Key Points
-
Orthologues (genes in different species arising from a common ancestral gene during speciation) are important in drug discovery for establishing assays and animal models, but a wider-ranging phylogenomic view can lend further insight into function. This can be accomplished not only through the detection of conserved functional elements, but also of functional shifts that could shed light on species differences that often adversely affect drug discovery projects.
-
Establishing orthology is best accomplished through full phylogenetic reconstructions, which then also provide a framework for assessing selective pressures that could signal functional shifts. The nature and extent of selection can be estimated, for example, on the basis of ratios of non-synonymous-to-synonymous nucleotide substitutions and evidence of selective sweeps in patterns of polymorphism.
-
Paralogues (homologous genes in the same species arising by duplication) are also important in drug discovery, not only for compiling classes of tractable targets and outlining selectivity issues, but also for the evolutionary relationship of paralogues to pleiotropy (multifunctionality) and functional redundancy of targets, phenomena that are critical to assessing druggability.
-
Pleiotropy and redundancy are in turn related to crosstalk and heteromery, increasingly prominent themes in drug discovery and (along with alternative transcription) sources of combinatoric diversity of function arising from the genome. Such phenomena also indicate the relevance of an evolutionary view of pathways and networks, whose elements can co-evolve in a way that can also be detected by phylogenomic means and further contribute to functional characterization.
-
Putative drug targets may be profitably viewed through a variety of phylogenomic 'property filters' related to evolutionary rates, selective pressures, degree and nature of paralogy, and factors reflecting pleiotropy such as size, breadth of expression and interaction potential.
Abstract
Phylogenomics, which advocates an evolutionary view of genomic data, has been useful in the prediction of protein function, of significant sequence and structural elements, and of protein interactions and other relationships. Although such information is important in characterizing individual pharmacological targets, evolutionary analyses also indicate new ways to view the overall space of gene products in terms of their suitability for therapeutic intervention. This view places increased emphasis on the comprehensive analysis of the evolutionary history of targets, in particular their orthology and paralogy relationships, the rate and nature of evolutionary change they have undergone, and their involvement in evolving pathways and networks.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Eisen, J. A., Kaiser, D. & Myers, R. M. Gastrogenomic delights: a moveable feast. Nature Med. 3, 1076 (1997).
Eisen, J. A. Phylogenomics: improving functional predictions for uncharacterized genes by evolutionary analysis. Genome Res. 8, 163–167 (1998). The first full description of the phylogenomic approach.
Casari, G., Sander, C. & Valencia, A. A method to predict functional residues in proteins. Nature Struct. Biol. 2, 171–178 (1995).
Mirney, L. A. & Gelfand, M. S. Using orthologous and paralogous proteins to identify specificity-determining residues in bacterial transcription factors. J. Mol. Biol. 321, 7–20 (2002).
Eisen, J. A. & Wu, M. Phylogenetic analysis and gene functional predictions: phylogenomics in action. Theor. Popul. Biol. 61, 481–487 (2002).
Hochachka, P. W. & Monge, C. Evolution of human hypoxia tolerance physiology. Adv. Exp. Med. Biol. 475, 25–43 (2000).
Barclay, A. N. Ig-like domains: evolution from simple interaction molecules to sophisticated antigen recognition. Proc. Natl Acad. Sci. USA 96, 14672–14674 (1999).
Jaaro, H., Beck, G., Conticello, S. G. & Fainzilber, M. Evolving better brains: a need for neurotrophins? Trends Neurosci. 24, 79–85 (2001).
Wilson, D. R. Evolutionary epidemiology and manic depression. Br. J. Med. Psychol. 71, 375–395 (1998).
Gammelgaard, A. Evolutionary biology and the concept of disease. Med. Health Care Philos. 3, 109–116 (2000).
Tatusov, R. L. et al. The COG database: new developments in phylogenetic classification of proteins from complete genomes. Nucleic Acids Res. 29, 22–28 (2001).
Gilks, W. R. et al. Modeling the percolation of annotation errors in a database of protein sequences. Bioinformatics 18, 1641–1649 (2002).
