nutrients and endogenous molecules into energy and biomass (proteins, nucleic acids and lipids). Naive T cells have a catabolic metabolism through which they use glucose, fatty acids and amino acids for ATP generation via the tricarboxylic acid cycle and oxidative phosphorylation. On antigen stimulation, the bioenergetic demands of a T cell increase dramatically and the cells transition into anabolic metabolism mediated by glycolysis and glutaminolysis.
- AMP-activated protein kinase
-
(AMPK). A group of serine/threonine kinases that are activated in response to energy depletion. AMPK is an important activator of fatty acid oxidation and a potent inhibitor of mTORC1.
- Regulatory T cells
-
(TReg cells). A subset of CD4+ T cells that expresses FOXP3 and is crucial for the maintenance of immune tolerance. Although TReg cells mainly develop in the thymus as a separate lineage of CD4+ T cells, known as naturally occurring TReg cells, a second subset of TReg cells (known as induced TReg cells) arises de novo from conventional T cells in the periphery following antigen stimulation in the presence of TGFβ.
- Invariant natural killer T cells
-
(iNKT cells). A subset of immune cells that shares properties with both T cells and natural killer cells.
- Central tolerance
-
A process that eliminates self-reactive lymphocytes during their development. For T cells, this occurs mainly through clonal deletion in the thymus.
- Peripheral tolerance
-
A process that downregulates the activation of self-reactive T cells in secondary lymphoid organs. Two of the most important mechanisms are suppression by regulatory T cells and the induction of T cell anergy.
- Catabolic metabolism
-
The breakdown of complex substances into simpler ones, which is often accompanied by ATP production. Examples include the oxidation of fatty acids and amino acids.
- Oxidative phosphorylation
-
A metabolic pathway that produces ATP from the oxidation of nutrients and the transfer of electrons in a two-step process in mitochondria. The first reaction involves the conversion of intermediate molecules (pyruvate and fatty acids) into acetyl-CoA, which enters the tricarboxylic acid cycle, yielding free electrons that are carried by NADH and FADH2. In the second reaction, electrons from NADH and FADH2 are transferred to the electron-transport chain, resulting in the movement of protons out of the mitochondrial matrix and the generation of an electrochemical potential for ATP synthesis.
- Fatty acid oxidation
-
An important metabolic process used to derive energy through the mobilization and oxidation of fatty acids, mainly in the mitochondrial matrix. Fatty acid oxidation is positively and negatively regulated by AMPK and mTOR, respectively.
- T cell anergy
-
A state of T cell unresponsiveness to antigen stimulation in which T cells fail to proliferate and produce interleukin-2.
- Autophagy
-
A recycling process in which the cell degrades cytoplasmic organelles and proteins in lysosomes.
Rights and permissions
About this article
Cite this article
Chi, H. Regulation and function of mTOR signalling in T cell fate decisions. Nat Rev Immunol 12, 325–338 (2012). https://doi.org/10.1038/nri3198
Published:
Issue Date:
DOI: https://doi.org/10.1038/nri3198
This article is cited by
-
mTOR-mediated differentiation and maintenance of suppressive T cells at the center stage of IPEX treatment
Immunologic Research (2024)
-
Combined Immunodeficiency Caused by a Novel Nonsense Mutation in LCK
Journal of Clinical Immunology (2024)
-
The transcription factor GCN4 contributes to maintaining intracellular amino acid contents under nitrogen-limiting conditions in the mushroom Ganoderma lucidum
Microbial Cell Factories (2023)
-
Amino acid metabolism in immune cells: essential regulators of the effector functions, and promising opportunities to enhance cancer immunotherapy
Journal of Hematology & Oncology (2023)
-
Regulation of c-SMAC formation and AKT-mTOR signaling by the TSG101-IFT20 axis in CD4+ T cells
Cellular & Molecular Immunology (2023)