Key Points
-
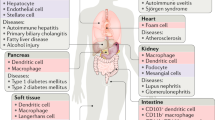
The clearance of apoptotic cells occurs throughout life in multi-cellular organisms; the failure to properly engulf and remove these dying cells has been linked to a break in self tolerance and autoimmunity.
-
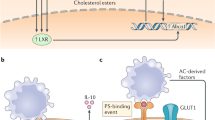
Recent evidence suggests that phagocytes sense the presence of apoptotic cells at the earliest stages of programmed cell death. Some of this sensing involves the secretion of soluble factors that help to recruit phagocytic cells (such as monocytes) to the dying cells. Such prompt removal of apoptotic cells is necessary before they lose their membrane integrity in the later stages of apoptosis, and potentially release their intracellular contents.
-
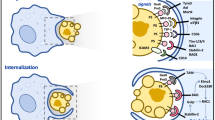
Phosphatidylserine (PtdSer) is one of the key 'eat-me' signals that is exposed on the surface of apoptotic cells. The exposed PtdSer can be recognized by bridging molecules that can in turn be bound by phagocytic receptors, or by receptors that can directly engage PtdSer.
-
Although the identity of phagocyte receptors that can directly engage PtdSer has remained controversial, recent studies have begun to resolve the conflicting issues and identify new PtdSer recognition receptors.
-
With respect to intracellular signalling within the phagocyte, the RAS homologue (RHO)-family GTPase RAC has a crucial role in the reorganization of the cytoskeleton during the engulfment of apoptotic cells.
-
Upon recognition of apoptotic targets, the intracellular signalling proteins DOCK180 and ELMO1 (engulfment and cell motility 1) function together to activate RAC within the phagocyte and thereby facilitate the uptake of apoptotic cells. The signalling pathway involving DOCK180–ELMO1–RAC that facilitates the uptake of apoptotic cells is evolutionarily conserved from worms to humans.
-
One of the hallmarks of the engulfment of apoptotic cells is that such uptake is immunologically silent, and the phagocytes that engulf apoptotic cells also produce anti-inflammatory cytokines. This is in contrast to the uptake of bacteria or other pathogens, which readily induces an inflammatory response. Defining the intracellular mechanisms within the phagocyte that are induced by the recognition of apoptotic cells that, in turn, inhibit inflammatory signalling, is one of the key areas of ongoing research.
-
When a phagocyte engulfs one or more apoptotic cells, it essentially doubles its cellular contents, including phospholipids, cholesterol, ATP and so on. How the phagocyte maintains homeostasis is beginning to be addressed. Early studies suggest a crucial role for the PtdSer exposed on apoptotic cells in regulating cholesterol efflux from the engulfing macrophages.
-
Future studies will probably focus on how the phagocyte responses are regulated and whether the mechanisms of such apoptotic-cell-mediated immunosuppression could be harnessed for treating chronic inflammatory diseases and autoimmunity.
Abstract
The clearance of apoptotic cells by phagocytes is an integral component of normal life, and defects in this process can have significant implications for self tolerance and autoimmunity. Recent studies have provided new insights into the engulfment process, including how phagocytes seek apoptotic cells, how they recognize and ingest these targets and how they maintain cellular homeostasis after the 'meal'. Several new factors that regulate engulfment have been identified, whereas the roles of some of the older players require revision. This Review focuses on these recent developments and attempts to highlight some of the important questions in this field.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Henson, P. M. & Hume, D. A. Apoptotic cell removal in development and tissue homeostasis. Trends Immunol. 27, 244–250 (2006).
Finnemann, S. C., Bonilha, V. L., Marmorstein, A. D. & Rodriguez-Boulan, E. Phagocytosis of rod outer segments by retinal pigment epithelial cells requires αvβ5 integrin for binding but not for internalization. Proc. Natl Acad. Sci. USA 94, 12932–12937 (1997).
Wood, W. et al. Mesenchymal cells engulf and clear apoptotic footplate cells in macrophageless PU.1 null mouse embryos. Development 127, 5245–5252 (2000).
Gregory, C. D. & Devitt, A. The macrophage and the apoptotic cell: an innate immune interaction viewed simplistically? Immunology 113, 1–14 (2004).
Aderem, A. & Underhill, D. M. Mechanisms of phagocytosis in macrophages. Annu. Rev. Immunol. 17, 593–623 (1999).
