Key Points
-
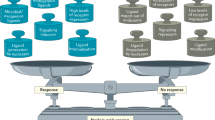
In innate immune responses, molecular patterns that are associated with invading pathogens are recognized by two classes of pattern-recognition receptor (PRR): transmembrane PRRs, namely Toll-like receptors (TLRs); and cytosolic PRRs, including retinoic-acid-inducible gene I (RIG-I) and melanoma-differentiation-associated gene 5 (MDA5). Signalling through these PRRs results in the activation of transcription factors that regulate genes encoding chemokines and other cytokines.
-
The interferon (IFN)-regulatory factor (IRF) family of transcription factors is crucial for the regulation of various aspects of immune responses, most notably those mediated by PRRs. The family comprises nine members, each of which contains a well-conserved DNA-binding domain that recognizes IFN-stimulated response elements (ISREs) in the promoters of target genes.
-
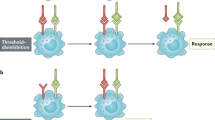
The activation of cytosolic PRRs typically elicits expression of type I IFN genes (the genes that encode IFNα and IFNβ). IRF7 functions as the master regulator of induction of these genes, and IRF3 contributes to this induction. Both IRF3 and IRF7 are activated by TBK1 (TANK (tumour-necrosis-factor-receptor-associated factor (TRAF)-family-member-associated nuclear factor (NF-κB) activator)-binding kinase 1), which phosphorylates these IRFs to convert them into an active form.
-
Signalling through TLRs is mainly mediated by two distinct adaptor molecules: MyD88 (myeloid differentiation primary-response protein 88) and TRIF (Toll/interleukin-1 receptor (TIR)-domain-containing adaptor protein inducing IFNβ). In the MyD88-dependent pathway, IRF4, IRF5 and IRF7 directly interact with MyD88 and regulate gene-expression programmes in this way. IRF7 is essential for the robust type I IFN gene induction that is elicited by ligation of TLR7 or TLR9, whereas IRF5 is required for the induction of pro-inflammatory cytokine genes. By contrast, IRF3 has an essential role in the TRIF-dependent pathway of type I IFN gene induction by TLR4.
-
IRFs interact with other transcription factors, such as NF-κB, and these interactions determine the specificity and magnitude of transcriptional events that are induced by PRR activation.
-
The aberrant activation of IRFs by PRRs has been implicated in the development of autoimmune diseases such as systemic lupus erythematosus.
Abstract
The interferon-regulatory factor (IRF) family of transcription factors was initially found to be involved in the induction of genes that encode type I interferons. IRFs have now been shown to have functionally diverse roles in the regulation of the immune system. Recently, the crucial involvement of IRFs in innate and adaptive immune responses has been gaining much attention, particularly with the discovery of their role in immunoregulation by Toll-like receptors and other pattern-recognition receptors.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Tjian, R. & Maniatis, T. Transcriptional activation: a complex puzzle with few easy pieces. Cell 77, 5–8 (1994).
Lenardo, M. J., Fan, C. M., Maniatis, T. & Baltimore, D. The involvement of NF-κB in β-interferon gene regulation reveals its role as widely inducible mediator of signal transduction. Cell 57, 287–294 (1989).
Li, Q. & Verma, I. M. NF-κB regulation in the immune system. Nature Rev. Immunol. 2, 725–734 (2002).
Taniguchi, T., Ogasawara, K., Takaoka, A. & Tanaka, N. IRF family of transcription factors as regulators of host defense. Annu. Rev. Immunol. 19, 623–655 (2001).
Lohoff, M. & Mak, T. W. Roles of interferon-regulatory factors in T-helper-cell differentiation. Nature Rev. Immunol. 5, 125–135 (2005).
Taniguchi, T. & Takaoka, A. A weak signal for strong responses: interferon-α/β revisited. Nature Rev. Mol. Cell Biol. 2, 378–386 (2001).
Platanias, L. C. Mechanisms of type-I- and type-II-interferon-mediated signalling. Nature Rev. Immunol. 5, 375–386 (2005).
Decker, T., Muller, M. & Stockinger, S. The Yin and Yang of type I interferon activity in bacterial infection. Nature Rev. Immunol. 5, 675–687 (2005).
Eroshkin, A. & Mushegian, A. Conserved transactivation domain shared by interferon regulatory factors and Smad morphogens. J. Mol. Med. 77, 403–405 (1999).
Akira, S., Uematsu, S. & Takeuchi, O. Pathogen recognition and innate immunity. Cell 124, 783–801 (2006).
Janeway, C. A. Jr. & Medzhitov, R. Innate immune recognition. Annu. Rev. Immunol. 20, 197–216 (2002).
Yoneyama, M. et al. The RNA helicase RIG-I has an essential function in double-stranded RNA-induced innate antiviral responses. Nature Immunol. 5, 730–737 (2004).
Yoneyama, M. et al. Shared and unique functions of the DExD/H-box helicases RIG-I, MDA5, and LGP2 in antiviral innate immunity. J. Immunol. 175, 2851–2858 (2005). References 12 and 13 were the first reports of the cytosolic PRR molecules involved in type I IFN gene induction by viruses or dsRNA.
Mamane, Y. et al. Interferon regulatory factors: the next generation. Gene 237, 1–14 (1999).
