Key Points
-
Protein–protein interactions represent a large and important class of targets for human therapeutics.
-
However, developing small-molecule antagonists of protein–protein interactions is challenging, owing to issues such as the general lack of small-molecule starting points for drug design, the typical flatness of the interface, the difficulty of distinguishing real from artefactual binding, and the size and character of typical small-molecule libraries.
-
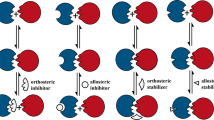
This article uses examples to describe general strategies in the development of small molecule antagonists of protein–protein interactions. Two types of antagonists are described: those that bind directly to the 'hot spot'of a protein–protein interface — a region that has a major contribution to high-affinity binding — and those that bind at allosteric sites distal from the protein–protein interface.
-
Finally, characteristics of programmes that have successfully identified small-molecule antagonists of protein–protein interactions are discussed. A high degree of validation is recommended for antagonists of protein–protein interactions owing to the nature of these targets, and a series of steps for validating and characterizing a new series of antagonists are presented.
Abstract
Protein–protein interactions have a key role in most biological processes, and offer attractive opportunities for therapeutic intervention. Developing small molecules that modulate protein–protein interactions is difficult, owing to issues such as the lack of well-defined binding pockets. Nevertheless, there has been important progress in this endeavour in recent years. Here, we use illustrative examples to discuss general strategies for addressing the challenges inherent in the discovery and characterization of small-molecule inhibitors of protein–protein interactions.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout









Similar content being viewed by others
References
Stockwin, L. & Holmes, S. Antibodies as therapeutic agents: vive la renaissance! Expert Opin. Biol. Ther. 3, 1133–1152 (2003).
Das, R. C. & Morrow, K. J. J. in Antibody Engineering Report (Drug and Market Development Publications, 2002).
Monoclonal Antibodies Report 2003: Meeting Clinical and Financial Expectations (Visiongain, 2003).
Berg, T. Modulation of protein–protein interactions with small organic molecules. Angew. Chem. Int. Ed. Engl. 42, 2462–2481 (2003).
Cochran, A. G. Protein–protein interfaces: mimics and inhibitors. Curr. Opin. Chem. Biol. 5, 654–659 (2001).
Gadek, T. R. & Nicholas, J. B. Small molecule antagonists of proteins. Biochem. Pharmacol. 65, 1–8 (2003).
Ockey, D. A. & Gadek, T. R. Inhibitors of protein–protein interactions. Expert Opin. Ther. Pat. 12, 393–400 (2002).
Sharma, S. K., Ramsey, T. M. & Bair, K. W. Protein–protein interactions: lessons learned. Curr. Med. Chem. Anti-Canc. Agents 2, 311–330 (2002).
Toogood, P. L. Inhibition of protein–protein association by small molecules: approaches and progress. J. Med. Chem. 45, 1–16 (2002).
Sulyok, G. A. et al. Solid-phase synthesis of a nonpeptide RGD mimetic library: new selective αvβ3 integrin antagonists. J. Med. Chem. 44, 1938–1950 (2001).
Goodman, S. L., Holzemann, G., Sulyok, G. A. & Kessler, H. Nanomolar small molecule inhibitors for αv(β)6, αv(β)5, and αv(β)3 integrins. J. Med. Chem. 45, 1045–1051 (2002).
Gibson, C. et al. Nonpeptidic α(v)β(3) Integrin antagonist libraries: on-bead screening and mass spectrometric identification without tagging. Angew. Chem. Int. Ed. Engl. 40, 165–169 (2001).
Hoekstra, W. J. & Poulter, B. L. Combinatorial chemistry techniques applied to nonpeptide integrin antagonists. Curr. Med. Chem. 5, 195–204 (1998).
Mousa, S. A. Anti-integrin as novel drug-discovery targets: potential therapeutic and diagnostic implications. Curr. Opin. Chem. Biol. 6, 534–541 (2002).
Scarborough, R. M. & Gretler, D. D. Platelet glycoprotein IIb-IIIa antagonists as prototypical integrin blockers: novel parenteral and potential oral antithrombotic agents. J. Med. Chem. 43, 3453–3473 (2000).
Eldred, C. D. & Judkins, B. D. Fibrinogen receptor antagonists: design and clinical applications. Prog. Med. Chem. 36, 29–90 (1999).
Ojima, I., Chakravarty, S. & Dong, Q. Antithrombotic agents: from RGD to peptide mimetics. Bioorg. Med. Chem. 3, 337–360 (1995).
Gadek, T. R. & McDowell, R. S. Discovery of small molecule leads in a biotechnology datastream. Drug Discov. Today 8, 545–550 (2003).
Jackson, D. Y. α4 integrin antagonists. Curr. Pharm. Des. 8, 1229–1253 (2002).
Jordan, M. A. Mechanism of action of antitumor drugs that interact with microtubules and tubulin. Curr. Med. Chem. Anti-Canc. Agents 2, 1–17 (2002).
