Key Points
-
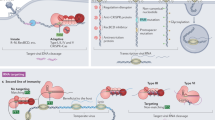
RNA-based antiviral immunity is active in diverse host species, which produce virus-derived small RNAs in infected cells and use them as specificity determinants to guide specific virus clearance.
-
Specific members of the Dicer family of proteins, which encode RNA helicase, double-stranded RNA (dsRNA)-binding and dsRNA-specific RNase domains, function as pattern recognition receptors (PRRs) in fungi, plants and invertebrates to detect viral dsRNA as a pathogen-associated molecular pattern (PAMP) and to further process it into virus-derived small interfering RNAs (siRNAs).
-
Viral siRNAs are structurally similar to host endogenous siRNAs with monophosphate groups at the 5′ end and 2′-O-methyl groups at the 3′ end. They guide specific clearance of the invading viral RNAs by members of the Argonaute (AGO) protein family.
-
Amplification of viral siRNAs by a host RNA-dependent RNA polymerase (RdRP) has an essential role in RNA-based antiviral immunity in plants. Distinct families of cellular RdRPs are conserved in all eukaryotes.
-
Recent deep sequencing has identified siRNA-like virus-derived small RNAs in mammalian cells infected with distinct RNA viruses. However, it is not clear whether these viral small RNAs function in RNA-based viral immunity or other aspects of viral immunity and pathogenesis.
-
Many nucleus-replicating DNA viruses that infect vertebrates and invertebrates encode up to 25 microRNAs (miRNAs) to regulate the expression of viral and host genes implicated in viral immunity and pathogenesis.
-
The recent discovery of virus-derived Piwi-interacting RNAs (piRNAs) in infected Drosophila melanogaster cells, which are larger than the 21-nucleotide viral siRNAs, suggests that piRNAs might have a novel antiviral function in addition to their role in genome defence against transposons and repeat elements.
Abstract
In eukaryotic RNA-based antiviral immunity, viral double-stranded RNA is recognized as a pathogen-associated molecular pattern and processed into small interfering RNAs (siRNAs) by the host ribonuclease Dicer. After amplification by host RNA-dependent RNA polymerases in some cases, these virus-derived siRNAs guide specific antiviral immunity through RNA interference and related RNA silencing effector mechanisms. Here, I review recent studies on the features of viral siRNAs and other virus-derived small RNAs from virus-infected fungi, plants, insects, nematodes and vertebrates and discuss the innate and adaptive properties of RNA-based antiviral immunity.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Takeuchi, O. & Akira, S. Innate immunity to virus infection. Immunol. Rev. 227, 75–81 (2009).
Franchi, L., Warner, N., Viani, K. & Nunez, G. Function of Nod-like receptors in microbial recognition and host defense. Immunol. Rev. 227, 106–128 (2009).
Yoneyama, M. & Fujita, T. RNA recognition and signal transduction by RIG-I-like receptors. Immunol. Rev. 227, 54–65 (2009).
Pfeffer, S. et al. Identification of virus-encoded microRNAs. Science 304, 734–736 (2004). This article reported the first evidence for virus-derived miRNAs.
Wu, Q. et al. Virus discovery by deep sequencing and assembly of virus-derived small silencing RNAs. Proc. Natl Acad. Sci. USA 107, 1606–1611 (2010). This paper reported the first evidence for virus-derived piRNAs in animals.
Kim, V. N., Han, J. & Siomi, M. C. Biogenesis of small RNAs in animals. Nature Rev. Mol. Cell Biol. 10, 126–139 (2009).
Carthew, R. W. & Sontheimer, E. J. Origins and mechanisms of miRNAs and siRNAs. Cell 136, 642–655 (2009).
Chen, X. Small RNAs and their roles in plant development. Annu. Rev. Cell Dev. Biol. 25, 21–44 (2009).
Hamilton, A. J. & Baulcombe, D. C. A species of small antisense RNA in posttranscriptional gene silencing in plants. Science 286, 950–952 (1999). References 9, 10 and 17 reported the first evidence for virus-derived siRNAs in plants, invertebrates and mammals, respectively.