Wathelet, M. G. et al. Virus infection induces the assembly of coordinately activated transcription factors on the IFN-β enhancer in vivo. Mol. Cell 1, 507–518 (1998).
Miyamoto, M. et al. Regulated expression of a gene encoding a nuclear factor, IRF-1, that specifically binds to IFN-β gene regulatory elements. Cell 54, 903–913 (1988). This paper reports the discovery of the first IRF-family member, IRF1.
Matsuyama, T. et al. Targeted disruption of IRF-1 or IRF-2 results in abnormal type I IFN gene induction and aberrant lymphocyte development. Cell 75, 83–97 (1993).
Takaoka, A. et al. Integral role of IRF-5 in the gene induction programme activated by Toll-like receptors. Nature 434, 243–249 (2005). This paper was the first report to show that IRF5 is essential for TLR-mediated induction of pro-inflammatory cytokine genes.
Weaver, B. K., Kumar, K. P. & Reich, N. C. Interferon regulatory factor 3 and CREB-binding protein/p300 are subunits of double-stranded RNA-activated transcription factor DRAF1. Mol. Cell. Biol. 18, 1359–1368 (1998).
Lin, R., Heylbroeck, C., Pitha, P. M. & Hiscott, J. Virus-dependent phosphorylation of the IRF-3 transcription factor regulates nuclear translocation, transactivation potential, and proteasome-mediated degradation. Mol. Cell. Biol. 18, 2986–2996 (1998).
Yoneyama, M. et al. Direct triggering of the type I interferon system by virus infection: activation of a transcription factor complex containing IRF-3 and CBP/p300. EMBO J. 17, 1087–1095 (1998).
Sato, M., Tanaka, N., Hata, N., Oda, E. & Taniguchi, T. Involvement of the IRF family transcription factor IRF-3 in virus-induced activation of the IFN-β gene. FEBS Lett. 425, 112–116 (1998). References 19–22 provide evidence for the phosphorylation-dependent activation of IRF3 during type I IFN gene induction.
Suhara, W. et al. Analyses of virus-induced homomeric and heteromeric protein associations between IRF-3 and coactivator CBP/p300. J. Biochem. (Tokyo) 128, 301–307 (2000).
Sato, M. et al. Positive feedback regulation of type I IFN genes by the IFN-inducible transcription factor IRF-7. FEBS Lett. 441, 106–110 (1998).
Marie, I., Durbin, J. E. & Levy, D. E. Differential viral induction of distinct interferon-α genes by positive feedback through interferon regulatory factor-7. EMBO J. 17, 6660–6669 (1998). References 24 and 25 describe the positive-feedback regulation of type I IFN gene induction, which involves the expression and activation of IRF7.
Lin, R., Mamane, Y. & Hiscott, J. Multiple regulatory domains control IRF-7 activity in response to virus infection. J. Biol. Chem. 275, 34320–34327 (2000).
Sato, M. et al. Distinct and essential roles of transcription factors IRF-3 and IRF-7 in response to viruses for IFN-α/β gene induction. Immunity 13, 539–548 (2000).
Honda, K. et al. IRF-7 is the master regulator of type-I interferon-dependent immune responses. Nature 434, 772–777 (2005). References 27 and 28 are gene-targeting studies of IRF3 and IRF7, respectively. Reference 28 provides definitive evidence that IRF7 is the main regulator of both cytosolic PRR- and TLR-mediated type I IFN gene induction.
Nakaya, T. et al. Gene induction pathways mediated by distinct IRFs during viral infection. Biochem. Biophys. Res. Commun. 283, 1150–1156 (2001).
Sharma, S. et al. Triggering the interferon antiviral response through an IKK-related pathway. Science 300, 1148–1151 (2003).
Fitzgerald, K. A. et al. IKKε and TBK1 are essential components of the IRF3 signaling pathway. Nature Immunol. 4, 491–496 (2003). References 30 and 31 were the first reports to identify that TBK1 and IKK ε are the protein kinases that activate IRF3 and IRF7.
Hemmi, H. et al. The roles of two IκB kinase-related kinases in lipopolysaccharide and double stranded RNA signaling and viral infection. J. Exp. Med. 199, 1641–1650 (2004).
Perry, A. K., Chow, E. K., Goodnough, J. B., Yeh, W. C. & Cheng, G. Differential requirement for TANK-binding kinase-1 in type I interferon responses to Toll-like receptor activation and viral infection. J. Exp. Med. 199, 1651–1658 (2004).
Uematsu, S. et al. Interleukin-1 receptor-associated kinase-1 plays an essential role for Toll-like receptor (TLR)7- and TLR9-mediated interferon-α induction. J. Exp. Med. 201, 915–923 (2005).
Hoshino, K. et al. IκB kinase-α is critical for interferon-α production induced by Toll-like receptors 7 and 9. Nature 440, 949–953 (2006). References 34 and 35 show that IRAK1 and IKK α are involved in the activation of IRF7.
Obata, Y. et al. Role of cyclophilin B in activation of interferon regulatory factor-3. J. Biol. Chem. 280, 18355–18360 (2005).
Saitoh, T. et al. Negative regulation of interferon-regulatory factor 3-dependent innate antiviral response by the prolyl isomerase Pin1. Nature Immunol. 7, 598–605 (2006).
