Key Points
-
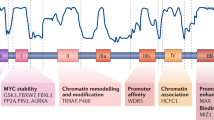
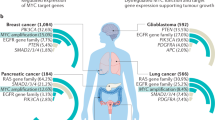
MYC is a pivotal player in normal cells, not only in proliferation but also in apoptosis and differentiation. MYC is deregulated in most tumour types and stages.
-
MYC expression is sometimes an addictive character of cancer and is therefore an Achilles heel for cancer cells. Initial studies indicate that MYC suppression can lead to apoptosis and cell death in cancer cells, while producing only rapidly reversible effects in normal cells, providing a wide therapeutic window for specific and efficacious anti-tumour treatment.
-
Transcriptionally induced negative supercoiling is crucial for controlling MYC expression. The distance that this negative supercoiling travels upstream from the transcriptional start sites can be large (>1 kb) and is constrained within the boundaries imposed by looping and other features of the chromatin.
-
There are two important elements in the MYC promoter that are dynamically affected by negative supercoiling: far upstream element (FUSE) and NHE III1.
-
FUSE, in concert with the effector proteins FUSE-binding protein (FBP), FBP-interacting repressor (FIR) and TFIIH, constitutes a cruise control system that can precisely control the rate of MYC promoter firing. NHE III1 can exist in three different forms, of which two (duplex and single stranded) constitute an on switch and the third (G-quadruplex and i-motif) constitutes an off switch.
-
There are defined strategies to suppress MYC expression by small-molecule targeting of the FUSE–FBP interface or by small-molecule stabilization of the G-quadruplex or i-motif structure in the on–off switch.
Abstract
MYC is deregulated in most tumour types, but an effective means to selectively target its aberrant expression is not yet available. Supercoiling that is induced by transcription has been demonstrated to have dynamic effects on DNA in the MYC promoter element: it converts duplex DNA to non-duplex DNA structures, even at considerable distances from the transcriptional start site. These non-duplex DNA structures, which control both turning on and off of transcription and the rate of transcription firing, are amenable to small-molecule targeting. This dynamic system provides a unique opportunity for the treatment of tumours in which MYC is an important oncogene.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout








Similar content being viewed by others
References
Hurley, L. H. DNA and associated processes as targets for cancer therapy. Nature Rev. Cancer 2, 188–200 (2002).
Zhang, J., Yang, P. L. & Gray, N. S. Targeting cancer with small molecule kinase inhibitors. Nature Rev. Cancer 9, 28–39 (2009).
Meyer, N. & Penn, L. Z. Reflecting on 25 years with MYC. Nature Rev. Cancer 8, 976–990 (2008). An excellent comprehensive review of the history of MYC from its first discovery through 25 years of data.
Adhikary, S. & Eilers, M. Transcriptional regulation and transformation by Myc proteins. Nature Rev. Mol. Cell Biol. 6, 635–645 (2005).
Dani, C. et al. Extreme instability of myc mRNA in normal and transformed human cells. Proc. Natl. Acad. Sci. USA 81, 7046–7050 (1984).
Hann, S. R., Thompson, C. B. & Eisenman, R. N. c-myc oncogene protein synthesis is independent of the cell cycle in human and avian cells. Nature 314, 366–369 (1985).
Kingston, R. E. Transcription control and differentiation: the HLH family, c-myc and C/EBP. Curr. Opin. Cell Biol. 1, 1081–1087 (1989).
Rabbitts, P. H. et al. Metabolism of c-myc gene products: c-myc mRNA and protein expression in the cell cycle. EMBO J. 4, 2009–2015 (1985).
Ramsay, G., Evan, G. I. & Bishop, J. M. The protein encoded by the human proto-oncogene c-myc. Proc. Natl. Acad. Sci. USA 81, 7742–7746 (1984).
Fernandez, P. C. et al. Genomic targets of the human c-Myc protein. Genes Dev. 17, 1115–1129 (2003).
Li, Z. et al. A global transcriptional regulatory role for c-Myc in Burkitt's lymphoma cells. Proc. Natl. Acad. Sci. USA 100, 8164–8169 (2003).