Li, H. W., Li, W. X. & Ding, S. W. Induction and suppression of RNA silencing by an animal virus. Science 296, 1319–1321 (2002).
Zamore, P. D., Tuschl, T., Sharp, P. A. & Bartel, D. P. RNAi: double-stranded RNA directs the ATP-dependent cleavage of mRNA at 21 to 23 nucleotide intervals. Cell 101, 25–33 (2000).
Hammond, S. M., Bernstein, E., Beach, D. & Hannon, G. J. An RNA-directed nuclease mediates post-transcriptional gene silencing in Drosophila cells. Nature 404, 293–296 (2000).
Lindbo, J. A., Silva-Rosales, L., Proebsting, W. M. & Dougherty, W. G. Induction of a highly specific antiviral state in transgenic plants: implications for regulation of gene expression and virus resistance. Plant Cell 5, 1749–1759 (1993).
Ratcliff, F., Harrison, B. D. & Baulcombe, D. C. A similarity between viral defense and gene silencing in plants. Science 276, 1558–1560 (1997).
Hammond, S. M., Boettcher, S., Caudy, A. A., Kobayashi, R. & Hannon, G. J. Argonaute2, a link between genetic and biochemical analyses of RNAi. Science 293, 1146–1150 (2001).
Pfeffer, S. et al. Identification of microRNAs of the herpesvirus family. Nature Methods 2, 269–276 (2005).
Parameswaran, P. et al. Six RNA viruses and forty-one hosts: viral small RNAs and modulation of small RNA repertoires in vertebrate and invertebrate systems. PLoS Pathog. 6, e1000764 (2010).
Aliyari, R. et al. Mechanism of induction and suppression of antiviral immunity directed by virus-derived small RNAs in Drosophila. Cell Host Microbe 4, 387–397 (2008). Use of deep sequencing in this study led to identification of the first viral dsRNA precursor of virus-derived siRNAs.
Vanitharani, R., Chellappan, P. & Fauquet, C. M. Short interfering RNA-mediated interference of gene expression and viral DNA accumulation in cultured plant cells. Proc. Natl Acad. Sci. USA 100, 9632–9636 (2003).
Zhang, X., Segers, G. C., Sun, Q., Deng, F. & Nuss, D. L. Characterization of hypovirus-derived small RNAs generated in the chestnut blight fungus by an inducible DCL-2-dependent pathway. J. Virol. 82, 2613–2619 (2008).
Yoo, B. C. et al. A systemic small RNA signaling system in plants. Plant Cell 16, 1979–2000 (2004).
Molnar, A. et al. Plant virus-derived small interfering RNAs originate predominantly from highly structured single-stranded viral RNAs. J. Virol. 79, 7812–7818 (2005).
Ho, T., Pallett, D., Rusholme, R., Dalmay, T. & Wang, H. A simplified method for cloning of short interfering RNAs from Brassica juncea infected with Turnip mosaic potyvirus and Turnip crinkle carmovirus. J. Virol. Methods 136, 217–223 (2006).
Donaire, L. et al. Deep-sequencing of plant viral small RNAs reveals effective and widespread targeting of viral genomes. Virology 392, 203–214 (2009).
Myles, K. M., Wiley, M. R., Morazzani, E. M. & Adelman, Z. N. Alphavirus-derived small RNAs modulate pathogenesis in disease vector mosquitoes. Proc. Natl Acad. Sci. USA 105, 19938–19943 (2008).
Brackney, D. E., Beane, J. E. & Ebel, G. D. RNAi targeting of West Nile virus in mosquito midguts promotes virus diversification. PLoS Pathog. 5, e1000502 (2009).
Flynt, A., Liu, N., Martin, R. & Lai, E. C. Dicing of viral replication intermediates during silencing of latent Drosophila viruses. Proc. Natl Acad. Sci. USA 106, 5270–5275 (2009).
Qi, X., Bao, F. S. & Xie, Z. Small RNA deep sequencing reveals role for Arabidopsis thaliana RNA-dependent RNA polymerases in viral siRNA biogenesis. PLoS ONE 4, e4971 (2009).