Huang, J. et al. SIKE is an IKKε/TBK1-associated suppressor of TLR3- and virus-triggered IRF-3 activation pathways. EMBO J. 24, 4018–4028 (2005).
Honda, K. et al. Selective contribution of IFN-α/β signaling to the maturation of dendritic cells induced by double-stranded RNA or viral infection. Proc. Natl Acad. Sci. USA 100, 10872–10877 (2003).
Kato, H. et al. Cell type-specific involvement of RIG-I in antiviral response. Immunity 23, 19–28 (2005).
Kato, H. et al. Differential roles of MDA5 and RIG-I helicases in the recognition of RNA viruses. Nature 441, 101–105 (2006).
Kawai, T. et al. IPS-1, an adaptor triggering RIG-I- and Mda5-mediated type I interferon induction. Nature Immunol. 6, 981–988 (2005).
Meylan, E. et al. Cardif is an adaptor protein in the RIG-I antiviral pathway and is targeted by hepatitis C virus. Nature 437, 1167–1172 (2005).
Xu, L. G. et al. VISA is an adapter protein required for virus-triggered IFN-β signaling. Mol. Cell 19, 727–740 (2005).
Seth, R. B., Sun, L., Ea, C. K. & Chen, Z. J. Identification and characterization of MAVS, a mitochondrial antiviral signaling protein that activates NF-κB and IRF 3. Cell 122, 669–682 (2005). References 42–45 report the discovery of the cytosolic adaptor molecule CARDIF/IPS1/MAVS/VISA.
Sun, Q. et al. The specific and essential role of MAVS in antiviral innate immune responses. Immunity 24, 633–642 (2006).
Kumar, H. et al. Essential role of IPS-1 in innate immune responses against RNA viruses. J. Exp. Med. 203, 1795–1803 (2006). References 46 and 47 describe gene-targeting studies of CARDIF/IPS1/MAVS/VISA and show that this adaptor has an essential role in the cytosolic pathway of type I IFN gene induction.
Oganesyan, G. et al. Critical role of TRAF3 in the Toll-like receptor-dependent and -independent antiviral response. Nature 439, 208–211 (2006).
Stockinger, S. et al. IFN regulatory factor 3-dependent induction of type I IFNs by intracellular bacteria is mediated by a TLR- and Nod2-independent mechanism. J. Immunol. 173, 7416–7425 (2004).
O'Connell, R. M. et al. Type I interferon production enhances susceptibility to Listeria monocytogenes infection. J. Exp. Med. 200, 437–445 (2004).
Stetson, D. B. & Medzhitov, R. Recognition of cytosolic DNA activates an IRF3-dependent innate immune response. Immunity 24, 93–103 (2006).
Ishii, K. J. et al. A Toll-like receptor-independent antiviral response induced by double-stranded B-form DNA. Nature Immunol. 7, 40–48 (2006).
Medzhitov, R., Preston-Hurlburt, P. & Janeway, C. A. Jr. A human homologue of the Drosophila Toll protein signals activation of adaptive immunity. Nature 388, 394–397 (1997).
Kurt-Jones, E. A. et al. Pattern recognition receptors TLR4 and CD14 mediate response to respiratory syncytial virus. Nature Immunol. 1, 398–401 (2000).
Hoshino, K., Kaisho, T., Iwabe, T., Takeuchi, O. & Akira, S. Differential involvement of IFN-β in Toll-like receptor-stimulated dendritic cell activation. Int. Immunol. 14, 1225–1231 (2002).
Kawai, T. et al. Lipopolysaccharide stimulates the MyD88-independent pathway and results in activation of IFN-regulatory factor 3 and the expression of a subset of lipopolysaccharide-inducible genes. J. Immunol. 167, 5887–5894 (2001). This paper provides evidence that there is a MyD88-independent pathway for activation of IRF3.
Sakaguchi, S. et al. Essential role of IRF-3 in lipopolysaccharide-induced interferon-β gene expression and endotoxin shock. Biochem. Biophys. Res. Commun. 306, 860–866 (2003).
Hoebe, K. et al. Upregulation of costimulatory molecules induced by lipopolysaccharide and double-stranded RNA occurs by Trif-dependent and Trif-independent pathways. Nature Immunol. 4, 1223–1229 (2003).
Werner, S. L., Barken, D. & Hoffmann, A. Stimulus specificity of gene expression programs determined by temporal control of IKK activity. Science 309, 1857–1861 (2005).
Ogawa, S. et al. Molecular determinants of crosstalk between nuclear receptors and Toll-like receptors. Cell 122, 707–721 (2005). Reference 60, together with references 101 and 102, provides evidence for the association of IRF3 with the NF-κ B component p65.
Kamijo, R. et al. Requirement for transcription factor IRF-1 in NO synthase induction in macrophages. Science 263, 1612–1615 (1994).
Alexopoulou, L., Holt, A. C., Medzhitov, R. & Flavell, R. A. Recognition of double-stranded RNA and activation of NF-κB by Toll-like receptor 3. Nature 413, 732–738 (2001).