Schreiber, S. L. & Crabtree, G. R. The mechanism of action of cyclosporin A and FK506. Immunol Today 13, 136–142 (1992).
Lo Conte, L., Chothia, C. & Janin, J. The atomic structure of protein–protein recognition sites. J. Mol. Biol. 285, 2177–2198 (1999).
Bogan, A. A. & Thorn, K. S. Anatomy of hot spots in protein interfaces. J. Mol. Biol. 280, 1–9 (1998).
Clackson, T. & Wells, J. A. A hot spot of binding energy in a hormone-receptor interface. Science 267, 383–386 (1995).
DeLano, W. L. Unraveling hot spots in binding interfaces: progress and challenges. Curr. Opin. Struct. Biol. 12, 14–20 (2002).
Ma, B., Elkayam, T., Wolfson, H. & Nussinov, R. Protein–protein interactions: structurally conserved residues distinguish between binding sites and exposed protein surfaces. Proc. Natl Acad. Sci. USA 100, 5772–5777 (2003).
Sundberg, E. J. & Mariuzza, R. A. Luxury accommodations: the expanding role of structural plasticity in protein–protein interactions. Structure Fold. Des. 8, R137–R142 (2000).
DeLano, W. L., Ultsch, M. H., de Vos, A. M. & Wells, J. A. Convergent solutions to binding at a protein–protein interface. Science 287, 1279–1283 (2000). The protein–protein hot spot on the Fc domain of immunoglobulin G is characterized by comparing several structures of the protein bound to its natural ligands and to a synthetic peptide. The hot spot is found to be more involuted and hydrophobic than most of the surface; in addition, the conformation of the binding site varies to complement each protein partner.
Teague, S. J. Implications of protein flexibility for drug discovery. Nature Rev. Drug Discov. 2, 527–541 (2003).
Luque, I. & Freire, E. Structural stability of binding sites: consequences for binding affinity and allosteric effects. Proteins S4, 63–71 (2000).
Ma, B., Shatsky, M., Wolfson, H. J. & Nussinov, R. Multiple diverse ligands binding at a single protein site: a matter of pre-existing populations. Protein Sci. 11, 184–197 (2002).
Sidhu, S. S., Fairbrother, W. J. & Deshayes, K. Exploring protein–protein interactions with phage display. Chembiochem. 4, 14–25 (2003). Recent review describing how phage display has been used to probe protein hot spots and identify novel peptide agonists/antagonists of protein–protein interactions.
Pillutla, R. C. et al. Peptides identify the critical hotspots involved in the biological activation of the insulin receptor. J. Biol. Chem. 277, 22590–22594 (2002).
Fairbrother, W. J. et al. Novel peptides selected to bind vascular endothelial growth factor target the receptor-binding site. Biochemistry 37, 17754–17764 (1998).
Lowman, H. B. Bacteriophage display and discovery of peptide leads for drug development. Annu. Rev. Biophys. Biomol. Struct. 26, 401–424 (1997).
Schaffer, M. L., Deshayes, K., Nakamura, G., Sidhu, S. & Skelton, N. J. Complex with a phage display-derived peptide provides insight into the function of insulin-like growth factor I. Biochemistry 42, 9324–9334 (2003).
Nakamura, G. R., Reynolds, M. E., Chen, Y. M., Starovasnik, M. A. & Lowman, H. B. Stable 'zeta' peptides that act as potent antagonists of the high-affinity IgE receptor. Proc. Natl Acad. Sci. USA 99, 1303–1308 (2002).
Wrighton, N. C. et al. Small peptides as potent mimetics of the protein hormone erythropoietin. Science 273, 458–464 (1996).
Cwirla, S. E. et al. Peptide agonist of the thrombopoietin receptor as potent as the natural cytokine. Science 276, 1696–1699 (1997).
Scott, J. K. et al. Evidence that a protein–protein interaction 'hot spot' on heterotrimeric G protein βγ subunits is used for recognition of a subclass of effectors. EMBO J. 20, 767–776 (2001).
McGovern, S. L., Helfand, B. T., Feng, B. & Shoichet, B. K. A specific mechanism of nonspecific inhibition. J. Med. Chem. 46, 4265–4272 (2003).
Seidler, J., McGovern, S. L., Doman, T. N. & Shoichet, B. K. Identification and prediction of promiscuous aggregating inhibitors among known drugs. J. Med. Chem. 46, 4477–4786 (2003).
McGovern, S. L., Caselli, E., Grigorieff, N. & Shoichet, B. K. A common mechanism underlying promiscuous inhibitors from virtual and high-throughput screening. J. Med. Chem. 45, 1712–1722 (2002). References 41–43 describe methods for identifying complexes that cause artefactual inhibition by an aggregating mechanism. This mechanism seems to be common at compound concentrations in the mid-micromolar range.
Carter, P. H. et al. Photochemically enhanced binding of small molecules to the tumor necrosis factor receptor-1 inhibits the binding of TNF-α. Proc. Natl Acad. Sci. USA 98, 11879–11884 (2001).