Wang, X. B. et al. RNAi-mediated viral immunity requires amplification of virus-derived siRNAs in Arabidopsis thaliana. Proc. Natl Acad. Sci. USA 107, 484–489 (2010). References 29 and 40 established the role of viral secondary siRNAs in RNA-based antiviral immunity.
Szittya, G. et al. Structural and functional analysis of viral siRNAs. PLoS Pathog. 6, e1000838 (2010).
Blevins, T. et al. Four plant Dicers mediate viral small RNA biogenesis and DNA virus induced silencing. Nucleic Acids Res. 34, 6233–6246 (2006).
Akbergenov, R. et al. Molecular characterization of geminivirus-derived small RNAs in different plant species. Nucleic Acids Res. 34, 462–471 (2006).
Ebhardt, H. A., Thi, E. P., Wang, M. B. & Unrau, P. J. Extensive 3′ modification of plant small RNAs is modulated by helper component-proteinase expression. Proc. Natl Acad. Sci. USA 102, 13398–13403 (2005).
Lozsa, R., Csorba, T., Lakatos, L. & Burgyan, J. Inhibition of 3′ modification of small RNAs in virus-infected plants require spatial and temporal co-expression of small RNAs and viral silencing-suppressor proteins. Nucleic Acids Res. 36, 4099–4107 (2008).
Aliyari, R. & Ding, S. W. RNA-based viral immunity initiated by the Dicer family of host immune receptors. Immunol. Rev. 227, 176–188 (2009).
Kreuze, J. F. et al. Complete viral genome sequence and discovery of novel viruses by deep sequencing of small RNAs: a generic method for diagnosis, discovery and sequencing of viruses. Virology 388, 1–7 (2009).
Meyers, B. C. et al. Criteria for annotation of plant microRNAs. Plant Cell 20, 3186–3190 (2008).
Molnar, A. et al. Small silencing RNAs in plants are mobile and direct epigenetic modification in recipient cells. Science 328, 872–875 (2010).
Lu, R. et al. Animal virus replication and RNAi-mediated antiviral silencing in Caenorhabditis elegans. Nature 436, 1040–1043 (2005). References 39, 56 and 57 established an antiviral role for RNAi in C. elegans.
Garcia-Ruiz, H. et al. Arabidopsis RNA-dependent RNA polymerases and dicer-like proteins in antiviral defense and small interfering RNA biogenesis during turnip mosaic virus infection. Plant Cell 22, 481–496 (2010).
Lee, Y. S. et al. Distinct roles for Drosophila Dicer-1 and Dicer-2 in the siRNA/miRNA silencing pathways. Cell 117, 69–81 (2004).
Galiana-Arnoux, D., Dostert, C., Schneemann, A., Hoffmann, J. A. & Imler, J. L. Essential function in vivo for Dicer-2 in host defense against RNA viruses in Drosophila. Nature Immunol. 7, 590–597 (2006).
van Rij, R. P. et al. The RNA silencing endonuclease Argonaute 2 mediates specific antiviral immunity in Drosophila melanogaster. Genes Dev. 20, 2985–2995 (2006).
Wang, X. H. et al. RNA interference directs innate immunity against viruses in adult Drosophila. Science 312, 452–454 (2006). References 42, 43, 44 and 59 identified a key antiviral role for the dsRNA–siRNA pathway initiated by Dicer-2 in adult D. melanogaster.
Sabin, L. R. et al. Ars2 regulates both miRNA- and siRNA-dependent silencing and suppresses RNA virus infection in Drosophila. Cell 138, 340–351 (2009).
Saleh, M. C. et al. Antiviral immunity in Drosophila requires systemic RNA interference spread. Nature 458, 346–350 (2009).
Catalanotto, C. et al. Redundancy of the two dicer genes in transgene-induced posttranscriptional gene silencing in Neurospora crassa. Mol. Cell. Biol. 24, 2536–2545 (2004).
Voinnet, O. Origin, biogenesis, and activity of plant microRNAs. Cell 136, 669–687 (2009).