Jones, D. T. & Swindells, M. B. Getting the most from PSI-BLAST. Trends Biochem. Sci. 27, 161–164 (2002).
George, R. A. & Heringa, J. Protein domain identification and improved sequence similarity searching using PSI-BLAST. Proteins 48, 672–681 (2002).
Holm, L. & Sander, C. Protein folds and families: sequence and structure alignments. Nucleic Acids Res. 27, 244–247 (1999).
Todd, A. E., Orengo, C. A. & Thornton, J. M. Plasticity of enzyme active sites. Trends Biochem. Sci. 27, 419–426 (2002).
Hou, J., Sims, G. E., Zhang, C. & Kim, S. H. A global representation of the protein fold space. Proc. Natl Acad. Sci. USA 100, 2386–2390 (2003).
Thornton, J. W. & DeSalle, R. A new method to localize and test the significance of incongruence: detecting domain shuffling in the nuclear receptor superfamily. Syst. Biol. 49, 183–201 (2000).
Koski, L. B. & Golding, G. B. The closest BLAST hit is often not the nearest neighbor. J. Mol. Evol. 52, 540–542 (2001).
Liao, D. Concerted evolution: molecular mechanism and biological implications. Am. J. Hum. Genet. 64, 24–30 (1999).
Amadou, C. Evolution of the MHC class I region: the framework hypothesis. Immunogenetics 49, 362–367 (1999).
Swofford, D. L., Olsen, G. J., Waddell, P. J. & Hillis, D. M. in Molecular Systematics (eds Hillis, D. M., Moritz, C. & Mable, B. K.) 407–514 (Sinauer Associates, Sunderland, 1996).
Storm, C. E. & Sonnhammer, E. L. Automated ortholog inference from phylogenetic trees and calculation of orthology reliability. Bioinformatics 18, 92–99 (2002).
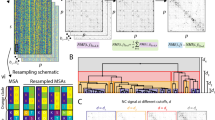
Zmasek, C. M. & Eddy, S. R. Analyzing proteomes by automated phylogenomics using resampled inference of orthologs. BMC Bioinformatics 3, 14 (2002).
Koonin, E. V., Mushegian, A. R. & Bork, P. Non-orthologous gene displacement. Trends Genet. 12, 334–336 (1996).
Brookfield, J. F. What determines the rate of sequence evolution? Curr. Biol. 10, R410–R411 (2000).
Lake, B. G. Coumarin metabolism, toxicity and carcinogenicity: relevance for human risk assessment. Food Chem. Toxicol. 37, 423–453 (1999).
Li, W. -H. Molecular Evolution (Sinauer Associates, Sunderland, 1997).
Messier, W. & Stewart, C. B. Episodic adaptive evolution of primate lysozymes. Nature 385, 151–154 (1997).
Yang, Z. PAML: a program package for phylogenetic analysis by maximum likelihood. Comput. Appl. Biosci. 13, 555–556 (1997).
Benner, S. A. et al. Functional inferences from reconstructed evolutionary biology involving rectified databases — an evolutionarily grounded approach to functional genomics. Res. Microbiol. 151, 97–106 (2000).
Gaucher, E. A. et al. Predicting functional divergence in protein evolution by site-specific rate shifts. Trends Biochem. Sci. 27, 315–321 (2002).
Lopez, P., Casane, D. & Philippe, H. Heterotachy, an important process in protein evolution. Mol. Biol. Evol. 19, 1–7 (2002).
Bamshad, M. & Wooding, S. P. Signatures of natural selection in the human genome. Nature Rev. Genet. 4, 99–111 (2003). An extensive and accessible review of evidence for selection in the human genome.
Smith, J. M. & Haigh, J. The hitch-hiking effect of a favourable gene. Genet. Res. Camb. 23, 23–35 (1974).
Przeworski, M. The signature of positive selection at randomly chosen loci. Genetics 160, 1179–1189 (2002).
de Groot, N. G. et al. Evidence for an ancient selective sweep in the MHC class I gene repertoire of chimpanzees. Proc. Natl Acad. Sci. USA 99, 11748–11753 (2002).
Akey, J. M. et al. Interrogating a high-density SNP map for signatures of natural selection. Genome Res. 12, 1805–1814 (2002).