Savill, J. & Fadok, V. Corpse clearance defines the meaning of cell death. Nature 407, 784–788 (2000).
Lauber, K., Blumenthal, S. G., Waibel, M. & Wesselborg, S. Clearance of apoptotic cells: getting rid of the corpses. Mol. Cell 14, 277–287 (2004).
Henson, P. M. Dampening inflammation. Nature Immunol. 6, 1179–1181 (2005).
Vandivier, R. W., Henson, P. M. & Douglas, I. S. Burying the dead: the impact of failed apoptotic cell removal (efferocytosis) on chronic inflammatory lung disease. Chest 129, 1673–1682 (2006).
Hoeppner, D. J., Hengartner, M. O. & Schnabel, R. Engulfment genes cooperate with ced-3 to promote cell death in Caenorhabditis elegans. Nature 412, 202–206 (2001).
Reddien, P. W., Cameron, S. & Horvitz, H. R. Phagocytosis promotes programmed cell death in C. elegans. Nature 412, 198–202 (2001). References 10 and 11 showed for the first time that the initiation of apoptosis in a given cell and its subsequent engulfment are intimately linked such that the failure to clear apoptotic cells arrested progression through the steps of programmed cell death.
Ravichandran, K. S. “Recruitment signals” from apoptotic cells: invitation to a quiet meal. Cell 113, 817–820 (2003).
Lauber, K. et al. Apoptotic cells induce migration of phagocytes via caspase-3-mediated release of a lipid attraction signal. Cell 113, 717–730 (2003). Using MCF7 cells, with or without caspase-3 expression, and transwell migration assays, this paper identified LPC as a find-me signal that is released from apoptotic cells and can attract monocytes.
Drobnik, W. et al. Plasma ceramide and lysophosphatidylcholine inversely correlate with mortality in sepsis patients. J. Lipid Res. 44, 754–761 (2003).
Koizumi, S. et al. UDP acting at P2Y6 receptors is a mediator of microglial phagocytosis. Nature 446, 1091–1095 (2007).
Scannell, M. et al. Annexin-1 and peptide derivatives are released by apoptotic cells and stimulate phagocytosis of apoptotic neutrophils by macrophages. J. Immunol. 178, 4595–4605 (2007).
Maderna, P., Yona, S., Perretti, M. & Godson, C. Modulation of phagocytosis of apoptotic neutrophils by supernatant from dexamethasone-treated macrophages and annexin-derived peptide Ac(2–26). J. Immunol. 174, 3727–3733 (2005).
Henson, P. M., Bratton, D. L. & Fadok, V. A. Apoptotic cell removal. Curr. Biol. 11, R795–R805 (2001).
Savill, J., Dransfield, I., Gregory, C. & Haslett, C. A blast from the past: clearance of apoptotic cells regulates immune responses. Nature Rev. Immunol. 2, 965–975 (2002).
Fadok, V. A., Bratton, D. L., Frasch, S. C., Warner, M. L. & Henson, P. M. The role of phosphatidylserine in recognition of apoptotic cells by phagocytes. Cell Death Differ. 5, 551–562 (1998).
Devitt, A. et al. Human CD14 mediates recognition and phagocytosis of apoptotic cells. Nature 392, 505–509 (1998).
Fadok, V. A., Warner, M. L., Bratton, D. L. & Henson, P. M. CD36 is required for phagocytosis of apoptotic cells by human macrophages that use either a phosphatidylserine receptor or the vitronectin receptor (αvβ3). J. Immunol. 161, 6250–6257 (1998).
Schlegel, R. A., Krahling, S., Callahan, M. K. & Williamson, P. CD14 is a component of multiple recognition systems used by macrophages to phagocytose apoptotic lymphocytes. Cell Death Differ. 6, 583–592 (1999).
Ezekowitz, R. A., Sastry, K., Bailly, P. & Warner, A. Molecular characterization of the human macrophage mannose receptor: demonstration of multiple carbohydrate recognition-like domains and phagocytosis of yeasts in Cos-1 cells. J. Exp. Med. 172, 1785–1794 (1990).
Savill, J., Dransfield, I., Hogg, N. & Haslett, C. Vitronectin receptor-mediated phagocytosis of cells undergoing apoptosis. Nature 343, 170–173 (1990).
Gordon, S. Macrophage-restricted molecules: role in differentiation and activation. Immunol. Lett. 65, 5–8 (1999).