Bouche, N., Lauressergues, D., Gasciolli, V. & Vaucheret, H. An antagonistic function for Arabidopsis DCL2 in development and a new function for DCL4 in generating viral siRNAs. EMBO J. 25, 3347–3356 (2006).
Deleris, A. et al. Hierarchical action and inhibition of plant dicer-like proteins in antiviral defense. Science 313, 68–71 (2006).
Diaz-Pendon, J. A., Li, F., Li, W. X. & Ding, S. W. Suppression of antiviral silencing by cucumber mosaic virus 2b protein in Arabidopsis is associated with drastically reduced accumulation of three classes of viral small interfering RNAs. Plant Cell 19, 2053–2063 (2007).
Fusaro, A. F. et al. RNA interference-inducing hairpin RNAs in plants act through the viral defence pathway. EMBO Rep. 7, 1168–1175 (2006). References 49–52 identified a key antiviral role for the RNA silencing pathway initiated by DCL4 and DCL2 in A. thaliana plants.
Moissiard, G. & Voinnet, O. RNA silencing of host transcripts by cauliflower mosaic virus requires coordinated action of the four Arabidopsis Dicer-like proteins. Proc. Natl Acad. Sci. USA 103, 19593–19598 (2006).
Raja, P., Wolf, J. N. & Bisaro, D. M. RNA silencing directed against geminiviruses: Post-transcriptional and epigenetic components. Biochim. Biophys. Acta 1799, 337–351 (2010).
Lu, R., Yigit, E., Li, W. X. & Ding, S. W. An RIG-I-like RNA helicase mediates antiviral RNAi downstream of viral siRNA biogenesis in Caenorhabditis elegans. PLoS Pathog. 5, e1000286 (2009).
Schott, D. H., Cureton, D. K., Whelan, S. P. & Hunter, C. P. An antiviral role for the RNA interference machinery in Caenorhabditis elegans. Proc. Natl Acad. Sci. USA 102, 18420–18424 (2005).
Wilkins, C. et al. RNA interference is an antiviral defence mechanism in Caenorhabditis elegans. Nature 436, 1044–1047 (2005).
Habayeb, M. S., Ekstrom, J. O. & Hultmark, D. Nora virus persistent infections are not affected by the RNAi machinery. PLoS ONE 4, e5731 (2009).
Zambon, R. A., Vakharia, V. N. & Wu, L. P. RNAi is an antiviral immune response against a dsRNA virus in Drosophila melanogaster. Cell. Microbiol. 8, 880–889 (2006).
Deddouche, S. et al. The DExD/H-box helicase Dicer-2 mediates the induction of antiviral activity in Drosophila. Nature Immunol. 9, 1425–1432 (2008).
Tabara, H., Yigit, E., Siomi, H. & Mello, C. C. The dsRNA binding protein RDE-4 interacts with RDE-1, DCR-1, and a DExX-box helicase to direct RNAi in C. elegans. Cell 109, 861–871 (2002).
Morel, J. B. et al. Fertile hypomorphic argonaute (ago1) mutants impaired in post-transcriptional gene silencing and virus resistance. Plant Cell 14, 629–639 (2002).
Qu, F., Ye, X. & Morris, T. J. Arabidopsis DRB4, AGO1, AGO7, and RDR6 participate in a DCL4-initiated antiviral RNA silencing pathway negatively regulated by DCL1. Proc. Natl Acad. Sci. USA 105, 14732–14737 (2008).
Sun, Q., Choi, G. H. & Nuss, D. L. A single Argonaute gene is required for induction of RNA silencing antiviral defense and promotes viral RNA recombination. Proc. Natl Acad. Sci. USA 106, 17927–17932 (2009).
Ding, S. W. & Voinnet, O. Antiviral immunity directed by small RNAs. Cell 130, 413–426 (2007).
Czech, B. et al. An endogenous small interfering RNA pathway in Drosophila. Nature 453, 798–802 (2008).
Vaucheret, H. Plant ARGONAUTES. Trends Plant Sci. 13, 350–358 (2008).