Enard, W. et al. Molecular evolution of FOXP2, a gene involved in speech and language. Nature 418, 869–872 (2002). Demonstrates the use of measures of selection to suggest a recent functional shift in a gene also associated with an inherited disorder.
DeLisi, L. E. Speech disorder in schizophrenia: review of the literature and exploration of its relation to the uniquely human capacity for language. Schizophr. Bull. 27, 481–496 (2001).
Olson, M. V. & Varki, A. Sequencing the chimpanzee genome: insights into human evolution and disease. Nature Rev. Genet. 4, 20–28 (2003). Makes a strong case for the utility of primate genomes in the study of human disease.
Rockman, M. V. & Wray, G. A. Abundant raw material for cis-regulatory evolution in humans. Mol. Biol. Evol. 19, 1991–2004 (2002).
Akashi, H. Gene expression and molecular evolution. Curr. Opin. Genet. Dev. 11, 660–666 (2001).
Duan, J. et al. Synonymous mutations in the human dopamine receptor D2 (DRD2) affect mRNA stability and synthesis of the receptor. Hum. Mol. Genet. 12, 205–216 (2003).
Hurst, L. D. & Pal, C. Evidence for purifying selection acting on silent sites in BRCA1. Trends Genet. 17, 62–65 (2001).
Durand, D. Vertebrate evolution: doubling and shuffling with a full deck. Trends Genet. 19, 2–5 (2003).
Samonte, R. V. & Eichler, E. E. Segmental duplications and the evolution of the primate genome. Nature Rev. Genet. 3, 65–72 (2002).
Bailey, J. A. et al. Recent segmental duplications in the human genome. Science 297, 1003–1007 (2002).
Friedman, R. & Hughes, A. L. The temporal distribution of gene duplication events in a set of highly conserved human gene families. Mol. Biol. Evol. 20, 154–161 (2003).
Smith G. D. et al. TRPV3 is a temperature-sensitive vanilloid receptor-like protein. Nature 418, 186–190 (2002).
Wise, A. et al. Molecular identification of high and low affinity receptors for nicotinic acid. J. Biol. Chem. 278, 9869–9874 (2003).
Vicker, N. et al. Novel angular benzophenazines: dual topoisomerase I and topoisomerase II inhibitors as potential anticancer agents. J. Med. Chem. 45, 721–739 (2002).
Xia, W. et al. Anti-tumor activity of GW572016: a dual tyrosine kinase inhibitor blocks EGF activation of EGFR/erbB2 and downstream Erk1/2 and AKT pathways. Oncogene 21, 6255–6263 (2002).
Lobell, R. B. et al. Evaluation of farnesyl:protein transferase and geranylgeranyl:protein transferase inhibitor combinations in preclinical models. Cancer Res. 61, 8758–8768 (2001).
Foley, C. L. & Kirby, R. S. 5α-reductase inhibitors: what's new? Curr. Opin. Urol. 13, 31–37 (2003).
Heath, R. J., White, S. W. & Rock, C. O. Lipid biosynthesis as a target for antibacterial agents. Prog. Lipid Res. 40, 467–497 (2001).
Goldstein, J. M. The new generation of antipsychotic drugs: how atypical are they? Int. J. Neuropsychopharmacol. 3, 339–349 (2000).
Hodgkin, J. Seven types of pleiotropy. Int. J. Dev. Biol. 42, 501–505 (1998). A thorough review and catalogue of manifestations of pleiotropy from a genetic perspective.
Jeffery, C. J. Moonlighting proteins. Trends Biochem. Sci. 24, 8–11 (1999).
Copley, S. D. Enzymes with extra talents: moonlighting functions and catalytic promiscuity. Curr. Opin. Chem. Biol. 7, 265–272 (2003).
Wistow, G. & Piatigorsky, J. Recruitment of enzymes as lens structural proteins. Science 236, 1554–1556 (1987).
Citron, B. A. et al. Identity of 4α-carbinolamine dehydratase, a component of the phenylalanine hydroxylation system, and DCoH, a transregulator of homeodomain proteins. Proc. Natl Acad. Sci. USA 89, 11891–11894 (1992).