Gregory, C. D., Devitt, A. & Moffatt, O. Roles of ICAM-3 and CD14 in the recognition and phagocytosis of apoptotic cells by macrophages. Biochem. Soc. Trans. 26, 644–649 (1998).
Gardai, S. J. et al. Cell-surface calreticulin initiates clearance of viable or apoptotic cells through trans-activation of LRP on the phagocyte. Cell 123, 321–334 (2005).
Ogden, C. A. et al. C1q and mannose binding lectin engagement of cell surface calreticulin and CD91 initiates macropinocytosis and uptake of apoptotic cells. J. Exp. Med. 194, 781–795 (2001). References 28 and 29 describe a role for calreticulin in decorating the surface of apoptotic cells and the subsequent recognition by CD91 (also known as LRP1). Reference 28 also describes the patchy appearance of exposed PtdSer on apoptotic cells, and the colocalization of calreticulin in similar patches.
Fadok, V. A. et al. Different populations of macrophages use either the vitronectin receptor or the phosphatidylserine receptor to recognize and remove apoptotic cells. J. Immunol. 149, 4029–4035 (1992).
Pradhan, D., Krahling, S., Williamson, P. & Schlegel, R. A. Multiple systems for recognition of apoptotic lymphocytes by macrophages. Mol. Biol. Cell 8, 767–778 (1997).
Fadok, V. A. et al. Exposure of phosphatidylserine on the surface of apoptotic lymphocytes triggers specific recognition and removal by macrophages. J. Immunol. 148, 2207–2216 (1992). This seminal paper identified PtdSer as a key eat-me signal exposed on apoptotic cells.
Anderson, H. A. et al. Serum-derived protein S binds to phosphatidylserine and stimulates the phagocytosis of apoptotic cells. Nature Immunol. 4, 87–91 (2003).
Nagata, K. et al. Identification of the product of growth arrest-specific gene 6 as a common ligand for Axl, Sky, and Mer receptor tyrosine kinases. J. Biol. Chem. 271, 30022–30027 (1996).
Hanayama, R. et al. Identification of a factor that links apoptotic cells to phagocytes. Nature 417, 182–187 (2002). This paper identified MFGE8 as a bridging molecule that can bind PtdSer on apoptotic cells and link it to α v β 3 -integrin on phagocytes.
Hanayama, R. et al. Autoimmune disease and impaired uptake of apoptotic cells in MFG-E8-deficient mice. Science 304, 1147–1150 (2004).
Arur, S. et al. Annexin I is an endogenous ligand that mediates apoptotic cell engulfment. Dev. Cell 4, 587–598 (2003).
Fadok, V. A., McDonald, P. P., Bratton, D. L. & Henson, P. M. Regulation of macrophage cytokine production by phagocytosis of apoptotic and post-apoptotic cells. Biochem. Soc. Trans. 26, 653–656 (1998).
Fadok, V. A. et al. A receptor for phosphatidylserine-specific clearance of apoptotic cells. Nature 405, 85–90 (2000).
Fadok, V. A. et al. Macrophages that have ingested apoptotic cells in vitro inhibit proinflammatory cytokine production through autocrine/paracrine mechanisms involving TGF-β, PGE2, and PAF. J. Clin. Invest. 101, 890–898 (1998).
Gallucci, S., Lolkema, M. & Matzinger, P. Natural adjuvants: endogenous activators of dendritic cells. Nature Med. 5, 1249–1255 (1999).
Sauter, B. et al. Consequences of cell death: exposure to necrotic tumor cells, but not primary tissue cells or apoptotic cells, induces the maturation of immunostimulatory dendritic cells. J. Exp. Med. 191, 423–434 (2000). References 41 and 42 suggested that necrotic cells promote the maturation of immature DCs, whereas the engulfment of apoptotic cells does not
Greenberg, M. E. et al. Oxidized phosphatidylserine-CD36 interactions play an essential role in macrophage-dependent phagocytosis of apoptotic cells. J. Exp. Med. 203, 2613–2625 (2006).
Kagan, V. E. et al. A role for oxidative stress in apoptosis: oxidation and externalization of phosphatidylserine is required for macrophage clearance of cells undergoing Fas-mediated apoptosis. J. Immunol. 169, 487–499 (2002). This report demonstrated that some of the PtdSer that is exposed on apoptotic cells can exist in an oxidized state.