Takeda, A., Iwasaki, S., Watanabe, T., Utsumi, M. & Watanabe, Y. The mechanism selecting the guide strand from small RNA duplexes is different among argonaute proteins. Plant Cell Physiol. 49, 493–500 (2008).
Zhang, X. et al. Cucumber mosaic virus-encoded 2b suppressor inhibits Arabidopsis Argonaute1 cleavage activity to counter plant defense. Genes Dev. 20, 3255–3268 (2006).
Chen, X. A microRNA as a translational repressor of apetala 2 in Arabidopsis flower development. Science 303, 2022–2025 (2004).
Brodersen, P. et al. Widespread translational inhibition by plant miRNAs and siRNAs. Science 320, 1185–1190 (2008).
Omarov, R. T., Ciomperlik, J. J. & Scholthof, H. B. RNAi-associated ssRNA-specific ribonucleases in Tombusvirus P19 mutant-infected plants and evidence for a discrete siRNA-containing effector complex. Proc. Natl Acad. Sci. USA 104, 1714–1719 (2007).
Pantaleo, V., Szittya, G. & Burgyan, J. Molecular bases of viral RNA targeting by viral small interfering RNA-programmed RISC. J. Virol. 81, 3797–3806 (2007).
Raja, P., Sanville, B. C., Buchmann, R. C. & Bisaro, D. M. Viral genome methylation as an epigenetic defense against geminiviruses. J. Virol. 82, 8997–9007 (2008).
Buchmann, R. C., Asad, S., Wolf, J. N., Mohannath, G. & Bisaro, D. M. Geminivirus AL2 and L2 proteins suppress transcriptional gene silencing and cause genome-wide reductions in cytosine methylation. J. Virol. 83, 5005–5013 (2009).
Seemanpillai, M., Dry, I., Randles, J. & Rezaian, A. Transcriptional silencing of geminiviral promoter-driven transgenes following homologous virus infection. Mol. Plant Microbe Interact. 16, 429–438 (2003).
Al Kaff, N. S. et al. Transcriptional and posttranscriptional plant gene silencing in response to a pathogen. Science 279, 2113–2115 (1998).
Rodriguez-Negrete, E. A., Carrillo-Tripp, J. & Rivera-Bustamante, R. F. RNA silencing against geminivirus: complementary action of posttranscriptional gene silencing and transcriptional gene silencing in host recovery. J. Virol. 83, 1332–1340 (2009).
Voinnet, O. Non-cell autonomous RNA silencing. FEBS Lett. 579, 5858–5871 (2005).
Molnar, A. et al. Small silencing RNAs in plants are mobile and direct epigenetic modification in recipient cells. Science 328, 872–875 (2010).
Dunoyer, P. et al. Small RNA duplexes function as mobile silencing signals between plant cells. Science 328, 912–916 (2010). References 80 and 81 demonstrated short- and long-distance transport of 21–24-nucleotide siRNAs in plant cells.
Lipardi, C. & Paterson, B. M. Identification of an RNA-dependent RNA polymerase in Drosophila involved in RNAi and transposon suppression. Proc. Natl Acad. Sci. USA 106, 15645–15650 (2009).
Mourrain, P. et al. Arabidopsis SGS2 and SGS3 genes are required for posttranscriptional gene silencing and natural virus resistance. Cell 101, 533–542 (2000).
Xie, Z., Fan, B., Chen, C. & Chen, Z. An important role of an inducible RNA-dependent RNA polymerase in plant antiviral defense. Proc. Natl Acad. Sci. USA 98, 6516–6521 (2001).
Schwach, F., Vaistij, F. E., Jones, L. & Baulcombe, D. C. An RNA-dependent RNA polymerase prevents meristem invasion by potato virus X and is required for the activity but not the production of a systemic silencing signal. Plant Physiol. 138, 1842–1852 (2005).
Yang, S. J., Carter, S. A., Cole, A. B., Cheng, N. H. & Nelson, R. S. A natural variant of a host RNA-dependent RNA polymerase is associated with increased susceptibility to viruses by Nicotiana benthamiana. Proc. Natl Acad. Sci. USA 101, 6297–6302 (2004).