Zeller, K. I. et al. Global mapping of c-Myc binding sites and target gene networks in human B cells. Proc. Natl. Acad. Sci. USA 103, 17834–17839 (2006).
Patel, J. H., Loboda, A. P., Showe, M. K., Showe, L. C. & McMahon, S. B. Analysis of genomic targets reveals complex functions of MYC. Nature Rev. Cancer 4, 562–568 (2004).
Fisher, F., Jayaraman, P. S. & Goding, C. R. C-myc and the yeast transcription factor PHO4 share a common CACGTG-binding motif. Oncogene 6, 1099–1104 (1991).
Kleine-Kohlbrecher, D., Adhikary, S. & Eilers, M. Mechanisms of transcriptional repression by Myc. Curr. Top. Microbiol. Immunol. 302, 51–62 (2006).
Cole, M. D. & Cowling, V. H. Transcription-independent functions of MYC: regulation of translation and DNA replication. Nature Rev. Mol. Cell Biol. 9, 810–815 (2008).
Morrish, F., Isern, N., Sadilek, M., Jeffrey, M. & Hockenbery, D. M. c-Myc activates multiple metabolic networks to generate substrates for cell-cycle entry. Oncogene 28, 2485–2491 (2009).
Shiue, C. N., Berkson, R. G. & Wright, A. P. c-Myc induces changes in higher order rDNA structure on stimulation of quiescent cells. Oncogene 28, 1833–1842 (2009).
Cheng, T. et al. Hematopoietic stem cell quiescence maintained by p21cip1/waf1. Science 287, 1804–1808 (2000).
Hoffman, B. & Liebermann, D. A. Apoptotic signaling by c-MYC. Oncogene 27, 6462–6472 (2008).
Wierstra, I. & Alves, J. The c-myc promoter: still MysterY and Challenge. Adv. Cancer Res. 99, 113–333 (2008). This article reviews all aspects of transcriptional control of MYC , including all converging pathways.
Lutz, W., Leon, J. & Eilers, M. Contributions of Myc to tumorigenesis. Biochim. Biophys. Acta 1602, 61–71 (2002). Review detailing models of, requirements for, and roles of MYC in tumorigenesis.
Cotter, F. E. & Zucca, E. Altered gene expression and oncogenesis of B-cell neoplasia. Ann. Oncol. 2, 335–342 (1991).
Kato, G. J. & Dang, C. V. Function of the c-Myc oncoprotein. FASEB J. 6, 3065–3072 (1992).
Eilers, M. & Eisenman, R. N. Myc's broad reach. Genes Dev. 22, 2755–2766 (2008).
Vita, M. & Henriksson, M. The Myc oncoprotein as a therapeutic target for human cancer. Semin. Cancer Biol. 16, 318–330 (2006). This paper is a good summary of the roles of MYC in cancer and normal cells. It includes a discussion of all MYC targeting strategies as of 2006.
Kelly, K. & Siebenlist, U. The role of c-myc in the proliferation of normal and neoplastic cells. J. Clin. Immunol. 5, 65–77 (1985).
Rapp, U. R. et al. MYC is a metastasis gene for non-small-cell lung cancer. PLoS One 4, e6029 (2009).
Dadiani, M. et al. Estrogen regulation of VEGF in breast cancer in vitro and in vivo: the role of estrogen receptorα and c-Myc. Endocr. Relat. Cancer 16, 819–834 (2009).
Kokai, E. et al. Myc regulates embryonic vascular permeability and remodeling. Circul. Res. 104, 1151–1159 (2009).
Huang, L. E. Carrot and stick: HIF-α engages c-Myc in hypoxic adaptation. Cell Death Differ. 15, 672–677 (2008).
Marcu, K. B., Bossone, S. A. & Patel, A. J. myc function and regulation. Annu. Rev. Biochem. 61, 809–860 (1992).
Spencer, C. A. & Groudine, M. Control of c-myc regulation in normal and neoplastic cells. Adv. Cancer Res. 56, 1–48 (1991).
D'Cruz, C. M. et al. c-MYC induces mammary tumorigenesis by means of a preferred pathway involving spontaneous Kras2 mutations. Nature Med. 7, 235–239 (2001).