Wang, T. et al. Toll-like receptor 3 mediates West Nile virus entry into the brain causing lethal encephalitis. Nature Med. 10, 1366–1373 (2004). References 62 and 63 are the initial reports on the role of TLR3 in the recognition of virus-derived and synthetic dsRNA.
Rudd, B. D. et al. Deletion of TLR3 alters the pulmonary immune environment and mucus production during respiratory syncytial virus infection. J. Immunol. 176, 1937–1942 (2006).
Tabeta, K. et al. Toll-like receptors 9 and 3 as essential components of innate immune defense against mouse cytomegalovirus infection. Proc. Natl Acad. Sci. USA 101, 3516–3521 (2004).
Flandin, J. F., Chano, F. & Descoteaux, A. RNA interference reveals a role for TLR2 and TLR3 in the recognition of Leishmania donovani promastigotes by interferon-γ-primed macrophages. Eur. J. Immunol. 36, 411–420 (2006).
Aksoy, E. et al. Double-stranded RNAs from the helminth parasite Schistosoma activate TLR3 in dendritic cells. J. Biol. Chem. 280, 277–283 (2005).
Yamamoto, M. et al. Role of adaptor TRIF in the MyD88-independent Toll-like receptor signaling pathway. Science 301, 640–643 (2003).
Sarkar, S. N. et al. Novel roles of TLR3 tyrosine phosphorylation and PI3 kinase in double-stranded RNA signaling. Nature Struct. Mol. Biol. 11, 1060–1067 (2004).
Sato, S. et al. Toll/IL-1 receptor domain-containing adaptor inducing IFN-β (TRIF) associates with TNF receptor-associated factor 6 and TANK-binding kinase 1, and activates two distinct transcription factors, NF-κB and IFN-regulatory factor-3, in the Toll-like receptor signaling. J. Immunol. 171, 4304–4310 (2003).
Gohda, J., Matsumura, T. & Inoue, J. TNFR-associated factor (TRAF) 6 is essential for MyD88-dependent pathway but not Toll/IL-1 receptor domain-containing adaptor-inducing IFN-β (TRIF)-dependent pathway in TLR signaling. J. Immunol. 173, 2913–2917 (2004).
Hacker, H. et al. Specificity in Toll-like receptor signalling through distinct effector functions of TRAF3 and TRAF6. Nature 439, 204–207 (2006).
Takeshita, F. et al. TRAF4 acts as a silencer in TLR-mediated signaling through the association with TRAF6 and TRIF. Eur. J. Immunol. 35, 2477–2485 (2005).
Su, X. et al. TNF receptor-associated factor-1 (TRAF1) negatively regulates Toll/IL-1 receptor domain-containing adaptor inducing IFN-β (TRIF)-mediated signaling. Eur. J. Immunol. 36, 199–206 (2006).
Sasai, M. et al. NF-κB-activating kinase-associated protein 1 participates in TLR3/Toll–IL-1 homology domain-containing adapter molecule-1-mediated IFN regulatory factor 3 activation. J. Immunol. 174, 27–30 (2005).
Nakano, H., Yanagita, M. & Gunn, M. D. CD11c+B220+Gr-1+ cells in mouse lymph nodes and spleen display characteristics of plasmacytoid dendritic cells. J. Exp. Med. 194, 1171–1178 (2001).
Asselin-Paturel, C. et al. Mouse type I IFN-producing cells are immature APCs with plasmacytoid morphology. Nature Immunol. 2, 1144–1150 (2001).
Colonna, M., Trinchieri, G. & Liu, Y. J. Plasmacytoid dendritic cells in immunity. Nature Immunol. 5, 1219–1226 (2004).
Wagner, H. The immunobiology of the TLR9 subfamily. Trends Immunol. 25, 381–386 (2004).
Diebold, S. S., Kaisho, T., Hemmi, H., Akira, S. & Reis e Sousa, C. Innate antiviral responses by means of TLR7-mediated recognition of single-stranded RNA. Science 303, 1529–1531 (2004).
Hemmi, H., Kaisho, T., Takeda, K. & Akira, S. The roles of Toll-like receptor 9, MyD88, and DNA-dependent protein kinase catalytic subunit in the effects of two distinct CpG DNAs on dendritic cell subsets. J. Immunol. 170, 3059–3064 (2003).
Krug, A. et al. Herpes simplex virus type 1 activates murine natural interferon-producing cells through Toll-like receptor 9. Blood 103, 1433–1437 (2004).
Lund, J., Sato, A., Akira, S., Medzhitov, R. & Iwasaki, A. Toll-like receptor 9-mediated recognition of herpes simplex virus-2 by plasmacytoid dendritic cells. J. Exp. Med. 198, 513–520 (2003).
Krug, A. et al. TLR9-dependent recognition of MCMV by IPC and DC generates coordinated cytokine responses that activate antiviral NK cell function. Immunity 21, 107–119 (2004).
Heil, F. et al. Species-specific recognition of single-stranded RNA via Toll-like receptor 7 and 8. Science 303, 1526–1529 (2004).
Lund, J. M. et al. Recognition of single-stranded RNA viruses by Toll-like receptor 7. Proc. Natl Acad. Sci. USA 101, 5598–5603 (2004).
Honda, K. et al. Role of a transductional–transcriptional processor complex involving MyD88 and IRF-7 in Toll-like receptor signaling. Proc. Natl Acad. Sci. USA 101, 15416–15421 (2004).