Sun, Y. J. et al. The crystal structure of a multifunctional protein: phosphoglucose isomerase/autocrine motility factor/neuroleukin. Proc. Natl Acad. Sci. USA 96, 5412–5417 (1999).
Gomez, A., Domedel, N., Cedano, J., Pinol, J. & Querol, E. Do current sequence analysis algorithms disclose multifunctional (moonlighting) proteins? Bioinformatics 19, 895–896 (2003).
Kousteni, S. et al. Nongenotropic, sex-nonspecific signaling through the estrogen or androgen receptors: dissociation from transcriptional activity. Cell 104, 719–730 (2002).
Hughes, A. L. Adaptive evolution after gene duplication. Trends Genet. 18, 433–434 (1994). Suggests that pleiotropy might precede paralogy in the evolution of novel gene function.
Brett, D. et al. Alternative splicing and genome complexity. Nature Genet. 30, 29–30 (2002).
Wagner, A. The role of population size, pleiotropy and fitness effects of mutations in the evolution of overlapping gene functions. Genetics 154, 1389–1401 (2000).
Gu, Z. et al. Role of duplicate genes in genetic robustness against null mutations. Nature 421, 63–66 (2003).
Zhou, F. C., Lesch, K. P. & Murphy, D. L. Serotonin uptake into dopamine neurons via dopamine transporters: a compensatory alternative. Brain Res. 942, 109–119 (2002).
Muoio, D. M. et al. Fatty acid homeostasis and induction of lipid regulatory genes in skeletal muscles of peroxisome proliferator-activated receptor (PPAR)-α knock-out mice. Evidence for compensatory regulation by PPAR-δ. J. Biol. Chem. 277, 26089–26097 (2002).
Troy, C. M. et al. Death in the balance: alternative participation of the caspase-2 and -9 pathways in neuronal death induced by nerve growth factor deprivation. J. Neurosci. 21, 5007–5016 (2001).
Zhang, J. et al. The tissue-specific, compensatory expression of cyclooxygenase-1 and -2 in transgenic mice. Prostaglandins Other Lipid Mediat. 67, 121–135 (2002).
Wang, L. et al. Redundant pathways for negative feedback regulation of bile acid production. Dev. Cell 2, 721–731 (2002).
Mesulam, M. M. et al. Acetylcholinesterase knockouts establish central cholinergic pathways and can use butyrylcholinesterase to hydrolyze acetylcholine. Neuroscience 110, 627–639 (2002).
Haddad, J. J. Cytokines and related receptor-mediated signaling pathways. Biochem. Biophys. Res. Commun. 297, 700–713 (2002).
Dumont, J. E., Pecasse, F. & Maenhaut, C. Crosstalk and specificity in signalling. Are we crosstalking ourselves into general confusion? Cell Signal. 13, 457–463 (2001).
Iwamoto, T. et al. STAT and SMAD signalling in cancer. Histol. Histopathol. 17, 887–895 (2002).
Takayanagi, H. et al. T-cell-mediated regulation of osteoclastogenesis by signalling cross-talk between RANKL and IFN-γ. Nature 408, 600–605 (2000).
Stork, P. J. & Schmitt, J. M. Crosstalk between cAMP and MAP kinase signaling in the regulation of cell proliferation. Trends Cell Biol. 12, 258–266 (2002).
Schwartz, M. A. & Ginsberg, M. H. Networks and crosstalk: integrin signalling spreads. Nature Cell Biol. 4, E65–E68 (2002).
Marshall, F. H. et al. GABAB receptors function as heterodimers. Biochem. Soc. Trans. 27, 530–535 (1999).
Angers, S., Salahpour, A. & Bouvier, M. Biochemical and biophysical demonstration of GPCR oligomerization in mammalian cells. Life Sci. 68, 2243–2250 (2002).
North, R. A. Molecular physiology of P2X receptors. Physiol. Rev. 82, 1013–1067 (2002).
Czirjak, G. & Enyedi, P. Formation of functional heterodimers between the TASK-1 and TASK-3 two-pore domain potassium channel subunits. J. Biol. Chem. 277, 5426–5432 (2002).