Pelengaris, S., Khan, M. & Evan, G. c-MYC: more than just a matter of life and death. Nature Rev. Cancer 2, 764–776 (2002).
Pelengaris, S., Khan, M. & Evan, G. I. Suppression of Myc-induced apoptosis in β cells exposes multiple oncogenic properties of Myc and triggers carcinogenic progression. Cell 109, 321–334 (2002).
Pelengaris, S., Littlewood, T., Khan, M., Elia, G. & Evan, G. Reversible activation of c-Myc in skin: induction of a complex neoplastic phenotype by a single oncogenic lesion. Mol. Cell 3, 565–577 (1999).
Kellen, J. A. Genomic effects of tamoxifen. Anticancer Res. 16, 3537–3541 (1996).
Mandlekar, S. & Kong, A. N. Mechanisms of tamoxifen-induced apoptosis. Apoptosis 6, 469–477 (2001).
Musgrove, E. A. et al. Identification of functional networks of estrogen- and c-Myc-responsive genes and their relationship to response to tamoxifen therapy in breast cancer. PLoS One 3, e2987 (2008).
Arango, D., Corner, G. A., Wadler, S., Catalano, P. J. & Augenlicht, L. H. c-myc/p53 interaction determines sensitivity of human colon carcinoma cells to 5-fluorouracil in vitro and in vivo. Cancer Res. 61, 4910–4915 (2001).
Weinstein, I. B. & Joe, A. Oncogene addiction. Cancer Res. 68, 3077–3080 (2008). An excellent review of the concept of oncogene addiction.
Soucek, L. et al. Modelling Myc inhibition as a cancer therapy. Nature 455, 679–683 (2008). This paper described the first concrete evidence that a therapeutic window exists for MYC-targeted therapies. It demonstrates the reversibility of all toxicities in normal cells in a short time frame, with the complete regression of developing neoplasias.
Shachaf, C. M. et al. Genomic and proteomic analysis reveals a threshold level of MYC required for tumor maintenance. Cancer Res. 68, 5132–5142 (2008). This paper addresses the question of a threshold for MYC knockdown that may be required for a therapeutic effect. It further highlights a potentially large window for MYC-targeted therapies.
Wu, C. H. et al. Cellular senescence is an important mechanism of tumor regression upon c-Myc inactivation. Proc. Natl. Acad. Sci. USA 104, 13028–13033 (2007).
Postow, L., Hardy, C. D., Arsuaga, J. & Cozzarelli, N. R. Topological domain structure of the Escherichia coli chromosome. Genes Dev. 18, 1766–1779 (2004).
Travers, A. & Muskhelishvili, G. DNA supercoiling—a global transcriptional regulator for enterobacterial growth? Nature Rev. Microbiol. 3, 157–169 (2005).
Dayn, A. et al. Formation of (dA-dT)n cruciforms in Escherichia coli cells under different environmental conditions. J. Bacteriol. 173, 2658–2664 (1991).
Krasilnikov, A. S., Podtelezhnikov, A., Vologodskii, A. & Mirkin, S. M. Large-scale effects of transcriptional DNA supercoiling in vivo. J. Mol. Biol. 292, 1149–1160 (1999).
Dayn, A., Malkhosyan, S. & Mirkin, S. M. Transcriptionally driven cruciform formation in vivo. Nucleic Acids Res. 20, 5991–5997 (1992).
Kouzine, F. & Levens, D. Supercoil-driven DNA structures regulate genetic transactions. Front. Biosci. 12, 4409–4423 (2007). An excellent review of the general area of supercoiling and transcriptional control that is very readable for the non-specialist.
Li, G., Levitus, M., Bustamante, C. & Widom, J. Rapid spontaneous accessibility of nucleosomal DNA. Nature Struct. Mol. Biol. 12, 46–53 (2005).
Bancaud, A. et al. Structural plasticity of single chromatin fibers revealed by torsional manipulation. Nature Struct. Mol. Biol. 13, 444–450 (2006).
Adachi, N. & Lieber, M. R. Bidirectional gene organization: a common architectural feature of the human genome. Cell 109, 807–809 (2002).