Kawai, T. et al. Interferon-α induction through Toll-like receptors involves a direct interaction of IRF7 with MyD88 and TRAF6. Nature Immunol. 5, 1061–1068 (2004). References 87 and 88 report the evidence for a direct interaction between MyD88 and IRF7.
Honda, K. et al. Spatiotemporal regulation of MyD88–IRF-7 signalling for robust type-I interferon induction. Nature 434, 1035–1040 (2005). This paper shows the importance of endosomal MyD88–IRF7 signalling for robust induction of type I IFN genes in pDCs.
Verthelyi, D., Ishii, K. J., Gursel, M., Takeshita, F. & Klinman, D. M. Human peripheral blood cells differentially recognize and respond to two distinct CpG motifs. J. Immunol. 166, 2372–2377 (2001).
Klinman, D. M. Immunotherapeutic uses of CpG oligodeoxynucleotides. Nature Rev. Immunol. 4, 249–258 (2004).
Shinohara, M. L. et al. Osteopontin expression is essential for interferon-α production by plasmacytoid dendritic cells. Nature Immunol. 7, 498–506 (2006).
Yanai, H. et al. IRF family transcription factors in type I interferon induction. Int. Congr. Ser. 1285, 104–113 (2005).
Durrer, P., Gaudin, Y., Ruigrok, R. W., Graf, R. & Brunner, J. Photolabeling identifies a putative fusion domain in the envelope glycoprotein of rabies and vesicular stomatitis viruses. J. Biol. Chem. 270, 17575–17581 (1995).
Brunetti, C. R., Dingwell, K. S., Wale, C., Graham, F. L. & Johnson, D. C. Herpes simplex virus gD and virions accumulate in endosomes by mannose 6-phosphate-dependent and -independent mechanisms. J. Virol. 72, 3330–3339 (1998).
Negishi, H. et al. Negative regulation of Toll-like-receptor signaling by IRF-4. Proc. Natl Acad. Sci. USA 102, 15989–15994 (2005).
Schoenemeyer, A. et al. The interferon regulatory factor, IRF5, is a central mediator of Toll-like receptor 7 signaling. J. Biol. Chem. 280, 17005–17012 (2005).
Zhao, J. et al. IRF-8/interferon (IFN) consensus sequence-binding protein is involved in Toll-like receptor (TLR) signaling and contributes to the cross-talk between TLR and IFN-γ signaling pathways. J. Biol. Chem. 281, 10073–10080 (2006).
Tsujimura, H. et al. Toll-like receptor 9 signaling activates NF-κB through IFN regulatory factor-8/IFN consensus sequence binding protein in dendritic cells. J. Immunol. 172, 6820–6827 (2004).
Marecki, S. & Fenton, M. J. The role of IRF-4 in transcriptional regulation. J. Interferon Cytokine Res. 22, 121–133 (2002).
Leung, T. H., Hoffmann, A. & Baltimore, D. One nucleotide in a κB site can determine cofactor specificity for NF-κB dimers. Cell 118, 453–464 (2004).
Wietek, C., Miggin, S. M., Jefferies, C. A. & O'Neill, L. A. Interferon regulatory factor-3-mediated activation of the interferon-sensitive response element by Toll-like receptor (TLR) 4 but not TLR3 requires the p65 subunit of NF-κB. J. Biol. Chem. 278, 50923–50931 (2003).
Kariko, K., Buckstein, M., Ni, H. & Weissman, D. Suppression of RNA recognition by Toll-like receptors: the impact of nucleoside modification and the evolutionary origin of RNA. Immunity 23, 165–175 (2005).
Koski, G. K. et al. Innate immune system discriminates between RNA containing bacterial versus eukaryotic structural features that prime for high-level IL-12 secretion by dendritic cells. J. Immunol. 172, 3989–3993 (2004).
Krieg, A. M. CpG motifs in bacterial DNA and their immune effects. Annu. Rev. Immunol. 20, 709–760 (2002).
Vollmer, J. et al. Immune stimulation mediated by autoantigen binding sites within small nuclear RNAs involves Toll-like receptors 7 and 8. J. Exp. Med. 202, 1575–1585 (2005).
Lau, C. M. et al. RNA-associated autoantigens activate B cells by combined B cell antigen receptor/Toll-like receptor 7 engagement. J. Exp. Med. 202, 1171–1177 (2005).
Leadbetter, E. A. et al. Chromatin–IgG complexes activate B cells by dual engagement of IgM and Toll-like receptors. Nature 416, 603–607 (2002).
Ronnblom, L. & Alm, G. V. Systemic lupus erythematosus and the type I interferon system. Arthritis Res. Ther. 5, 68–75 (2003).
Boule, M. W. et al. Toll-like receptor 9-dependent and -independent dendritic cell activation by chromatin–immunoglobulin G complexes. J. Exp. Med. 199, 1631–1640 (2004).
Blanco, P., Palucka, A. K., Gill, M., Pascual, V. & Banchereau, J. Induction of dendritic cell differentiation by IFN-α in systemic lupus erythematosus. Science 294, 1540–1543 (2001).
Yasuda, K. et al. Endosomal translocation of vertebrate DNA activates dendritic cells via TLR9-dependent and -independent pathways. J. Immunol. 174, 6129–6136 (2005).