Kagan, V. E. et al. Oxidative lipidomics of apoptosis: redox catalytic interactions of cytochrome c with cardiolipin and phosphatidylserine. Free Radic. Biol. Med. 37, 1963–1985 (2004).
Bose, J. et al. The phosphatidylserine receptor has essential functions during embryogenesis but not in apoptotic cell removal. J. Biol. 3, 21 (2004). This seminal paper, through the knockout of the putative Ptdsr1 gene, revealed the lack of an obvious role for the gene product of Ptdsr1 in the engulfment of apoptotic cells and demonstrated that the Ptdsr1 gene does not encode the putative receptor for PtdSer.
Cikala, M. et al. The phosphatidylserine receptor from Hydra is a nuclear protein with potential Fe(II) dependent oxygenase activity. BMC Cell Biol. 5, 26 (2004).
Mitchell, J. E. et al. The presumptive phosphatidylserine receptor is dispensable for innate anti-inflammatory recognition and clearance of apoptotic cells. J. Biol. Chem. 281, 5718–5725 (2006).
Kunisaki, Y. et al. Defective fetal liver erythropoiesis and T lymphopoiesis in mice lacking the phosphatidylserine receptor. Blood 103, 3362–3364 (2004).
Li, M. O., Sarkisian, M. R., Mehal, W. Z., Rakic, P. & Flavell, R. A. Phosphatidylserine receptor is required for clearance of apoptotic cells. Science 302, 1560–1563 (2003). In this paper, the authors knocked out the putative Ptdsr1 gene and reported a defect in apoptotic-cell clearance. However, it now appears that the intended knockout of the Ptdsr1 gene may have also deleted the expression of a neighbouring gene.
Gardai, S. J., Bratton, D. L., Ogden, C. A. & Henson, P. M. Recognition ligands on apoptotic cells: a perspective. J. Leukoc. Biol. 79, 896–903 (2006).
Park, D. et al. BAI1 is an engulfment receptor for apoptotic cells upstream of the ELMO/Dock180/Rac module. Nature 24 October 2007 (doi:10.1038/nature06329).
Wu, Y., Tibrewal, N. & Birge, R. B. Phosphatidylserine recognition by phagocytes: a view to a kill. Trends Cell Biol. 16, 189–197 (2006).
Kiss, R. S., Elliott, M. R., Ma, Z., Marcel, Y. L. & Ravichandran, K. S. Apoptotic cells induce a phosphatidylserine-dependent homeostatic response from phagocytes. Curr. Biol. 16, 2252–2258 (2006). This paper demonstrated a key role for PtdSer that is exposed on apoptotic cells in stimulating cholesterol efflux from the phagocytes that come in contact with apoptotic cells.
Chimini, G. Apoptosis: repulsive encounters. Nature 418, 139–141 (2002).
Brown, S. et al. Apoptosis disables CD31-mediated cell detachment from phagocytes promoting binding and engulfment. Nature 418, 200–203 (2002). This work identified CD31 as one type of signal (loosely termed a 'don't-eat-me' signal) that prevents the engulfment of live cells by phagocytes.
Ellis, R. E., Jacobson, D. M. & Horvitz, H. R. Genes required for the engulfment of cell corpses during programmed cell death in Caenorhabditis elegans. Genetics 129, 79–94 (1991).
Gumienny, T. L. & Hengartner, M. O. How the worm removes corpses: the nematode C. elegans as a model system to study engulfment. Cell Death Differ. 8, 564–568 (2001).
Kinchen, J. M. & Hengartner, M. O. Tales of cannibalism, suicide, and murder: Programmed cell death in C. elegans. Curr. Top. Dev. Biol. 65, 1–45 (2005).
Reddien, P. W. & Horvitz, H. R. CED-2/CrkII and CED-10/Rac control phagocytosis and cell migration in Caenorhabditis elegans. Nature Cell Biol. 2, 131–136 (2000).
Wu, Y. C. & Horvitz, H. R. The C. elegans cell corpse engulfment gene ced-7 encodes a protein similar to ABC transporters. Cell 93, 951–960 (1998).
Wu, Y. C. & Horvitz, H. R. C. elegans phagocytosis and cell-migration protein CED-5 is similar to human DOCK180. Nature 392, 501–504 (1998).