Trinklein, N. D. et al. An abundance of bidirectional promoters in the human genome. Genome Res. 14, 62–66 (2004).
Champoux, J. J. DNA topoisomerases: structure, function, and mechanism. Annu. Rev. Biochem. 70, 369–413 (2001).
Wang, Z. & Droge, P. Differential control of transcription-induced and overall DNA supercoiling by eukaryotic topoisomerases in vitro. EMBO J. 15, 581–589 (1996).
Kramer, P. R. & Sinden, R. R. Measurement of unrestrained negative supercoiling and topological domain size in living human cells. Biochemistry 36, 3151–3158 (1997).
Kouzine, F., Sanford, S., Elisha-Feil, Z. & Levens, D. The functional response of upstream DNA to dynamic supercoiling in vivo. Nature Struct. Mol. Biol. 15, 146–154 (2008). This paper provides the in vivo proof that the functional response in the MYC promoter is directly related to transcriptionally induced supercoiling. This paper will become a classic in the regulation of eukaryotic gene expression.
Michelotti, G. A. et al. Multiple single-stranded cis elements are associated with activated chromatin of the human c-myc gene in vivo. Mol. Cell. Biol. 16, 2656–2669 (1996).
Avigan, M. I., Strober, B. & Levens, D. A far upstream element stimulates c-myc expression in undifferentiated leukemia cells. J. Biol. Chem. 265, 18538–18545 (1990).
Bazar, L. et al. Targeted melting and binding of a DNA regulatory element by a transactivator of c-myc. J. Biol. Chem. 270, 8241–8248 (1995).
Sun, D. & Hurley, L. H. The importance of negative superhelicity in inducing the formation of G-quadruplex and i-motif structures in the c-Myc promoter: implications for drug targeting and control of gene expression. J. Med. Chem. 52, 2863–2874 (2009). This paper addresses the question of how the G-quadruplex and i-motifs can form from duplex DNA and how this relates to the control of MYC gene transcription.
Sinden, R. R. DNA Structure and Function (Academic Press, San Diego, 1994).
Wittig, B., Dorbic, T. & Rich, A. Transcription is associated with Z-DNA formation in metabolically active permeabilized mammalian cell nuclei. Proc. Natl. Acad. Sci. USA 88, 2259–2263 (1991).
Liu, J. et al. The FUSE/FBP/FIR/TFIIH system is a molecular machine programming a pulse of c-myc expression. EMBO J. 25, 2119–2130 (2006). An important paper on the time course of events involving the interplay of transcription factors, chromatin and negative supercoiling after serum activation.
Liu, J. et al. Defective interplay of activators and repressors with TFIIH in xeroderma pigmentosum. Cell 104, 353–363 (2001).
Michelotti, E. F., Tomonaga, T., Krutzsch, H. & Levens, D. Cellular nucleic acid binding protein regulates the CT element of the human c-myc protooncogene. J. Biol. Chem. 270, 9494–9499 (1995).
Braddock, D. T., Baber, J. L., Levens, D. & Clore, G. M. Molecular basis of sequence-specific single-stranded DNA recognition by KH domains: solution structure of a complex between hnRNP K KH3 and single-stranded DNA. EMBO J. 21, 3476–3485 (2002).
Braddock, D. T., Louis, J. M., Baber, J. L., Levens, D. & Clore, G. M. Structure and dynamics of KH domains from FBP bound to single-stranded DNA. Nature 415, 1051–1056 (2002).
Michelotti, E. F., Michelotti, G. A., Aronsohn, A. I. & Levens, D. Heterogeneous nuclear ribonucleoprotein K is a transcription factor. Mol. Cell. Biol. 16, 2350–2360 (1996). In vivo identification, using chemical and enzymatic probes, of the elements that will prove to be essential for the control of MYC transcription.
Lynch, M. et al. hnRNP K binds a core polypyrimidine element in the eukaryotic translation initiation factor 4E (eIF4E) promoter, and its regulation of eIF4E contributes to neoplastic transformation. Mol. Cell. Biol. 25, 6436–6453 (2005).
Simonsson, T., Pecinka, P. & Kubista, M. DNA tetraplex formation in the control region of c-myc. Nucleic Acids Res. 26, 1167–1172 (1998).