Liu, Y. & Eisenberg, D. 3D domain swapping: as domains continue to swap. Protein Sci. 11, 1285–1299 (2002).
Waxman, D. & Peck, J. R. Pleiotropy and the preservation of perfection. Science 279, 1210–1213 (1998).
Galis, F., van Dooren, T. J. & Metz, J. A. Conservation of the segmented germband stage: robustness or pleiotropy? Trends Genet. 18, 504–509 (2002).
Lipman, D. J. et al. The relationship of protein conservation and sequence length. BMC Evol. Biol. 2, 20 (2002).
Duret, L. & Mouchiroud, D. Determinants of substitution rates in mammalian genes: expression pattern affects selection intensity but not mutation rate. Mol. Biol. Evol. 17, 68–74 (2000).
Hastings, K. E. M. Strong evolutionary conservation of broadly expressed protein isoforms in the troponin I gene family and other vertebrate gene families. J. Mol. Evol. 42, 631–640 (1996).
Moskowitz, D. W. Is angiotensin I-converting enzyme a “master” disease gene? Diabetes Technol. Ther. 4, 683–711 (2002).
Viner, J. L., Umar, A. & Hawk, E. T. Chemoprevention of colorectal cancer: problems, progress, and prospects. Gastroenterol. Clin. North Am. 31, 971–999 (2002).
Horowitz, N. H. in Evolving Genes and Proteins (eds Bryson, V. & Vogel, H. J.) 15–23 (Academic Press, New York, 1965).
Belfaiza, J. et al. Evolution of biosynthetic pathways: two enzymes catalyzing consecutive steps in methionine biosynthesis originate from a common ancestor and possess a similar regulatory region. Proc. Natl Acad. Sci. USA 83, 867–871 (1986).
Wilmanns, M. et al. Structural conservation in parallel β/α-barrel enzymes that catalyze three sequential reactions in the pathway of tryptophan biosynthesis. Biochemistry 30, 9161–9169 (1991).
Fani, R., Lio, P., Chiarelli, I. & Bazzicalupo, M. The evolution of the histidine biosynthetic genes in prokaryotes: a common ancestor for the hisA and hisF genes. J. Mol. Evol. 38, 489–495 (1994).
Alves, R., Chaleil, R. A. & Sternberg, M. J. Evolution of enzymes in metabolism: a network perspective. J. Mol. Biol. 320, 751–770 (2002).
Copley, R. R. & Bork, P. Homology among (βα)8 barrels: implications for the evolution of metabolic pathways. J. Mol. Biol. 303, 627–641 (2000).
Forst, C. V. & Schulten, K. Phylogenetic analysis of metabolic pathways. J. Mol. Evol. 52, 471–489 (2001).
Wagner, A. Robustness against mutations in genetic networks of yeast. Nature Genet. 24, 355–361 (2001).
Grange, R. W. et al. Functional and molecular adaptations in skeletal muscle of myoglobin-mutant mice. Am. J. Physiol. Cell Physiol. 281, C1487–C1494 (2001).
de Groof, A. J., Oerlemans, F. T., Jost, C. R. & Wieringa, B. Changes in glycolytic network and mitochondrial design in creatine kinase-deficient muscles. Muscle Nerve 24, 1188–1196 (2001).
Zheng, T. S. et al. Deficiency in caspase-9 or caspase-3 induces compensatory caspase activation. Nature Med. 6, 1241–1247 (2001).
Putcha, G. V. et al. Intrinsic and extrinsic pathway signaling during neuronal apoptosis: lessons from the analysis of mutant mice. J. Cell Biol. 157, 441–453 (2002).
Marcotte, E. M. et al. Detecting protein function and protein–protein interactions from genome sequences. Science 285, 751–753 (1999). Shows that products of genes that fuse in the course of evolution also tend to interact or participate in common pathways in species where they remain unfused.
Pellegrini, M. et al. Assigning protein functions by comparative genome analysis: protein phylogenetic profiles. Proc. Natl Acad. Sci. USA 96, 4285–4288 (1999).