Wiekowski, M. et al. Characterization of potential antagonists of human interleukin 5 demonstrates their cross-reactivity with receptors for interleukin 3 and granulocyte–macrophage colony-stimulating factor. Eur. J. Biochem. 246, 625–632 (1997). References 44 and 45 describe the characterization of covalent inhibitors of protein–protein interactions. Interestingly, neither compound was initially intended for use as a covalent modifier, but this mechanism was identified through careful analysis.
Way, J. C. Covalent modification as a strategy to block protein–protein interactions with small-molecule drugs. Curr. Opin. Chem. Biol. 4, 40–46 (2000).
Woska, J. R. Jr, Morelock, M. M., Jeanfavre, D. D. & Bormann, B. J. Characterization of molecular interactions between intercellular adhesion molecule-1 and leukocyte function-associated antigen-1. J. Immunol. 156, 4680–4685 (1996).
Boehm, H. J. et al. Novel inhibitors of DNA gyrase: 3D structure based biased needle screening, hit validation by biophysical methods, and 3D guided optimization. A promising alternative to random screening. J. Med. Chem. 43, 2664–2664 (2000). This paper describes the efficient use of biophysical screens to assign priority to a series of early-stage compounds; half of the initial series are determined to inhibit by an acceptable mechanism.
Nelson, B. H. & Willerford, D. M. Biology of the interleukin-2 receptor. Adv. Immunol. 70, 1–81 (1998).
Berard, J. L., Velez, R. L., Freeman, R. B. & Tsunoda, S. M. A review of interleukin-2 receptor antagonists in solid organ transplantation. Pharmacotherapy 19, 1127–1137 (1999).
Waldmann, T. A. & O'Shea, J. The use of antibodies against the IL-2 receptor in transplantation. Curr. Opin. Immunol. 10, 507–512 (1998).
Zurawski, S. M. et al. Definition and spatial location of mouse interleukin-2 residues that interact with its heterotrimeric receptor. EMBO J. 12, 5113–5119 (1993).
Sauve, K. et al. Localization in human interleukin 2 of the binding site to the α chain (p55) of the interleukin 2 receptor. Proc. Natl Acad. Sci. USA 88, 4636–4640 (1991).
Emerson, S. D. et al. NMR characterization of interleukin-2 in complexes with the IL-2Rα receptor component, and with low molecular weight compounds that inhibit the IL-2/IL-Rα interaction. Protein Sci. 12, 811–822 (2003).
Tilley, J. W. et al. Identification of a small molecule inhibitor of the IL-2/IL-2Rα receptor interaction which binds to IL-2. J. Am. Chem. Soc. 119, 7589–7590 (1997). References 54 and 55 describe an early and well-characterized small-molecule inhibitor of a protein–protein interaction.
Arkin, M. R. et al. Binding of small molecules to an adaptive protein:protein interface. Proc. Natl Acad. Sci. USA 100, 1603–1608 (2003).
Braisted, A. C. et al. Discovery of a potent small molecule IL-2 inhibitor through fragment assembly. J. Am. Chem. Soc. 125, 3714–3715 (2003).
Hyde, J., Braisted, A. C., Randal, M. & Arkin, M. R. Discovery and characterization of cooperative ligand binding in the adaptive region of interleukin-2. Biochemistry 42, 6475–6483 (2003).
Raimundo, B. C. et al. Discovery of small-molecule inhibitors of interleukin-2 using fragment assembly. J. Med. Chem. (in the press). References 56–59 (and 62 below) detail an interdisciplinary approach, integrating fragment-based discovery, medicinal chemistry, and structural biology, to discovering inhibitors of the IL-2/IL-2R interaction.
McKay, D. B. Unraveling the structure of interleukin-2: reply. Science 257, 412–413 (1992).
Mott, H. R. et al. The solution structure of the F42A mutant of human interleukin 2. J. Mol. Biol. 247, 979–994 (1995).
Thanos, C., Randal, M. & Wells, J. A. Potent small-molecule binding to a dynamic hot spot on IL-2. J. Am. Chem. Soc. 125, 15280–15281 (2003).
Harding, F. A., McArthur, J. G., Gross, J. A., Raulet, D. H. & Allison, J. P. CD28-mediated signalling co-stimulates murine T cells and prevents induction of anergy in T-cell clones. Nature 356, 607–609 (1992).
Walunas, T. L. et al. CTLA-4 can function as a negative regulator of T cell activation. Immunity 1, 405–413 (1994).
van der Merwe, P. A. & Davis, S. J. Molecular interactions mediating T cell antigen recognition. Annu. Rev. Immunol. 21, 659–684 (2003).
Collins, A. V. et al. The interaction properties of costimulatory molecules revisited. Immunity 17, 201–210 (2002).
Egen, J. G., Kuhns, M. S. & Allison, J. P. CTLA-4: new insights into its biological function and use in tumor immunotherapy. Nature Immunol. 3, 611–618 (2002).