Siddiqui-Jain, A., Grand, C. L., Bearss, D. J. & Hurley, L. H. Direct evidence for a G-quadruplex in a promoter region and its targeting with a small molecule to repress c-MYC transcription. Proc. Natl. Acad. Sci. USA 99, 11593–11598 (2002). Proof-of-principle evidence that stabilization of the G-quadruplex in the MYC promoter by small molecules causes the inhibition of MYC transcription.
Seenisamy, J. et al. The dynamic character of the G-quadruplex element in the c-MYC promoter and modification by TMPyP4. J. Am. Chem. Soc. 126, 8702–8709 (2004).
Phan, A. T., Modi, Y. S. & Patel, D. J. Propeller-type parallel-stranded G-quadruplexes in the human c-myc promoter. J. Am. Chem. Soc. 126, 8710–8716 (2004).
Ambrus, A., Chen, D., Dai, J., Jones, R. A. & Yang, D. Solution structure of the biologically relevant G-quadruplex element in the human c-MYC promoter. Implications for G-quadruplex stabilization. Biochemistry 44, 2048–2058 (2005).
Kumar, P., Verma, A., Maiti, S., Gargallo, R. & Chowdhury, S. Tetraplex DNA transitions within the human c-myc promoter detected by multivariate curve resolution of fluorescence resonance energy transfer. Biochemistry 44, 16426–16434 (2005).
Freyer, M. W. et al. Biophysical studies of the c-MYC NHE III1 promoter: model quadruplex interactions with a cationic porphyrin. Biophys. J. 92, 2007–2015 (2007).
Gueron, M. & Leroy, J. L. The i-motif in nucleic acids. Curr. Opin. Struct. Biol. 10, 326–331 (2000).
Simonsson, T., Pribylova, M. & Vorlickova, M. A nuclease hypersensitive element in the human c-myc promoter adopts several distinct i-tetraplex structures. Biochem. Biophys. Res. Commun. 278, 158–166 (2000).
Bugaut, A. & Balasubramanian, S. A sequence-independent study of the influence of short loop lengths on the stability and topology of intramolecular DNA G-quadruplexes. Biochemistry 47, 689–697 (2008).
Hazel, P., Huppert, J., Balasubramanian, S. & Neidle, S. Loop-length-dependent folding of G-quadruplexes. J. Am. Chem. Soc. 126, 16405–16415 (2004).
Risitano, A. & Fox, K. R. Influence of loop size on the stability of intramolecular DNA quadruplexes. Nucleic Acids Res. 32, 2598–2606 (2004).
Dexheimer, T. S. et al. NM23-H2 may play an indirect role in transcriptional activation of c-myc gene expression but does not cleave the nuclease hypersensitive element III1 . Mol. Cancer Ther. 8, 1363–1377 (2009). Identification of an enzyme (NM23-H2) that unfolds G-quadruplexes to stimulate MYC transcription.
Thakur, R. K. et al. Metastases suppressor NM23-H2 interaction with G-quadruplex DNA within c-MYC promoter nuclease hypersensitive element induces c-MYC expression. Nucleic Acids Res. 37, 172–183 (2009). This paper provides additional evidence that enzymes able to recognize the MYC G-quadruplex can either unfold or fold this secondary DNA structure.
González, V., Guo, K., Hurley, L. H. & Sun, D. Identification and characterization of nucleolin as a c-MYC G-quadruplex-binding protein. J. Biol. Chem. 284, 23622–23635 (2009). This paper provides experimental evidence for the role of nucleolin in facilitating the formation of the MYC G-quadruplex and suppressing MYC transcription, as well as a potential explanation for how Quarfloxin, a first-in-class G-quadruplex drug in Phase II clinical trials, exerts its effects through nucleolin and the MYC G-quadruplex.
Kang, H. J. & Park, H. J. Novel molecular mechanism for actinomycin D activity as an oncogenic promoter G-quadruplex binder. Biochemistry 48, 7392–7398 (2009).