Marcotte, E. M., Xenarios, I., van der Bliek, A. M. & Eisenberg, D. Localizing proteins in the cell from their phylogenetic profiles. Proc. Natl Acad. Sci. USA 97, 12115–12120 (2000).
Goh, C. S. et al. Co-evolution of proteins with their interaction partners. J. Mol. Biol. 299, 283–293 (2000).
Goh, C. S. & Cohen, F. E. Co-evolutionary analysis reveals insights into protein–protein interactions. J. Mol. Biol. 324, 177–192 (2002).
Bafna, V., Hannenhalli, S., Rice, K. & Vawter, L. Ligand-receptor pairing via tree comparison. J. Comput. Biol. 7, 59–70 (2000).
Pazos, F. & Valencia, A. Similarity of phylogenetic trees as indicator of protein–protein interaction. Protein Eng. 14, 609–614 (2001).
Koretke, K. K. et al. Evolution of two-component signal transduction. Mol. Biol. Evol. 17, 1956–1970 (2000).
Fraser, H. B. et al. Evolutionary rate in the protein interaction network. Science 296, 750–752 (2002).
Jordan, I. K., Wolf, Y. I. & Koonin, E. V. No simple dependence between protein evolution rate and the number of protein–protein interactions: only the most prolific interactors tend to evolve slowly. BMC Evol. Biol. 3, 1 (2003).
Fraser, H. B., Wall, D. P. & Hirsh, A. E. A simple dependence between protein evolution rate and the number of protein–protein interactions. BMC Evol. Biol. 3, 11 (2003).
Maslov, S. & Sneppen, K. Specificity and stability in topology of protein networks. Science 296, 910–913 (2002).
Featherstone, D. E. & Broadie, K. Wrestling with pleiotropy: genomic and topological analysis of the yeast expression network. Bioessays 24, 267–274 (2002).
Ohno, S. Evolution by Gene and Genome Duplication (Springer, Berlin, 1970). The classic statement of the theory that duplicated genes are released from selective pressure and are therefore free to rapidly evolve new function.
Wilson, A. C., Carlson, S. S. & White, T. J. Biochemical evolution. Annu. Rev. Biochem. 46, 573–639 (1977).
Jordan, I. K., Rogozin, I. B., Wolf, Y. I. & Koonin, E. V. Essential genes are more evolutionarily conserved than are nonessential genes in bacteria. Genome Res. 12, 962–968 (2002).
Hirsh, A. E. & Fraser, H. B. Protein dispensability and rate of evolution. Nature 411, 1046–1049 (2001).
Pal, C., Papp, B. & Hurst, L. D. Genomic function: rate of evolution and gene dispensability. Nature 421, 496–497 (2003).
Hirsh, A. E. & Fraser, H. B. Genomic function: Rate of evolution and gene dispensability. Nature 421, 497–498 (2003).
Hurst, L. D. & Smith, N. G. C. Do essential genes evolve slowly? Curr. Biol. 9, 747–750 (1999).
Conant, G. C. & Wagner, A. GenomeHistory: a software tool and its application to fully sequenced genomes. Nucleic Acids Res. 30, 3378–3386 (2002).
Schrag, J. D., Winkler, F. K. & Cygler, M. Pancreatic lipases: evolutionary intermediates in a positional change of catalytic carboxylates? J. Biol. Chem. 267, 4300–4303 (1992).
Zhang, J., Dyer, K. D. & Rosenberg, H. F. Evolution of the rodent eosinophil-associated RNase gene family by rapid gene sorting and positive selection. Proc. Natl Acad. Sci. USA 97, 4701–4706 (2000).
Wooding, S. P. et al. DNA sequence variation in a 3.7-kb noncoding sequence 5' of the CYP1A2 gene: implications for human population history and natural selection. Am. J. Hum. Genet. 71, 528–542 (2002).
Altschul, S. F., Gish, W., Miller, W., Myers, E. W. & Lipman, D. J. Basic local alignment search tool. J. Mol. Biol. 215, 403–410 (1990).
Bromham, L. & Penn, D. The modern molecular clock. Nature Rev. Genet. 4, 216–224 (2003).
Mangel, M. & Samaniego, F. J. Abraham Wald's work on aircraft survivability. J. Amer. Statistical Assoc. 79, 259–270 (1984).