Darlington, P. J. et al. Surface cytotoxic T lymphocyte-associated antigen 4 partitions within lipid rafts and relocates to the immunological synapse under conditions of inhibition of T cell activation. J. Exp. Med. 195, 1337–1347 (2002).
Stamper, C. C. et al. Crystal structure of the B7-1/CTLA-4 complex that inhibits human immune responses. Nature 410, 608–611 (2001).
Ikemizu, S. et al. Structure and dimerization of a soluble form of B7-1. Immunity 12, 51–60 (2000).
Erbe, D. V., Wang, S., Xing, Y. & Tobin, J. F. Small molecule ligands define a binding site on the immune regulatory protein B7. 1. J. Biol. Chem. 277, 7363–7368 (2002).
Green, N. J. et al. Structure-activity studies of a series of dipyrazolo[3,4-b:3',4'-d]pyridin-3-ones binding to the immune regulatory protein B7.1. Bioorg. Med. Chem. 11, 2991–3013 (2003). References 71 and 72 report the chemical SAR and biochemical characterization of small-molecule inhibitors of B7-1.
Wang, S. et al. Antibodies to B7. 1 define the GFCC'C' face of the N-terminal domain as critical for co-stimulatory interactions. Immunol. Lett. 83, 77–83 (2002).
Huang, Z. The chemical biology of apoptosis. Exploring protein–protein interactions and the life and death of cells with small molecules. Chem. Biol. 9, 1059–1072 (2002). Review of small-molecule inhibitors of the Bcls and other targets in the apoptotic pathway.
Burlacu, A. Regulation of apoptosis by Bcl-2 family proteins. J. Cell. Mol. Med. 7, 249–257 (2003).
Gross, A., McDonnell, J. M. & Korsmeyer, S. J. BCL-2 family members and the mitochondria in apoptosis. Genes Dev. 13, 1899–1911 (1999).
Cory, S. & Adams, J. M. The Bcl2 family: regulators of the cellular life-or-death switch. Nature Rev. Cancer 2, 647–656 (2002).
Huang, D. C. & Strasser, A. BH3-only proteins — essential initiators of apoptotic cell death. Cell 103, 839–842 (2000).
Sattler, M. et al. Structure of Bcl-XL-Bak peptide complex: recognition between regulators of apoptosis. Science 275, 983–986 (1997).
Kim, K. M. et al. Biophysical characterization of recombinant human Bcl-2 and its interactions with an inhibitory ligand, antimycin A. Biochemistry 40, 4911–4922 (2001).
Wang, J. L. et al. Structure-based discovery of an organic compound that binds Bcl-2 protein and induces apoptosis of tumor cells. Proc. Natl Acad. Sci. USA 97, 7124–7129 (2000).
Degterev, A. et al. Identification of small-molecule inhibitors of interaction between the BH3 domain and Bcl-XL . Nature Cell Biol. 3, 173–182 (2001).
Tzung, S. P. et al. Antimycin A mimics a cell-death-inducing Bcl-2 homology domain 3. Nature Cell Biol. 3, 183–191 (2001).
Enyedy, I. J. et al. Discovery of small-molecule inhibitors of Bcl-2 through structure-based computer screening. J. Med. Chem. 44, 4313–4124 (2001).
Lugovskoy, A. A. et al. A novel approach for characterizing protein ligand complexes: molecular basis for specificity of small-molecule Bcl-2 inhibitors. J. Am. Chem. Soc. 124, 1234–1240 (2002).
Kutzki, O. et al. Development of a potent Bcl-XL antagonist based on α-helix mimicry. J. Am. Chem. Soc. 124, 11838–11839 (2002).
Jahnke, W. et al. Second-site NMR screening and linker design. Curr. Top. Med. Chem. 3, 69–80 (2003).
Jahnke, W. et al. Second-site NMR screening with a spin-labeled first ligand. J. Am. Chem. Soc. 122, 7394–7395 (2000). References 84–88 outline a range of NMR-based methods to characterize the binding of small-molecule inhibitors of BCL-X L.
Vassilev, L. T. et al. In vivo activation of the p53 pathway by small-molecule antagonists of MDM2. Science 303, 844–848 (2004). This report describes the activity of 'Nutlins,' small molecules that bind in the p53-helix-binding groove of MDM2. The X-ray crystal structure of Nutlin-2 bound to MDM2 shows the compound bound analogously to p53; compounds were also shown to be active in cell-based and xenograft models of cancer.
Zhao, J. et al. The initial evaluation of non-peptidic small-molecule HDM2 inhibitors based on p53-HDM2 complex structure. Cancer Lett. 183, 69–77 (2002).
Stoll, R. et al. Chalcone derivatives antagonize interactions between the human oncoprotein MDM2 and p53. Biochemistry 40, 336–344 (2001).
Duncan, S. J. et al. Isolation and structure elucidation of Chlorofusin, a novel p53-MDM2 antagonist from a Fusarium sp. J. Am. Chem. Soc. 123, 554–560 (2001).