Ou, T. M. et al. Stabilization of G-quadruplex DNA and down-regulation of oncogene c-myc by quindoline derivatives. J. Med. Chem. 50, 1465–1474 (2007). The first independent confirmation that small molecules that stabilize the MYC G-quadruplex can suppress MYC transcription.
Huth, J. R. et al. NMR-driven discovery of benzoylanthranilic acid inhibitors of far upstream element binding protein binding to the human oncogene c-myc promoter. J. Med. Chem. 47, 4851–4857 (2004). A joint paper from the Levens laboratory and Abbott Pharmaceuticals demonstrating an approach to modulation of MYC gene transcription.
Hanahan, D. & Weinberg, R. A. The hallmarks of cancer. Cell 100, 57–70 (2000).
Cogoi, S. et al. Identification of a new G-quadruplex motif in the KRAS promoter and design of pyrene-modified G4-decoys with antiproliferative activity in pancreatic cancer cells. J. Med. Chem. 52, 564–568 (2009).
Xu, Y. & Sugiyama, H. Formation of the G-quadruplex and i-motif structures in retinoblastoma susceptibility genes (Rb). Nucleic Acids Res. 34, 949–954 (2006).
Dexheimer, T. S., Sun, D. & Hurley, L. H. Deconvoluting the structural and drug-recognition complexity of the G-quadruplex-forming region upstream of the bcl-2 P1 promoter. J. Am. Chem. Soc. 128, 5404–5415 (2006).
Guo, K., Gokhale, V., Hurley, L. H. & Sun, D. Intramolecularly folded G-quadruplex and i-motif structures in the proximal promoter of the vascular endothelial growth factor gene. Nucleic Acids Res. 36, 4598–4608 (2008).
Qin, Y., Rezler, E. M., Gokhale, V., Sun, D. & Hurley, L. H. Characterization of the G-quadruplexes in the duplex nuclease hypersensitive element of the PDGF-A promoter and modulation of PDGF-A promoter activity by TMPyP4. Nucleic Acids Res. 35, 7698–7713 (2007).
Palumbo, S. L., Ebbinghaus, S. W. & Hurley, L. H. Formation of a unique end-to-end stacked pair of G-quadruplexes in the hTERT core promoter with implications for inhibition of telomerase by G-quadruplex-interactive ligands. J. Am. Chem. Soc. 131, 10878–10891 (2009).
Huppert, J. L. & Balasubramanian, S. Prevalence of quadruplexes in the human genome. Nucleic Acids Res. 33, 2908–2916 (2005).
Huppert, J. L. & Balasubramanian, S. G-quadruplexes in promoters throughout the human genome. Nucleic Acids Res. 35, 406–413 (2007). A bioinformatics study showing the enrichment of G-quadruplex-forming sequences clustered around transcriptional start sites.
Verma, A. et al. Genome-wide computational and expression analyses reveal G-quadruplex DNA motifs as conserved cis-regulatory elements in human and related species. J. Med. Chem. 51, 5641–5649 (2008).
Verma, A., Yadav, V. K., Basundra, R., Kumar, A. & Chowdhury, S. Evidence of genome-wide G4 DNA-mediated gene expression in human cancer cells. Nucleic Acids Res. 37, 4194–4204 (2009).
Rawal, P. et al. Genome-wide prediction of G4 DNA as regulatory motifs: role in Escherichia coli global regulation. Genome Res. 16, 644–655 (2006).
Hurley, L. H. DNA and associated targets for drug design. J. Med. Chem. 32, 2027–2033 (1989).
Weber, A. et al. The FUSE binding proteins FBP1 and FBP3 are potential c-myc regulators in renal, but not in prostate and bladder cancer. BMC Cancer 8, 369 (2008).
Drygin, D. et al. Quarfloxin (CX-3543) disrupts the Nucleolin/rDNA quadruplex complexes, inhibits the elongation by RNA Polymerase I and exhibits potent antitumor activity in models of cancer. Proc. Am. Assoc. Cancer Res. Abstr. 3301 (2008).
Bates, P. J., Laber, D. A., Miller, D. M., Thomas, S. D. & Trent, J. O. Discovery and development of the G-rich oligonucleotide AS1411 as a novel treatment for cancer. Exp. Mol. Path. 86, 151–164 (2009).