Hardison, R. C., Oeltjen, J. & Miller, W. Long human–mouse sequence alignments reveal novel regulatory elements: a reason to sequence the mouse genome. Genome Res. 7, 959–966 (1997).
Wasserman, W. W., Palumbo, M., Thompson, W., Fickett, J. W. & Lawrence, C. E. Human–mouse genome comparisons to locate regulatory sites. Nature Genet. 26, 225–228 (2000).
Bofelli, D. et al. Phylogenetic shadowing of primate sequences to find functional regions of the human genome. Science 299, 1391–1394 (2003).
Fitch, W. M. Distinguishing homologous from analogous proteins. Syst. Zool. 19, 99–113 (1970). The origin of the terms 'orthologue' and 'paralogue'.
Van Valen, L. A new evolutionary law. Evol. Theory 1, 1–30 (1973).
Black, C. G. & Coppel, R. L. Synonymous and non-synonymous mutations in a region of the Plasmodium chabaudi genome and evidence for selection acting on a malaria vaccine candidate. Mol. Biochem. Parasitol. 111, 447–451 (2000).
Woolhouse, M. E., Webster, J. P., Domingo, E., Charlesworth, B. & Levin, B. R. Biological and biomedical implications of the co-evolution of pathogens and their hosts. Nature Genet. 32, 569–577 (2002).
Enard, W. et al. Intra- and interspecific variation in primate gene expression patterns. Science 296, 340–343 (2002). Introduces the notion of phylogenetic analysis of overall gene expression patterns.
Tavazoie, S. et al. Systematic determination of genetic network architecture. Nature Genet. 22, 281–285 (1999).
Wang, Y., Schnegelsberg, P. N., Dausman, J. & Jaenisch, R. Functional redundancy of the muscle-specific transcription factors Myf5 and myogenin. Nature 379, 823–825 (1996).
Tong, A. H. et al. A combined experimental and computational strategy to define protein interaction networks for peptide recognition modules. Science 295, 321–324 (2002).
Ajay, A., Walters, W. P. & Murcko M. A. Can we learn to distinguish between “drug-like” and “nondrug-like” molecules? J. Med. Chem. 41, 3314–3324 (1998).
Muegge, I., Heald, S. L. & Brittelli, D. Simple selection criteria for drug-like chemical matter. J. Med. Chem. 44, 1841–1846 (2001).
Lipinski, C. A., Lombardo, F., Dominy, B. W. & Feeney, P. J. Experimental and computational approaches to estimate solubility and permeability in drug discovery and development settings. Adv. Drug Deliv. Rev. 23, 4–25 (1997).
Veber, D. F. et al. Molecular properties that influence oral bioavailability of drug candidates. J. Med. Chem. 45, 2615–2623 (2002).
Hopkins, A. L. & Groom, C. R. The druggable genome. Nature Rev. Drug Discov. 1, 727–730 (2002). An influential review that helps establish a view of targets as having measurable properties (their drug-binding domain content) making them generally suitable for therapeutic intervention.
Acknowledgements
The author thanks J. R. Brown, K. Rice, and N. Odendahl for many helpful comments on the manuscript.
Author information
Authors and Affiliations
Related links
Related links
DATABASE
LocusLink
FURTHER INFORMATION
PHYLogeny Inference Package (PHYLIP)
Phylogenetic Analysis Using Parsimony (PAUP)
Glossary
- PHYLOGENOMICS
-
The application to genomics of principles and techniques from evolutionary biology, to achieve a better understanding of gene function. 'Pharmacophylogenomics' is the use of phylogenomics in aid of drug discovery, through improved target selection and validation.
- HOMOLOGUES
-
Genes that are similar by virtue of having derived from the same ancestral gene. The similarity might be evident in the DNA sequences of the genes, or in the sequence and/or structure of the gene products. Similarity does not guarantee homology, as unrelated sequences can undergo convergent evolution.
- ORTHOLOGUES
-
Homologous genes in different species arising from a common ancestral gene at the time of speciation (Box 2). Orthology does not guarantee common function, as function can change over time and vary in different evolutionary lineages.