Wagner, H., Heit, A., Schmitz, F. & Bauer, S. Targeting split vaccines to the endosome improves vaccination. Curr. Opin. Biotechnol. 15, 538–542 (2004).
Graham, R. R. et al. A common haplotype of interferon regulatory factor 5 (IRF5) regulates splicing and expression and is associated with increased risk of systemic lupus erythematosus. Nature Genet. 38, 550–555 (2006). This paper reports that variants of IRF5 can confer risk of SLE.
Kimura, T. et al. Involvement of the IRF-1 transcription factor in antiviral responses to interferons. Science 264, 1921–1924 (1994).
Ko, J., Gendron-Fitzpatrick, A. & Splitter, G. A. Susceptibility of IFN regulatory factor-1 and IFN consensus sequence binding protein-deficient mice to brucellosis. J. Immunol. 168, 2433–2440 (2002).
Taki, S. et al. Multistage regulation of TH1-type immune responses by the transcription factor IRF-1. Immunity 6, 673–679 (1997).
Ogasawara, K. et al. Requirement for IRF-1 in the microenvironment supporting development of natural killer cells. Nature 391, 700–703 (1998).
Penninger, J. M. et al. The interferon regulatory transcription factor IRF-1 controls positive and negative selection of CD8+ thymocytes. Immunity 7, 243–254 (1997).
Tanaka, N. et al. Cooperation of the tumour suppressors IRF-1 and p53 in response to DNA damage. Nature 382, 816–818 (1996).
Tamura, T. et al. An IRF-1-dependent pathway of DNA damage-induced apoptosis in mitogen-activated T lymphocytes. Nature 376, 596–599 (1995).
Hida, S. et al. CD8+ T cell-mediated skin disease in mice lacking IRF-2, the transcriptional attenuator of interferon-α/β signaling. Immunity 13, 643–655 (2000).
Honda, K., Mizutani, T. & Taniguchi, T. Negative regulation of IFN-α/β signaling by IFN regulatory factor 2 for homeostatic development of dendritic cells. Proc. Natl Acad. Sci. USA 101, 2416–2421 (2004).
Lohoff, M. et al. Deficiency in the transcription factor interferon regulatory factor (IRF)-2 leads to severely compromised development of natural killer and T helper type 1 cells. J. Exp. Med. 192, 325–336 (2000).
Rengarajan, J. et al. Interferon regulatory factor 4 (IRF4) interacts with NFATc2 to modulate interleukin 4 gene expression. J. Exp. Med. 195, 1003–1012 (2002).
Lohoff, M. et al. Dysregulated T helper cell differentiation in the absence of interferon regulatory factor 4. Proc. Natl Acad. Sci. USA 99, 11808–11812 (2002).
Mittrucker, H. W. et al. Requirement for the transcription factor LSIRF/IRF4 for mature B and T lymphocyte function. Science 275, 540–543 (1997).
Lu, R., Medina, K. L., Lancki, D. W. & Singh, H. IRF-4, 8 orchestrate the pre-B-to-B transition in lymphocyte development. Genes Dev. 17, 1703–1708 (2003).
Tamura, T. et al. IFN regulatory factor-4 and -8 govern dendritic cell subset development and their functional diversity. J. Immunol. 174, 2573–2581 (2005).
Fehr, T. et al. Crucial role of interferon consensus sequence binding protein, but neither of interferon regulatory factor 1 nor of nitric oxide synthesis for protection against murine listeriosis. J. Exp. Med. 185, 921–931 (1997).
Giese, N. A. et al. Interferon (IFN) consensus sequence-binding protein, a transcription factor of the IFN regulatory factor family, regulates immune responses in vivo through control of interleukin 12 expression. J. Exp. Med. 186, 1535–1546 (1997).
Scharton-Kersten, T., Contursi, C., Masumi, A., Sher, A. & Ozato, K. Interferon consensus sequence binding protein-deficient mice display impaired resistance to intracellular infection due to a primary defect in interleukin 12 p40 induction. J. Exp. Med. 186, 1523–1534 (1997).
Holtschke, T. et al. Immunodeficiency and chronic myelogenous leukemia-like syndrome in mice with a targeted mutation of the ICSBP gene. Cell 87, 307–317 (1996).
Schiavoni, G. et al. ICSBP is essential for the development of mouse type I interferon-producing cells and for the generation and activation of CD8α+ dendritic cells. J. Exp. Med. 196, 1415–1425 (2002).
Harada, H. et al. Regulation of IFN-α/β genes: evidence for a dual function of the transcription factor complex ISGF3 in the production and action of IFN-α/β. Genes Cells 1, 995–1005 (1996).
Schulz, O. et al. Toll-like receptor 3 promotes cross-priming to virus-infected cells. Nature 433, 887–892 (2005).
Lee, J. et al. Molecular basis for the immunostimulatory activity of guanine nucleoside analogs: activation of Toll-like receptor 7. Proc. Natl Acad. Sci. USA 100, 6646–6651 (2003).
Hornung, V. et al. Sequence-specific potent induction of IFN-α by short interfering RNA in plasmacytoid dendritic cells through TLR7. Nature Med. 11, 263–270 (2005).