Yip, C. C. & Ottensmeyer, P. Three-dimensional structural interactions of insulin and its receptor. J. Biol. Chem. 278, 27329–27332 (2003).
Shimaoka, M. & Springer, T. A. Therapeutic antagonists and conformational regulation of integrin function. Nature Rev. Drug Discov. 2, 703–716 (2003). Reviews the small-molecule inhibitors of integrins. Describes how I-domain antagonists regulate ICAM1 binding and provides a clear summary of the authors' model for the mechanism of 'α/β-allosteric inhibitors' (see reference 112).
Wiesmann, C. & de Vos, A. M. Nerve growth factor: structure and function. Cell. Mol. Life Sci. 58, 748–759 (2001).
Alderton, W. K., Cooper, C. E. & Knowles, R. G. Nitric oxide synthases: structure, function and inhibition. Biochem J. 357, 593–615 (2001).
Humphries, M. J. Integrin structure. Biochem. Soc. Trans. 28, 311–339 (2000).
Yusuf-Makagiansar, H., Anderson, M. E., Yakovleva, T. V., Murray, J. S. & Siahaan, T. J. Inhibition of LFA-1/ICAM-1 and VLA-4/VCAM-1 as a therapeutic approach to inflammation and autoimmune diseases. Med. Res. Rev. 22, 146–167 (2002).
Shimaoka, M. et al. Structures of the α L I domain and its complex with ICAM-1 reveal a shape-shifting pathway for integrin regulation. Cell 112, 99–111 (2003).
Randi, A. M. & Hogg, N. I domain of β 2 integrin lymphocyte function-associated antigen-1 contains a binding site for ligand intercellular adhesion molecule-1. J. Biol. Chem. 269, 12395–12398 (1994).
Liu, G. Small molecule antagonists of the LFA-1/ICAM-1 interaction as potential therapeutic agents. Expert Opin. Ther. Patents 11, 1383–1393 (2001).
Sanfilippo, P. J. et al. Novel thiazole based heterocycles as inhibitors of LFA-1/ICAM-1 mediated cell adhesion. J. Med. Chem. 38, 1057–1059 (1995).
Hogg, N., Laschinger, M., Giles, K. & McDowall, A. T-cell integrins: more than just sticking points. J. Cell Sci. 116, 4695–4705 (2003).
Carman, C. V. & Springer, T. A. Integrin avidity regulation: are changes in affinity and conformation underemphasized? Curr. Opin. Cell Biol. 15, 547–556 (2003).
Huth, J. R. et al. NMR and mutagenesis evidence for an I domain allosteric site that regulates lymphocyte function-associated antigen 1 ligand binding. Proc. Natl Acad. Sci. USA 97, 5231–5236 (2000).
Lupher, M. L. Jr et al. Cellular activation of leukocyte function-associated antigen-1 and its affinity are regulated at the I domain allosteric site. J. Immunol. 167, 1431–1439 (2001).
Lu, C., Shimaoka, M., Zang, Q., Takagi, J. & Springer, T. A. Locking in alternate conformations of the integrin αLβ2 I domain with disulfide bonds reveals functional relationships among integrin domains. Proc. Natl Acad. Sci. USA 98, 2393–2398 (2001).
Kallen, J. et al. Structural basis for LFA-1 inhibition upon lovastatin binding to the CD11a I-domain. J. Mol. Biol. 292, 1–9 (1999). Presents the only published X-ray crystal structure of an LFA1 antagonist bound to the I-domain. This research group has used a variety of methods to demonstrate how I-domain inhibitors function and to show their biological relevance.
Last-Barney, K. et al. Binding site elucidation of hydantoin-based antagonists of LFA-1 using multidisciplinary technologies: evidence for the allosteric inhibition of a protein–protein interaction. J. Am. Chem. Soc. 123, 5643–5650 (2001).
Liu, G. et al. Novel p-arylthio cinnamides as antagonists of leukocyte function-associated antigen-1/intracellular adhesion molecule-1 interaction. 2. Mechanism of inhibition and structure-based improvement of pharmaceutical properties. J. Med. Chem. 44, 1202–1010 (2001).
Gadek, T. R. et al. Generation of an LFA-1 antagonist by the transfer of the ICAM-1 immunoregulatory epitope to a small molecule. Science 295, 1086–1089 (2002).
Shimaoka, M., Salas, A., Yang, W., Weitz-Schmidt, G. & Springer, T. A. Small molecule integrin antagonists that bind to the β2 subunit I-like domain and activate signals in one direction and block them in the other. Immunity 19, 391–402 (2003).
Welzenbach, K., Hommel, U. & Weitz-Schmidt, G. Small molecule inhibitors induce conformational changes in the I domain and the I-like domain of lymphocyte function-associated antigen-1. Molecular insights into integrin inhibition. J. Biol. Chem. 277, 10590–10598 (2002).
Weitz-Schmidt, G. et al. Statins selectively inhibit leukocyte function antigen-1 by binding to a novel regulatory integrin site. Nature Med. 7, 687–692 (2001).