Wu, Y. C., Tsai, M. C., Cheng, L. C., Chou, C. J. & Weng, N. Y. C. elegans CED-12 acts in the conserved CrkII/DOCK180/Rac pathway to control cell migration and cell corpse engulfment. Dev. Cell 1, 491–502 (2001).
Zhou, Z., Caron, E., Hartwieg, E., Hall, A. & Horvitz, H. R. The C. elegans PH domain protein CED-12 regulates cytoskeletal reorganization via a Rho/Rac GTPase signaling pathway. Dev. Cell 1, 477–489 (2001).
Zhou, Z., Hartwieg, E. & Horvitz, H. R. CED-1 is a transmembrane receptor that mediates cell corpse engulfment in C. elegans. Cell 104, 43–56 (2001).
Yu, X., Odera, S., Chuang, C. H., Lu, N. & Zhou, Z. C. elegans dynamin mediates the signaling of phagocytic receptor CED-1 for the engulfment and degradation of apoptotic cells. Dev. Cell 10, 743–757 (2006).
Ridley, A. J. Rho family proteins: coordinating cell responses. Trends Cell Biol. 11, 471–477 (2001).
Erwig, L. P. et al. Differential regulation of phagosome maturation in macrophages and dendritic cells mediated by Rho GTPases and ezrin-radixin-moesin (ERM) proteins. Proc. Natl Acad. Sci. USA 103, 12825–12830 (2006). This paper revealed some of the steps that are involved in the maturation of phagosomes that contain apoptotic cells, and a role for RHO proteins.
Nakaya, M., Tanaka, M., Okabe, Y., Hanayama, R. & Nagata, S. Opposite effects of rho family GTPases on engulfment of apoptotic cells by macrophages. J. Biol. Chem. 281, 8836–8842 (2006).
Tosello-Trampont, A. C., Nakada-Tsukui, K. & Ravichandran, K. S. Engulfment of apoptotic cells is negatively regulated by Rho-mediated signaling. J. Biol. Chem. 278, 49911–49919 (2003).
Riento, K. & Ridley, A. J. Rocks: multifunctional kinases in cell behaviour. Nature Rev. Mol. Cell Biol. 4, 446–456 (2003).
Olazabal, I. et al. Rho-kinase and myosin-II control phagocytic cup formation during CR, but not FcγR, phagocytosis. Curr. Biol. 12, 1413–1418 (2002).
Hoffmann, P. R. et al. Phosphatidylserine (PS) induces PS receptor-mediated macropinocytosis and promotes clearance of apoptotic cells. J. Cell Biol. 155, 649–660 (2001).
Albert, M., Kim, J. & Birge, R. αvβ5 integrin recruits the CrkII–Dock180–Rac1 complex for phagocytosis of apoptotic cells. Nature Cell Biol. 2, 899–905 (2000).
Leverrier, Y. & Ridley, A. J. Requirement for Rho GTPases and PI-3 kinase during apoptotic cell phagocytosis by macrophages. Curr. Biol. 11, 196–199 (2001).
Tosello-Trampont, A. C. et al. Identification of two signaling submodules within the CrkII/ELMO/Dock180 pathway regulating engulfment of apoptotic cells. Cell Death Differ. 14, 963–972 (2007).
Leverrier, Y. et al. Cutting edge: the Wiskott–Aldrich syndrome protein is required for efficient phagocytosis of apoptotic cells. J. Immunol. 166, 4831–4834 (2001).
Lu, M. & Ravichandran, K. S. Dock180–ELMO cooperation in Rac activation. Methods Enzymol. 406, 388–402 (2006).
Gumienny, T. L. et al. CED-12/ELMO, a novel member of the CrkII/Dock180/Rac pathway, is required for phagocytosis and cell migration. Cell 107, 27–41 (2001). References 63, 64 and 79 cloned and identified CED-12, the C. elegans homologue of ELMO1, as being involved in engulfment and placed the ced-12 gene upstream of RAC1.
Tosello-Trampont, A. C., Brugnera, E. & Ravichandran, K. S. Evidence for a conserved role for CRKII and Rac in engulfment of apoptotic cells. J. Biol. Chem. 276, 13797–13802 (2001).
Hasegawa, H. et al. DOCK180, a major CRK-binding protein, alters cell morphology upon translocation to the cell membrane. Mol. Cell Biol. 16, 1770–1776 (1996).