Qu, F. et al. RDR6 has a broad-spectrum but temperature-dependent antiviral defense role in Nicotiana benthamiana. J. Virol. 79, 15209–15217 (2005).
Xie, Z. et al. Genetic and functional diversification of small RNA pathways in plants. PLoS Biol. 2, e104 (2004).
Donaire, L. et al. Structural and genetic requirements for the biogenesis of tobacco rattle virus-derived small interfering RNAs. J. Virol. 82, 5167–5177 (2008).
Vaistij, F. E. & Jones, L. Compromised virus-induced gene silencing in RDR6-deficient plants. Plant Physiol. 149, 1399–1407 (2009).
Yigit, E. et al. Analysis of the C. elegans Argonaute family reveals that distinct Argonautes act sequentially during RNAi. Cell 127, 747–757 (2006).
Li, F. & Ding, S. W. Virus counterdefense: diverse strategies for evading the RNA-silencing immunity. Annu. Rev. Microbiol. 60, 503–531 (2006).
Diaz-Pendon, J. A. & Ding, S. W. Direct and indirect roles of viral suppressors of RNA silencing in pathogenesis. Annu. Rev. Phytopathol. 46, 303–326 (2008).
Ruiz-Ferrer, V. & Voinnet, O. Roles of plant small RNAs in biotic stress responses. Annu. Rev. Plant Biol. 60, 485–510 (2009).
Fukunaga, R. & Doudna, J. A. dsRNA with 5′ overhangs contributes to endogenous and antiviral RNA silencing pathways in plants. EMBO J. 28, 545–555 (2009).
Elkashef, S. & Ding, S. W. Possible new RNA intermediate in RNA silencing. Nature Chem. Biol. 5, 278–279 (2009).
Glick, E. et al. Interaction with host SGS3 is required for suppression of RNA silencing by tomato yellow leaf curl virus V2 protein. Proc. Natl Acad. Sci. USA 105, 157–161 (2008).
Lakatos, L., Szittya, G., Silhavy, D. & Burgyan, J. Molecular mechanism of RNA silencing suppression mediated by p19 protein of tombusviruses. EMBO J. 23, 876–884 (2004).
Vogler, H. et al. Modification of small RNAs associated with suppression of RNA silencing by tobamovirus replicase protein. J. Virol. 81, 10379–10388 (2007).
Cuellar, W. J. et al. Elimination of antiviral defense by viral RNase III. Proc. Natl Acad. Sci. USA 106, 10354–10358 (2009).
Goto, K., Kobori, T., Kosaka, Y., Natsuaki, T. & Masuta, C. Characterization of silencing suppressor 2b of cucumber mosaic virus based on examination of its small RNA-binding abilities. Plant Cell Physiol. 48, 1050–1060 (2007).
Haas, G. et al. Nuclear import of CaMV P6 is required for infection and suppression of the RNA silencing factor DRB4. EMBO J. 27, 2102–2112 (2008).
Bortolamiol, D., Pazhouhandeh, M., Marrocco, K., Genschik, P. & Ziegler-Graff, V. The Polerovirus F box protein P0 targets ARGONAUTE1 to suppress RNA silencing. Curr. Biol. 17, 1615–1621 (2007).
Baumberger, N., Tsai, C. H., Lie, M., Havecker, E. & Baulcombe, D. C. The Polerovirus silencing suppressor P0 targets ARGONAUTE proteins for degradation. Curr. Biol. 17, 1609–1614 (2007).
Nayak, A. et al. Cricket paralysis virus antagonizes Argonaute 2 to modulate antiviral defense in Drosophila. Nature Struct. Mol. Biol. 17, 547–554 (2010).
Szittya, G., Molnar, A., Silhavy, D., Hornyik, C. & Burgyan, J. Short defective interfering RNAs of tombusviruses are not targeted but trigger post-transcriptional gene silencing against their helper virus. Plant Cell 14, 359–372 (2002).