Wattanasin, S. et al. 1,4-Diazepane-2-ones as novel inhibitors of LFA-1. Bioorg. Med. Chem. Lett 13, 499–502 (2003).
Kelly, T. A. et al. Cutting edge: a small molecule antagonist of LFA-1-mediated cell adhesion. J. Immunol. 163, 5173–5177 (1999). The series of compounds first described in this paper have been used to elucidate mechanisms of LFA1 function and have led to drug development candidates.
Liu, G. et al. Discovery of novel p-arylthio cinnamides as antagonists of leukocyte function-associated antigen-1/intracellular adhesion molecule-1 interaction. 1. Identification of an additional binding pocket based on an anilino diaryl sulfide lead. J. Med. Chem. 43, 4025–4040 (2000).
Pei, Z. et al. Discovery of potent antagonists of leukocyte function-associated antigen-1/intercellular adhesion molecule-1 interaction. 3. Amide (C-ring) structure-activity relationship and improvement of overall properties of arylthio cinnamides. J. Med. Chem. 44, 2913–2920 (2001).
Winn, M. et al. Discovery of novel p-arylthio cinnamides as antagonists of leukocyte function-associated antigen-1/intercellular adhesion molecule-1 interaction. 4. Structure-activity relationship of substituents on the benzene ring of the cinnamide. J. Med. Chem. 44, 4393–4403 (2001). The compound series described in this paper and previous papers in this series (references 117 and 118) led to a clinical candidate for psoriasis.
Link, J. T. et al. Discovery and SAR of diarylsulfide cyclopropylamide LFA-1/ICAM-1 interaction antagonists. Bioorg. Med. Chem. Lett. 11, 973–976 (2001).
Legge, G. B. et al. NMR solution structure of the inserted domain of human leukocyte function associated antigen-1. J. Mol. Biol. 295, 1251–1264 (2000).
Qu, A. & Leahy, D. J. Crystal structure of the I-domain from the CD11a/CD18 (LFA-1, α L β 2) integrin. Proc. Natl Acad. Sci. USA 92, 10277–10281 (1995).
Qu, A. & Leahy, D. J. The role of the divalent cation in the structure of the I domain from the CD11a/CD18 integrin. Structure 4, 931–942 (1996).
Davidson, W. et al. Characterization of the allosteric inhibition of a protein–protein interaction by mass spectrometry. J. Am. Soc. Mass. Spectrom. 14, 8–13 (2003).
Woska, J. R. Jr et al. A small-molecule antagonist of LFA-1 blocks a conformational change important for LFA-1 function. J. Leukoc. Biol. 70, 329–334 (2001).
Woska, J. R. Jr et al. Small molecule LFA-1 antagonists compete with an anti-LFA-1 monoclonal antibody for binding to the CD11a I domain: development of a flow-cytometry-based receptor occupancy assay. J. Immunol. Methods 277, 101–115 (2003).
Burdick, D. J. et al. N-Benzoyl amino acids as LFA-1/ICAM inhibitors 1: amino acid structure-activity relationship. Bioorg. Med. Chem. Lett 13, 1015–1018 (2003).
Thibault, G. Sodium dodecyl sulfate-stable complexes of echistatin and RGD-dependent integrins: a novel approach to study integrins. Mol. Pharmacol. 58, 1137–1145 (2000).
Ignarro, L. J., Cirino, G., Casini, A. & Napoli, C. Nitric oxide as a signaling molecule in the vascular system: an overview. J. Cardiovasc. Pharmacol. 34, 879–886 (1999).
Gyurko, R., Leupen, S. & Huang, P. L. Deletion of exon 6 of the neuronal nitric oxide synthase gene in mice results in hypogonadism and infertility. Endocrinology 143, 2767–2774 (2002).
Huang, Z. et al. Effects of cerebral ischemia in mice deficient in neuronal nitric oxide synthase. Science 265, 1883–1885 (1994).
Nathan, C. Inducible nitric oxide synthase: what difference does it make? J. Clin. Invest. 100, 2417–2423 (1997).
Bogdan, C. Nitric oxide and the immune response. Nature Immunol. 2, 907–916 (2001).
Vallance, P. & Leiper, J. Blocking NO synthesis: how, where and why? Nature Rev. Drug Discov. 1, 939–950 (2002).
Mete, A. & Connolly, S. Inhibitors of the NOS enzymes: a patent review. IDrugs 6, 57–65 (2003).
Lirk, P., Hoffmann, G. & Rieder, J. Inducible nitric oxide synthase — time for reappraisal. Curr. Drug. Targets Inflamm. Allergy 1, 89–108 (2002).
McMillan, K. et al. Allosteric inhibitors of inducible nitric oxide synthase dimerization discovered via combinatorial chemistry. Proc. Natl Acad. Sci. USA 97, 1506–1511 (2000). Presents the biochemical and structural characterization of a potent inhibitor of iNOS dimerization. Pyridines are known ligands for the haem cofactor in iNOS, but previous members have not been characterized as allosteric dimerization inhibitors.