Brugnera, E. et al. Unconventional Rac-GEF activity is mediated through the Dock180–ELMO complex. Nature Cell Biol. 4, 574–582 (2002). This paper identified the new RAC-GEF domain within the DOCK180 family of proteins and the regulation of the nucleotide-exchange activity by ELMO1 during the engulfment of apoptotic cells.
Grimsley, C. M. et al. Dock180 and ELMO1 proteins cooperate to promote evolutionarily conserved Rac-dependent cell migration. J. Biol. Chem. 279, 6087–6097 (2004).
deBakker, C. D. et al. Phagocytosis of apoptotic cells is regulated by a UNC-73/TRIO-MIG-2/RhoG signaling module and armadillo repeats of CED-12/ELMO. Curr. Biol. 14, 2208–2216 (2004).
Cote, J. F. & Vuori, K. Identification of an evolutionarily conserved superfamily of DOCK180-related proteins with guanine nucleotide exchange activity. J. Cell Sci. 115, 4901–4913 (2002).
Meller, N., Irani-Tehrani, M., Kiosses, W. B., Del Pozo, M. A. & Schwartz, M. A. Zizimin1, a novel Cdc42 activator, reveals a new GEF domain for Rho proteins. Nature Cell Biol. 4, 639–647 (2002).
Meller, N., Merlot, S. & Guda, C. CZH proteins: a new family of Rho-GEFs. J. Cell Sci. 118, 4937–4946 (2005).
Lu, M. et al. A steric-inhibition model for regulation of nucleotide exchange via the Dock180 family of GEFs. Curr. Biol. 15, 371–377 (2005).
Lu, M. et al. PH domain of ELMO functions in trans to regulate Rac activation via Dock180. Nature Struct. Mol. Biol. 11, 756–762 (2004).
Blangy, A. et al. TrioGEF1 controls Rac- and Cdc42-dependent cell structures through the direct activation of rhoG. J. Cell Sci. 113, 729–739 (2000).
Katoh, H. & Negishi, M. RhoG activates Rac1 by direct interaction with the Dock180-binding protein Elmo. Nature 424, 461–464 (2003).
Grimsley, C. M., Lu, M., Haney, L. B., Kinchen, J. M. & Ravichandran, K. S. Characterization of a novel interaction between ELMO1 and ERM proteins. J. Biol. Chem. 281, 5928–5937 (2006).
Silva, E., Au-Yeung, H. W., Van Goethem, E., Burden, J. & Franc, N. C. Requirement for a Drosophila E3-ubiquitin ligase in phagocytosis of apoptotic cells. Immunity 27, 585–596 (2007). This paper identifies several new players in the engulfment of apoptotic cells by macrophages in a D. melanogaster model system, and also describes the possibility that the ubiquitylation of engulfed targets influences subsequent engulfment.
Makino, Y. et al. Elmo1 inhibits ubiquitylation of Dock180. J. Cell Sci. 119, 923–932 (2006).
Scott, R. S. et al. Phagocytosis and clearance of apoptotic cells is mediated by MER. Nature 411, 207–211 (2001).
Botto, M. et al. Homozygous C1q deficiency causes glomerulonephritis associated with multiple apoptotic bodies. Nature Genet. 19, 56–59 (1998).
Franz, S. et al. Apoptosis and autoimmunity: when apoptotic cells break their silence. Curr. Rheumatol. Rep. 8, 245–247 (2006).
Asano, K. et al. Masking of phosphatidylserine inhibits apoptotic cell engulfment and induces autoantibody production in mice. J. Exp. Med. 200, 459–467 (2004).
Bondanza, A. et al. Inhibition of phosphatidylserine recognition heightens the immunogenicity of irradiated lymphoma cells in vivo. J. Exp. Med. 200, 1157–1165 (2004).
Devitt, A. et al. Persistence of apoptotic cells without autoimmune disease or inflammation in CD14−/− mice. J. Cell Biol. 167, 1161–1170 (2004). This work showed that persistent, uncleared apoptotic bodies in CD14-deficient mice did not lead to an immune response to self antigens, and challenged the idea that uncleared apoptotic cells always promote autoimmunity.
Lucas, M. et al. Requirements for apoptotic cell contact in regulation of macrophage responses. J. Immunol. 177, 4047–4054 (2006). This work revealed that only macrophages that have come in contact with apoptotic cells are sensitive to suppression by TGFβ that is produced by cells that have engaged and/or engulfed apoptotic cells.