Hassani-Mehraban, A., Brenkman, A. B., van den Broek, N. J., Goldbach, R. & Kormelink, R. RNAi-mediated transgenic Tospovirus resistance broken by intraspecies silencing suppressor protein complementation. Mol. Plant Microbe Interact. 22, 1250–1257 (2009).
Chellappan, P., Vanitharani, R., Pita, J. & Fauquet, C. M. Short interfering RNA accumulation correlates with host recovery in DNA virus-infected hosts, and gene silencing targets specific viral sequences. J. Virol. 78, 7465–7477 (2004).
Umbach, J. L. et al. MicroRNAs expressed by herpes simplex virus 1 during latent infection regulate viral mRNAs. Nature 454, 780–783 (2008).
Jurak, I. et al. Numerous conserved and divergent microRNAs expressed by herpes simplex viruses 1 and 2. J. Virol. 84, 4659–4672 (2010).
Buck, A. H. et al. Discrete clusters of virus-encoded microRNAs are associated with complementary strands of the genome and the 7.2-kilobase stable intron in murine cytomegalovirus. J. Virol. 81, 13761–13770 (2007).
Dolken, L. et al. Mouse cytomegalovirus microRNAs dominate the cellular small RNA profile during lytic infection and show features of posttranscriptional regulation. J. Virol. 81, 13771–13782 (2007).
Umbach, J. L. & Cullen, B. R. The role of RNAi and microRNAs in animal virus replication and antiviral immunity. Genes Dev. 23, 1151–1164 (2009).
Sullivan, C. S., Grundhoff, A. T., Tevethia, S., Pipas, J. M. & Ganem, D. SV40-encoded microRNAs regulate viral gene expression and reduce susceptibility to cytotoxic T cells. Nature 435, 682–686 (2005).
Sullivan, C. S. et al. Murine Polyomavirus encodes a microRNA that cleaves early RNA transcripts but is not essential for experimental infection. Virology 387, 157–167 (2009).
Hussain, M., Taft, R. J. & Asgari, S. An insect virus-encoded microRNA regulates viral replication. J. Virol. 82, 9164–9170 (2008).
Wu, Q. F. Wang, X. B. and Ding, S. W. Viral suppressors of RNA-based viral immunity: host targets. Cell Host Microbe 8, 12–15 (2010).
Acknowledgements
Work in my laboratory is supported by grants from the US National Institutes of Health, the United States Department of Agriculture, California Citrus Research Board and UC Discovery. Because of space limitations I have often cited reviews rather than primary research papers. I apologize to those investigators whose original papers have not been cited.
Author information
Authors and Affiliations
Ethics declarations
Competing interests
The author declares no competing financial interests.
Supplementary information
Supplementary information S1 (box)
miRNAs and piRNAs in antiviral immunity (PDF 183 kb)
Related links
Glossary
- Pathogen-associated molecular patterns
-
(PAMPs). Molecular patterns that are found in pathogens but not mammalian cells. Examples include various microbial products, such as bacterial lipopolysaccharides, hypomethylated DNA, flagellin and double-stranded RNA, which bind to Toll-like receptors.
- Pattern recognition receptors
-
(PRRs). Host receptors (such as Toll-like receptors (TLRs), NOD-like receptors (NLRs) or RIG-I-like receptors (RLRs)) that can sense pathogen-associated molecular patterns and initiate signalling cascades that lead to an innate immune response. These can be membrane bound (such as TLRs) or soluble cytoplasmic receptors (such as RIG-I, melanoma differentiation-associated gene 5 (MDA5) and NLRs).
- Small interfering RNAs
-
(siRNAs). 21–24-nucleotide double-stranded RNAs with two-nucleotide 3′ overhangs and 5′-monophosphate and 3′-hydroxyl termini. They are processed from long double-stranded RNA precursors by Dicer. Plant and animal genomes encode many siRNAs with complete or extensive sequence complementarity to endogenous mRNA transcripts.
- MicroRNAs
-
(miRNAs). 21–25-nucleotide single-stranded RNAs with 5′-monophosphate and 3′-hydroxyl termini. They are processed by Dicer from a structured region of single-stranded nuclear transcripts. The seed region of miRNAs corresponds to nucleotides 2–8 and seed pairing has a crucial role in the recognition of the target mRNA.