Ohtsuka, M. et al. PPA250 [3-(2,4-difluorophenyl)-6-[2-[4-(1H-imidazol-1-ylmethyl) phenoxy]ethoxy]-2-phenylpyridine], a novel orally effective inhibitor of the dimerization of inducible nitric-oxide synthase, exhibits an anti-inflammatory effect in animal models of chronic arthritis. J. Pharmacol. Exp. Ther. 303, 52–57 (2002).
Blasko, E. et al. Mechanistic studies with potent and selective inducible nitric-oxide synthase dimerization inhibitors. J. Biol. Chem. 277, 295–302 (2002).
Enkhbaatar, P. et al. Inducible nitric oxide synthase dimerization inhibitor prevents cardiovascular and renal morbidity in sheep with combined burn and smoke inhalation injury. Am. J. Physiol. Heart Circ. Physiol. 285, H2430–H2436 (2003).
Enkhbaatar, P. et al. The inducible nitric oxide synthase inhibitor BBS-2 prevents acute lung injury in sheep after burn and smoke inhalation injury. Am. J. Respir. Crit. Care Med. 167, 1021–1026 (2003).
Szabolcs, M. J. et al. Effects of selective inhibitors of nitric oxide synthase-2 dimerization on acute cardiac allograft rejection. Circulation 106, 2392–2396 (2002).
Chao, M. et al. Neurotrophin receptors: mediators of life and death. Brain Res. Brain Res. Rev. 26, 295–301 (1998).
Ibanez, C. F. et al. Disruption of the low affinity receptor-binding site in NGF allows neuronal survival and differentiation by binding to the trk gene product. Cell 69, 329–341 (1992).
Ryden, M. & Ibanez, C. F. A second determinant of binding to the p75 neurotrophin receptor revealed by alanine-scanning mutagenesis of a conserved loop in nerve growth factor. J. Biol. Chem. 272, 33085–33091 (1997).
Wiesmann, C., Ultsch, M. H., Bass, S. H. & de Vos, A. M. Crystal structure of nerve growth factor in complex with the ligand-binding domain of the TrkA receptor. Nature 401, 184–188 (1999).
Niederhauser, O. et al. NGF ligand alters NGF signaling via p75(NTR) and trkA. J. Neurosci. Res. 61, 263–272 (2000). In characterizing an inhibitor of nerve growth factor, the authors use analytical ultracentrifugation to determine the binding partner and to estimate binding stoichiometry.
Lee, A., Huang, L. & Ellman, J. A. General solid-phase method for the preparation of mechanism-based cysteine protease inhibitors. J. Am. Chem. Soc. 121, 9907–9914 (1999).
Otto, H. -H. & Schirmeister, T. Cysteine proteases and their inhibitors. Chem. Rev. 97, 133–172 (1997).
Tsai, C. J., Lin, S. L., Wolfson, H. J. & Nussinov, R. Protein–protein interfaces: architectures and interactions in protein–protein interfaces and in protein cores. Their similarities and differences. Crit. Rev. Biochem. Mol. Biol. 31, 127–152 (1996).
Stites, W. E. Protein–protein interactions: interface structure, binding thermodynamics, and mutational analysis. Chem. Rev. 97, 1233–1250 (1997).
Hu, Z., Ma, B., Wolfson, H. & Nussinov, R. Conservation of polar residues as hot spots at protein interfaces. Proteins 39, 331–342 (2000).
Atwell, S., Ultsch, M., De Vos, A. M. & Wells, J. A. Structural plasticity in a remodeled protein–protein interface. Science 278, 1125–1128 (1997).
Shuker, S. B., Hajduk, P. J., Meadows, R. P. & Fesik, S. W. Discovering high-affinity ligands for proteins: SAR by NMR. Science 274, 1531–1534 (1996). The first paper describing the SAR by NMR technique used by this laboratory and one of the first demonstrations of fragment-based drug discovery.
Erlanson, D. A. et al. Site-directed ligand discovery. Proc. Natl Acad. Sci. USA 97, 9367–9372 (2000).
Hann, M. M., Leach, A. R. & Harper, G. Molecular complexity and its impact on the probability of finding leads for drug discovery. J. Chem. Inf. Comput. Sci. 41, 856–864 (2001).
Oprea, T. I., Davis, A. M., Teague, S. J. & Leeson, P. D. Is there a difference between leads and drugs? A historical perspective. J. Chem. Inf. Comput. Sci. 41, 1308–1315 (2001).
Lepre, C. A. et al. Applications of SHAPES screening in drug discovery. Comb. Chem. High. Throughput Screen. 5, 583–590 (2002).
Fejzo, J. et al. The SHAPES strategy: an NMR-based approach for lead generation in drug discovery. Chem. Biol. 6, 755–769 (1999).
Hajduk, P. J., Meadows, R. P. & Fesik, S. W. NMR-based screening in drug discovery. Q. Rev. Biophys. 32, 211–240 (1999).