Stuart, L. M., Takahashi, K., Shi, L., Savill, J. & Ezekowitz, R. A. Mannose-binding lectin-deficient mice display defective apoptotic cell clearance but no autoimmune phenotype. J. Immunol. 174, 3220–3226 (2005).
Voll, R. E. et al. Immunosuppressive effects of apoptotic cells. Nature 390, 350–351 (1997).
Kim, S., Elkon, K. B. & Ma, X. Transcriptional suppression of interleukin-12 gene expression following phagocytosis of apoptotic cells. Immunity 21, 643–653 (2004).
Huynh, M. L., Fadok, V. A. & Henson, P. M. Phosphatidylserine-dependent ingestion of apoptotic cells promotes TGF-β1 secretion and the resolution of inflammation. J. Clin. Invest. 109, 41–50 (2002). Using a model of lung inflammation, this work revealed a crucial in vivo role for the secretion of TGFβ1 by macrophages that engulf apoptotic cells using a PtdSer-dependent mechanism.
Cvetanovic, M. & Ucker, D. S. Innate immune discrimination of apoptotic cells: repression of proinflammatory macrophage transcription is coupled directly to specific recognition. J. Immunol. 172, 880–889 (2004).
Gerbod-Giannone, M. C. et al. TNFα induces ABCA1 through NF-κB in macrophages and in phagocytes ingesting apoptotic cells. Proc. Natl Acad. Sci. USA 103, 3112–3117 (2006).
Beaven, S. W. & Tontonoz, P. Nuclear receptors in lipid metabolism: targeting the heart of dyslipidemia. Annu. Rev. Med. 57, 313–329 (2006).
Choudhury, R. P., Lee, J. M. & Greaves, D. R. Mechanisms of disease: macrophage-derived foam cells emerging as therapeutic targets in atherosclerosis. Nature Clin. Pract. Cardiovasc. Med. 2, 309–315 (2005).
Condeelis, J. & Pollard, J. W. Macrophages: obligate partners for tumor cell migration, invasion, and metastasis. Cell 124, 263–266 (2006).
Vance, J. E. & Steenbergen, R. Metabolism and functions of phosphatidylserine. Prog. Lipid Res. 44, 207–234 (2005).
Tang, X., Halleck, M. S., Schlegel, R. A. & Williamson, P. A subfamily of P-type ATPases with aminophospholipid transporting activity. Science 272, 1495–1497 (1996).
Zullig, S. et al. Aminophospholipid translocase TAT-1 promotes phosphatidylserine exposure during C. elegans apoptosis. Curr. Biol. 17, 994–999 (2007).
Sahu, S. K., Gummadi, S. N., Manoj, N. & Aradhyam, G. K. Phospholipid scramblases: an overview. Arch. Biochem. Biophys. 462, 103–314 (2007).
Venegas, V. & Zhou, Z. Two alternative mechanisms that regulate the presentation of apoptotic cell engulfment signal in Caenorhabditis elegans. Mol. Biol. Cell 18, 3180–3192 (2007).
Wang, X. et al. C. elegans mitochondrial factor WAH-1 promotes phosphatidylserine externalization in apoptotic cells through phospholipid scramblase SCRM-1. Nature Cell Biol. 9, 541–549 (2007).
Williamson, P. & Schlegel, R. A. Transbilayer phospholipid movement and the clearance of apoptotic cells. Biochim. Biophys. Acta 1585, 53–63 (2002).
Miyanishi, M. et al. Identification of Tim4 as a phosphatidylserine receptor. Nature 24 October 2007 (doi: 10.1038/nature06307).
Park, S.-Y. et al. Rapid cell corpse clearance by stabilin-2, a membrane phosphatidylserine receptor. Cell Death Differ. 26 October 2007 (doi: 10.1038/sj.cdd.4402242).
Acknowledgements
The authors wish to thank colleagues in the engulfment field and members of our laboratories for many insightful discussions. This work was supported by grants GM064709 and GM069998 (to K.S.R.) and AI48672 (to U.L.).
Author information
Authors and Affiliations
Corresponding author
Related links
Glossary
- Apoptosis
-
A common type of cell death, also referred to as programmed cell death. Many physiological and developmental stimuli induce apoptosis. This mechanism is frequently used to delete unwanted, superfluous or potentially harmful cells during development and as part of normal homeostasis. By displaying markers for their removal at the earliest stages after the initiation of programmed cell death, apoptotic cells facilitate their removal in a 'safe package', without leakage of cellular contents.