- Piwi-interacting RNAs
-
(piRNAs). 24–31-nucleotide single-stranded RNAs with 5′-monophosphate and 3′-hydroxyl termini. They are independent of Dicer for biogenesis, bind to the Piwi subfamily of Argonaute (AGO) proteins for function and are found in animals but not in plants, possibly because plants do not encode any AGO protein in the Piwi subfamily.
- RNA interference
-
(RNAi). Specific gene silencing that is induced by long double-stranded RNA or small interfering RNAs. It is used widely to knock down gene expression in plants and animals.
- RNA silencing
-
Specific gene silencing guided by all classes of small silencing RNAs such as siRNAs, miRNAs and piRNAs.
- Piwi domain
-
The highly conserved carboxy-terminal domain of Argonaute (AGO) proteins, which contains an RNase H motif. The catalytic centre consists of a DDH triad that functions as a metal coordinating site. AGO binding to a target RNA that is highly complementary to the loaded small interfering RNA brings the scissile phosphate, opposite nucleotides 10 and 11 of the small RNA guide, into the enzyme active site, allowing cleavage of the target RNA to leave 5′-monophosphate and 3′-hydroxyl termini.
- RNA-induced silencing complex
-
(RISC). The effector complex of RNA silencing that contains at least two components: a single-stranded small interfering RNA or microRNA and an AGO protein.
- Positive-strand RNA virus
-
((+)RNA virus). These viruses use RNA as genetic material. Their virions contain single-stranded genomic RNA that functions directly as mRNA and is sufficient to initiate viral infection after entry into a host cell. Tobacco mosaic virus, poliovirus and hepatitis C virus are examples.
- Subgenomic RNA
-
RNA transcripts of the viral RNA genome that contain only part of the sequence present in the entire genome and usually function as mRNA.
- Deep sequencing
-
Also referred to as next-generation sequencing. This includes the Roche 454, Illumina and other high-throughput DNA sequencing platforms.
- dsRNA uptake pathway
-
The endocytic pathway that mediates cell entry of double-stranded RNA in insect cells.
- Systemic silencing
-
RNA silencing that occurs in tissues distant from the site where RNA silencing is initially induced, as a result of non-cell autonomous spread of RNA silencing in plants and Caenorhabditis elegans.
- Secondary siRNAs
-
Small interfering RNAs (siRNAs) produced by processes that require a cellular RNA-dependent RNA polymerase, in contrast to primary siRNAs that are diced from exogenous double-stranded RNA (dsRNA) or dsRNA synthesized by viral RNA-dependent RNA polymerase.
- Satellite RNA
-
Non-coding linear or circular RNA molecules of a few hundred nucleotides in length that are replicated and packaged into virions by a 'helper virus', but have no significant sequence homology with the 'helper virus' genome. Different strains of satellite RNA may attenuate or intensify the disease symptoms induced by the helper virus.
Rights and permissions
About this article
Cite this article
Ding, SW. RNA-based antiviral immunity. Nat Rev Immunol 10, 632–644 (2010). https://doi.org/10.1038/nri2824
Published:
Issue Date:
DOI: https://doi.org/10.1038/nri2824
This article is cited by
-
Reduced insulin/IGF-1 signalling upregulates two anti-viral immune pathways, decreases viral load and increases survival under viral infection in C. elegans
GeroScience (2024)
-
Small interfering RNA (siRNA)-based therapeutic applications against viruses: principles, potential, and challenges
Journal of Biomedical Science (2023)
-
Identification of ceRNA-vsiRNA-mRNA network for exploring the mechanism underlying pathogenesis of sugarcane mosaic virus in resistant and susceptible maize inbred lines
Phytopathology Research (2023)
-
Nationwide genomic surveillance reveals the prevalence and evolution of honeybee viruses in China
Microbiome (2023)
-
3′ UTR is critical for viral RNA accumulation of jasmine virus H
Phytopathology Research (2023)