Judge, A. D. et al. Sequence-dependent stimulation of the mammalian innate immune response by synthetic siRNA. Nature Biotechnol. 23, 457–462 (2005).
Hemmi, H. et al. A Toll-like receptor recognizes bacterial DNA. Nature 408, 740–745 (2000). This paper reports on the discovery and function of TLR9.
Lin, R. & Hiscott, J. A role for casein kinase II phosphorylation in the regulation of IRF-1 transcriptional activity. Mol. Cell. Biochem. 191, 169–180 (1999).
Kondo, T. et al. Identification and characterization of nucleophosmin/B23/numatrin which binds the anti-oncogenic transcription factor IRF-1 and manifests oncogenic activity. Oncogene 15, 1275–1281 (1997).
Dornan, D. et al. Interferon regulatory factor 1 binding to p300 stimulates DNA-dependent acetylation of p53. Mol. Cell. Biol. 24, 10083–10098 (2004).
Reily, M. M., Pantoja, C., Hu, X., Chinenov, Y. & Rogatsky, I. The GRIP1:IRF3 interaction as a target for glucocorticoid receptor-mediated immunosuppression. EMBO J. 25, 108–117 (2006).
Mamane, Y., Sharma, S., Petropoulos, L., Lin, R. & Hiscott, J. Posttranslational regulation of IRF-4 activity by the immunophilin FKBP52. Immunity 12, 129–140 (2000).
Kim, Y. M. et al. Roles of IFN consensus sequence binding protein and PU.1 in regulating IL-18 gene expression. J. Immunol. 163, 2000–2007 (1999).
Qing, J. et al. Transforming growth factor β/Smad3 signaling regulates IRF-7 function and transcriptional activation of the β interferon promoter. Mol. Cell. Biol. 24, 1411–1425 (2004).
Darnell, J. E. Jr., Kerr, I. M. & Stark, G. R. Jak–STAT pathways and transcriptional activation in response to IFNs and other extracellular signaling proteins. Science 264, 1415–1421 (1994).
Acknowledgements
We thank E. Barsoumian, A. Takaoka, Y. Ohba and H. Yanai for valuable discussion and advice. This work was supported by Kakenhi (Grants-in-Aid for Scientific Research) on the Priority Area 'Integrative Research Toward the Conquest of Cancer', from the Ministry of Education, Culture, Sports, Science and Technology (Japan).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Related links
Glossary
- Helix–turn–helix motif
-
A structural motif that can bind DNA. It comprises two α-helices joined by a short strand of amino acids, and it is found in many proteins that regulate gene expression.
- IFN-stimulated response element
-
(ISRE). A common DNA motif that is found in the promoters of genes that are regulated by type I interferons (IFNs). It is bound by IFN-regulatory factors (IRFs) and was initially known as the IRF enhancer (IRFE). The consensus sequence is GAAANNGAAAG/CT/C, where N denotes any nucleotide.
- Type I IFNs
-
A family of cytokines that includes interferon (IFNα; which is encoded by 13 functional genes in humans and 14 in mice) and IFNβ.
- Pathogen-associated molecular patterns
-
(PAMPs). Molecular patterns that are found in pathogens but not in mammalian cells. Examples include terminally mannosylated and polymannosylated compounds (which bind the mannose receptor) and various microbial components, such as bacterial lipopolysaccharide, hypomethylated DNA, flagellin and double-stranded RNA (all of which bind Toll-like receptors).
- Nucleotide-binding oligomerization-domain proteins
-
(NOD proteins). Members of a family that includes the apoptosis regulator APAF1 (apoptotic-protease-activating factor 1), mammalian NOD-LRR proteins (also known as NACHT-LRR proteins or CATERPILLERs) and plant disease-resistance gene products. Several NOD proteins have been implicated in the induction of nuclear factor-κB activity and in the activation of caspases.
- Virus-responsive elements
-
(VREs). The promoter of the gene that encodes interferon-β (IFNβ) contains at least four regulatory cis elements — positive-regulatory domain I (PRDI), PRDII, PRDIII and PRDIV — which are involved in virus-mediated gene induction. IFN-regulatory factors (IRFs) bind PRDI and PRDIII (which are IFN-stimulated response elements, ISREs), whereas nuclear factor-κB and activator protein 1 (AP1) bind PRDII and PRDIV, respectively. By contrast, the promoters of the genes that encode IFNα contain only IRF-binding elements, and these are known as PRDI-like and PRDIII-like elements (PRD-LEs).
- Mouse embryonic fibroblasts
-
(MEFs). A well-defined cell type that has been widely used to identify the consequences of ablation or ectopic expression of a gene of interest. In addition, MEFs are known to allow infection with various viruses and to express type I IFN genes effectively. They therefore provide a simple model for the study of innate immunity to viral infections.
- Latent form
-
A protein that is inactive in the absence of additional modification(s), such as phosphorylation or ubiquitylation.
- Holocomplex
-
A complex that consists of subunits, each of which cannot carry out a reaction by itself but can carry out the reaction as a complex.
- Histone acetyltransferase
-
A protein that acetylates core histones, resulting in important regulatory effects on chromatin structure and assembly, and on gene transcription.