- Necrosis
-
A form of cell death that frequently results from toxic injury, hypoxia or stress. Necrosis involves the loss of cell integrity and release of cell contents into the interstitium. This form of cell death usually occurs together with inflammation. Depending on the context, the self antigens that are released by necrosis could become immunogenic.
- Tingible-body macrophages
-
(TBMs). A type of macrophage that is specifically located in the germinal centres, where B cells and T cells cooperate in the production of antibody responses. TBMs are found in close proximity to follicular dendritic cells (FDCs), and engulf lymphocytes that undergo apoptosis in the germinal centres. TBMs can also have an inhibitory effect on the B-cell-mediated stimulation of T-cell responses when added to ex vivo co-cultures.
- RHO-family GTPases
-
A subfamily of small (∼21 kDa) GTP-binding proteins, which are members of the RAS superfamily of GTPases and have a key role in the rearrangement of the cytoskeleton. The nucleotide-bound state of these GTPases is generally regulated by guanine-nucleotide-exchange factors (GEFs), which catalyse GDP–GTP exchange, and GTPase-activating proteins, which facilitate the hydrolysis of the bound GTP.
- Membrane ruffling
-
A region of the plasma membrane that undergoes rapid morphological reorganization owing to the reorganization of the actin cytoskeleton and other cytoskeletal elements. Such ruffling and formation of the lamellae are seen on the phagocyte at the site of contact with the apoptotic cell and precede the uptake of targets.
- Macropinocytosis
-
A type of endocytosis (or phagocytosis) that occurs during the engulfment of apoptotic cells. During macropinocytosis, large droplets of fluid are trapped within the membrane protrusions (ruffles) or phagocytic arms.
- Docker domain
-
(DOCK180 region involved in RAC activation domain.) A new type of GEF domain, first discovered in DOCK180. There are currently 11 proteins with a Docker domain in the human genome, thereby defining a superfamily of proteins. This family of proteins is linked to the engulfment of apoptotic cells, cell migration, neuronal development and tumorigenesis.
- Ubiquitylation
-
The attachment of the small protein ubiquitin to lysine residues that are present in other proteins. This tags these proteins for rapid cellular degradation.
- Systemic lupus erythematosus
-
(SLE). A chronic systemic autoimmune disease that is characterized by rashes, arthritis, kidney disease and central-nervous-system involvement. It is mediated by antibodies that are specific for double-stranded DNA and other nuclear antigens.
- Atherosclerotic lesions
-
Lesions that initially start as low-grade inflammation in the vessel wall, with recruitment of monocytes that differentiate into macrophages. With further intracellular lipid accumulation and formation of foam cells, 'atheromas' develop, which are made up of cells (or cell debris) that contain cholesterol and lipids, calcium deposition and variable degree of fibrous connective tissue. The later-stage lesions contain a core of extracellular lipid surrounding the cholesterol-laden cells, many of which undergo apoptosis or necrosis.
- Foam cells
-
Macrophages that localize to sites of early stage inflammation in the vessel wall, which subsequently ingest oxidized low-density lipoprotein and slowly become overloaded with lipids. They are called foam cells because of their appearance, including numerous cytoplasmic vesicles containing cholesterol and other lipids. Foam cells eventually die and attract more macrophages, and further propagate the inflammation in the vessel wall.
Rights and permissions
About this article
Cite this article
Ravichandran, K., Lorenz, U. Engulfment of apoptotic cells: signals for a good meal. Nat Rev Immunol 7, 964–974 (2007). https://doi.org/10.1038/nri2214
Issue Date:
DOI: https://doi.org/10.1038/nri2214
This article is cited by
-
Proteomics analysis of methionine enkephalin upregulated macrophages against infection by the influenza-A virus
Proteome Science (2023)
-
Internalization of apoptotic cells during efferocytosis requires Mertk-mediated calcium influx
Cell Death & Disease (2023)
-
Patients with ACPA-positive and ACPA-negative rheumatoid arthritis show different serological autoantibody repertoires and autoantibody associations with disease activity
Scientific Reports (2023)
-
Externalized phosphatidylinositides on apoptotic cells are eat-me signals recognized by CD14
Cell Death & Differentiation (2022)
-
Old plasma dilution reduces human biological age: a clinical study
GeroScience (2022)