- PARALOGUES
-
Homologous genes in the same species arising by duplication (Box 2).
- PHYLOGENETIC RECONSTRUCTION
-
The attempt to recreate the evolutionary history of a set of orthologues and/or paralogues (or, more generally, any set of measurable characters) and portray it in tree form. A number of different methods and algorithms are used for this purpose, and are the subject of much technical debate, but in the final analysis certainty as to ancestral forms is not possible.
- PLEIOTROPY
-
The property of a gene or gene product by which it exhibits multiple phenotypic effects or possesses multiple functions.
- REDUNDANCY
-
The property by which more than one gene or gene product is able to produce a given phenotype or function.
- BLAST
-
Basic Local Alignment Search Tool, the most widely used bioinformatics algorithm130. It efficiently searches sequence databases for the entries most similar to a query sequence. Recent, more advanced, versions and related tools are specially adapted to finding distant homologues, for which sequence similarity is not obvious but typically some structural similarity is retained.
- INCONGRUENT EVOLUTION
-
Apparent topological differences in the phylogenetic trees of individual genes relative to that of the species, or of individual domains or regions within genes relative to each other. This can arise from phenomena such as domain shuffling or horizontal transmission of genes between species.
- CONCERTED EVOLUTION
-
Greater-than-expected similarity seen in members of gene families within a species relative to that seen between species. This can arise from phenomena related to physical mechanisms of replication and recombination that tend to maintain uniformity between (often tandem) copies.
- SYNTENY
-
The property of genes of being found on the same chromosome. The ordering of orthologues on chromosomes is often conserved between related species over extended segments, indicating a common ancestry of those segments; this phenomenon is referred to as conservation of synteny. (To describe the orthologues or regions of the different species as being syntenic to each other is a common misuse of the term.)
- MOLECULAR CLOCK
-
The hypothesis that, except for the effects of functional constraints on gene products, sequence substitutions occur at a constant rate on an evolutionary timescale. It is closely tied to the 'neutral theory' of evolution, which asserts that most such mutations are selectively neutral and driven only by random drift. Although subject to certain caveats and continuing debate, the notion of the molecular clock has proven to be an important and useful tool in many contexts131.
- NON-SYNONYMOUS SUBSTITUTION
-
A nucleotide substitution that results in an amino acid change.
- SYNONYMOUS SUBSTITUTION
-
A 'silent' nucleotide substitution, often in the third codon position, that does not result in an amino acid change.
- GENE SHARING (RECRUITMENT)
-
An adaptation of a gene to serve an additional unrelated function, generally in a different tissue and presumably by the incorporation of alternative regulatory elements at the same locus. It is one proposed mechanism for establishing pleiotropy.
- CROSSTALK
-
The interaction of elements of distinct signalling or regulatory pathways such that an input to one pathway has some effect on the output of the other.
- HETEROMERY
-
The physical association of distinct but often similar macromolecules, as when a pair of protein subunits combine to form a heterodimer. A combination of identical subunits is called homomery.
- DOMAIN SWAPPING
-
The symmetric exchange of portions of polypeptides (ranging up to entire domains), by partial unfolding, between subunits of a multimeric (usually dimeric) assemblage, such that the exchanged portions occupy positions in their counterpart subunits analogous to those they would assume in the monomers.
Rights and permissions
About this article
Cite this article
Searls, D. Pharmacophylogenomics: genes, evolution and drug targets. Nat Rev Drug Discov 2, 613–623 (2003). https://doi.org/10.1038/nrd1152
Issue Date:
DOI: https://doi.org/10.1038/nrd1152
This article is cited by
-
Understanding the tree of life: an overview of tree-reading skill frameworks
Evolution: Education and Outreach (2019)
-
Gene tree species tree reconciliation with gene conversion
Journal of Mathematical Biology (2019)
-
Evolutionary Perspectives of Genotype–Phenotype Factors in Leishmania Metabolism
Journal of Molecular Evolution (2018)
-
Reconciled rat and human metabolic networks for comparative toxicogenomics and biomarker predictions
Nature Communications (2017)
-
The evolutionary rate of antibacterial drug targets
BMC Bioinformatics (2013)