- Immunophilin family
-
A family of cis–trans peptidylprolyl isomerases that includes cyclophilins and FK506-binding proteins (FKBPs). These proteins were originally discovered as cellular receptors for immunosuppressive drugs, including cyclosporin A and FK506. The complexes that form between immunophilins and their cognate ligands are the functional modules for immunosuppression. Immunophilins are now known to function at the crossroads of protein folding and trafficking, and signal transduction.
- RNA-helicase domain
-
A protein domain that is found in many RNA-binding proteins that are required for mRNA synthesis, pre-mRNA splicing, ribosome biogenesis and RNA decay. This domain can unwind double-stranded RNA using energy derived from the hydrolysis of ATP.
- Caspase-recruitment domain
-
(CARD). A protein domain that is found in certain initiator caspases (for example, mammalian caspase-9) and their adaptor proteins (for example, apoptotic-protease-activating factor 1, APAF1). This domain mediates protein–protein interaction.
- B form of DNA
-
(B-DNA). DNA with a right-handed double helix. This is the conformation that is normally seen in solution and is thought to be the conformation of most DNA in vivo. It also formed the basis of the model described by James Watson and Francis Crick.
- Z form of DNA
-
(Z-DNA). DNA with a left-handed double helix. This conformation occurs as a consequence of methylation. It is found mainly in genes that are undergoing transcription. It is present only transiently, because the cessation of transcription results in rapid conversion of Z-DNA to the B form of DNA (the normal conformation), through the activity of topoisomerases.
- Plasmacytoid DCs
-
(pDCs). A subset of dendritic cells (DCs) that was named 'plasmacytoid' because their appearance under the microscope is similar to that of plasmablasts. In humans, these DCs can be derived from lineage (Lin)- haematopoietic stem cells from the peripheral blood. These DCs are the main producers of type I interferons in response to viral infections.
- CpG-A
-
Also known as D-type CpG. Synthetic oligodeoxynucleotides with the following three features: poly(G) sequences at the 3′ end; a central palindromic sequence; and CG dinucleotides within the palindrome. The poly(G) tails on CpG-A can interact with each other, resulting in the formation of G tetrads and clusters. The structure of CpG-A is interpreted by plasmacytoid DCs as a molecular pattern that indicates infection with a DNA virus, and this recognition elicits robust production of type I interferons. However, identical sequences have not been found in the genomes of DNA viruses.
- Death domain
-
A protein domain that is found in many proteins that are involved in signalling and apoptosis. This domain mediates protein–protein interaction.
- Endotoxic shock
-
A clinical condition that is induced by hyperreactivity of the innate immune system to bacterial lipopolysaccharide (LPS). It is mediated by the inflammatory cytokines interleukin-1 and tumour-necrosis factor, both of which are produced in high amounts following sustained activation of Toll-like receptor 4 by LPS.
- CpG-B
-
Also known as K-type CpG. Synthetic oligodeoxynucleotides that contain a CpG motif(s) on a phosphorothioate backbone. Analogous to bacterial infection, CpG-B triggers the differentiation of both plasmacytoid and conventional DCs, as well as the proliferation and activation of B cells. However, identical sequences have not been found in the genomes of bacteria.
- ETS/ISRE composite DNA motif
-
(ETS/interferon-stimulated-response-element composite DNA motif). A motif found in numerous genes that are essential for the proper function of macrophages and B cells. Interferon-regulatory factor 4 (IRF4) or IRF8 binds this motif following interaction with the transcription factor PU.1. The consensus sequence is GGAAGTGAAA, with the PU.1-binding core motif (at the 5′ end) and the IRF-binding core motif (at the 3′ end) underlined.
- REL-homology domain
-
(RHD). A conserved domain of ∼300 amino acids that is found in the amino-terminal region of nuclear factor-κB (NF-κB)-family members. It contains motifs that are responsible for dimerization, nuclear translocation and binding to NF-κB-binding motifs that are present in DNA.
- Systemic lupus erythematosus
-
(SLE). An autoimmune disease in which autoantibodies specific for DNA, RNA or proteins associated with nucleic acids form immune complexes. These complexes damage small blood vessels, especially in the kidneys. Patients with SLE generally have abnormal B- and T-cell function.
Rights and permissions
About this article
Cite this article
Honda, K., Taniguchi, T. IRFs: master regulators of signalling by Toll-like receptors and cytosolic pattern-recognition receptors. Nat Rev Immunol 6, 644–658 (2006). https://doi.org/10.1038/nri1900
Issue Date:
DOI: https://doi.org/10.1038/nri1900
This article is cited by
-
Interferon Regulatory Factor 5 Gene Polymorphisms and mRNA Expression Levels Are Associated with Neuromyelitis Optica Spectrum Disorder
Molecular Neurobiology (2024)
-
Sex differences in the percentage of IRF5 positive B cells are associated with higher production of TNF-α in women in response to TLR9 in humans
Biology of Sex Differences (2023)
-
Pan-cancer analysis of TASL: a novel immune infiltration-related biomarker for tumor prognosis and immunotherapy response prediction
BMC Cancer (2023)
-
Granulocyte-macrophage colony-stimulating factor suppresses induction of type I interferon in infants with severe pneumonia
Pediatric Research (2023)
-
Origins and diversification of animal innate immune responses against viral infections
Nature Ecology & Evolution (2023)