Hajduk, P. J. et al. High-throughput nuclear magnetic resonance-based screening. J. Med. Chem. 42, 2315–2317 (1999).
Hajduk, P. J. et al. Design of adenosine kinase inhibitors from the NMR-based screening of fragments. J. Med. Chem. 43, 4781–4786 (2000).
Nienaber, V. L. et al. Discovering novel ligands for macromolecules using X-ray crystallographic screening. Nature Biotechnol. 18, 1105–1108 (2000).
Carr, R. & Jhoti, H. Structure-based screening of low-affinity compounds. Drug Discov. Today 7, 522–527 (2002).
Erlanson, D. A. et al. Discovery of a new phosphotyrosine mimetic for PTP1B using breakaway tethering. J. Am. Chem. Soc. 125, 5602–5603 (2003).
Erlanson, D. A. et al. In situ assembly of enzyme inhibitors using extended tethering. Nature Biotechnol. 21, 308–314 (2003).
Lebowitz, J., Lewis, M. S. & Schuck, P. Modern analytical ultracentrifugation in protein science: a tutorial review. Protein. Sci. 11, 2067–2079 (2002).
Philo, J. S. Sedimentation equilibrium analysis of mixed associations using numberical contraints to impose mass or signal conservation. Meth. Enzymol. 321, 100–120 (2000).
Arkin, M. & Lear, J. D. A new data analysis method to determine binding constants of small molecules to proteins using equilibrium analytical ultracentrifugation with absorption optics. Anal. Biochem. 299, 98–107 (2001).
Mayer, M. & Meyer, B. Group epitope mapping by saturation transfer difference NMR to identify segments of a ligand in direct contact with a protein receptor. J. Am. Chem. Soc. 123, 6108–6117 (2001).
Hajduk, P. J., Olejniczak, E. T. & Fesik, S. W. One-Dimensional relaxtion- and diffusion-edited NMR methods for screening compounds that bind to macromolecules. J. Am. Chem. Soc. 119, 12257–12261 (1997).
Campbell, A. P. & Sykes, B. D. The two-dimensional transferred nuclear Overhauser effect: theory and practice. Annu. Rev. Biophys. Biomol. Struct. 22, 99–122 (1993).
Day, Y. S. & Myszka, D. G. Characterizing a drug's primary binding site on albumin. J. Pharm. Sci. 92, 333–343 (2003).
Rich, R. L. & Myszka, D. G. Survey of the year 2000 commercial optical biosensor literature. J. Mol. Recognit. 14, 273–294 (2001).
Leavitt, S. & Freire, E. Direct measurement of protein binding energetics by isothermal titration calorimetry. Curr. Opin. Struct. Biol. 11, 560–566 (2001).
Hajduk, P. J. et al. High-throughput nuclear magnetic resonance-based screening. J. Med. Chem. 42, 2315–2317 (1999).
Blundell, T. L., Jhoti, H. & Abell, C. High-throughput crystallography for lead discovery in drug design. Nature Rev. Drug Discov. 1, 45–54 (2002).
Acknowledgements
All images derived from X-ray crystallography data were prepared using Pymol (Delano Scientific, Belmont, California, USA).
Author information
Authors and Affiliations
Ethics declarations
Competing interests
M.R.A. and J.A.W. are employed by Sunesis Pharmaceuticals, which uses proprietary target-directed fragment discovery technologies.
Glossary
- ALLOSTERIC PROTEIN
-
A protein containing two or more topologically distinct binding sites that interact functionally with each other.
- SURFACE PLASMON RESONANCE
-
A method for measuring binding interactions between a surface-immobilized molecule and a solution-phase analyte. The technology measures changes in refractive index caused by a change in mass at the surface.
- ANTIGEN-PRESENTING CELL
-
An immune system cell that takes up antigens, processes them, and presents them to T cells, which are then stimulated to mount an immune response.
- APOPTOSIS
-
Programmed cell death.
- ISOTHERMAL CALORIMETRY
-
A method for measuring the change in heat that occurs when a protein–ligand complex is formed. From these data, the thermodynamic parameters for the interaction can be determined.
Rights and permissions
About this article
Cite this article
Arkin, M., Wells, J. Small-molecule inhibitors of protein–protein interactions: progressing towards the dream. Nat Rev Drug Discov 3, 301–317 (2004). https://doi.org/10.1038/nrd1343
Issue Date:
DOI: https://doi.org/10.1038/nrd1343
This article is cited by
-
Structure-based drug design and molecular dynamics studies of an allosteric modulator targeting the protein–protein interaction site of PDK1
Journal of Molecular Modeling (2024)
-
Single-molecule fingerprinting of protein-drug interaction using a funneled biological nanopore
Nature Communications (2023)
-
A cryptic pocket in Ebola VP35 allosterically controls RNA binding
Nature Communications (2022)
-
Comprehensive exploration of chemical space using trisubstituted carboranes
Scientific Reports (2021)
-
SARS-CoV-2 nucleocapsid and Nsp3 binding: an in silico study
Archives of Microbiology (2021)