Bates, P., Mergny, J.-L. & Yang, D. Quartets in G-major. The first international meeting on quadruplex DNA. EMBO Reports 8, 1003–1010 (2007).
Lavelle, C. DNA torsional stress propagates through chromatin fiber and participates in transcriptional regulation. Nature Struct. Mol. Biol. 15, 123–125 (2008).
Gunaratnam, M. et al. G-quadruplex compounds and cis-platin act synergistically to inhibit cancer cell growth in vitro and in vivo. Biochem. Pharmacol. 78, 115–122 (2009).
Phan, A. T., Kuryavyi, V., Burge, S., Neidle, S. & Patel, D. J. Structure of an unprecedented G-quadruplex scaffold in the human c-kit promoter. J. Am. Chem. Soc. 129, 4386–4392 (2007).
Todd, A. K., Haider, S. M., Parkinson, G. N. & Neidle, S. Sequence occurrence and structural uniqueness of a G-quadruplex in the human c-kit promoter. Nucleic Acids Res. 35, 5799–5808 (2007).
Cogoi, S., Paramasivam, M., Spolaore, B. & Xodo, L. E. Structural polymorphism within a regulatory element of the human KRAS promoter: formation of G4-DNA recognized by nuclear proteins. Nucleic Acids Res. 36, 3765–3780 (2008).
Paramasivam, M. et al. Protein hnRNP A1 and its derivative Up1 unfold quadruplex DNA in the human KRAS promoter: implications for transcription. Nucleic Acids Res. 37, 2841–2853 (2009).
Xodo, L., Paramasivam, M., Membrino, A. & Cogoi, S. Protein hnRNPA1 binds to a critical G-rich element of KRAS and unwinds G-quadruplex structures: implications in transcription. Nucleic Acids Symp. Ser. (Oxf.) 52, 159–160 (2008).
Nesbit, C. E., Tersak, J. M. & Prochownik, E. V. MYC oncogenes and human neoplastic disease. Oncogene 18, 3004–3016 (1999).
Nilsson, J. A. & Cleveland, J. L. Myc pathways provoking cell suicide and cancer. Oncogene 22, 9007–9021 (2003).
Oster, S. K., Ho, C. S., Soucie, E. L. & Penn, L. Z. The myc oncogene: MarvelouslY Complex. Adv. Cancer Res. 84, 81–154 (2002).
Pelengaris, S. & Khan, M. The many faces of c-MYC. Arch. Biochem. Biophys. 416, 129–136 (2003).
Felsher, D. W. & Bishop, J. M. Reversible tumorigenesis by MYC in hematopoietic lineages. Mol. Cell 4, 199–207 (1999).
Karlsson, A. et al. Genomically complex lymphomas undergo sustained tumor regression upon MYC inactivation unless they acquire novel chromosomal translocations. Blood 101, 2797–2803 (2003).
Jain, M. et al. Sustained loss of a neoplastic phenotype by brief inactivation of MYC. Science 297, 102–104 (2002).
Pelengaris, S. et al. Brief inactivation of c-Myc is not sufficient for sustained regression of c-Myc-induced tumours of pancreatic islets and skin epidermis. BMC Biol. 2, 26 (2004).
Shachaf, C. M. et al. MYC inactivation uncovers pluripotent differentiation and tumour dormancy in hepatocellular cancer. Nature 431, 1112–1117 (2004).
Boxer, R. B., Jang, J. W., Sintasath, L. & Chodosh, L. A. Lack of sustained regression of c-MYC-induced mammary adenocarcinomas following brief or prolonged MYC inactivation. Cancer Cell 6, 577–586 (2004).
Liu, J. et al. The FBP interacting repressor targets TFIIH to inhibit activated transcription. Mol. Cell 5, 331–341 (2000).
He, L. et al. Loss of FBP function arrests cellular proliferation and extinguishes c-myc expression. EMBO J. 19, 1034–1044 (2000).
Hui, A. B., Lo, K. W., Yin, X. L., Poon, W. S. & Ng, H. K. Detection of multiple gene amplifications in glioblastoma multiforme using array-based comparative genomic hybridization. Lab. Invest. 81, 717–723 (2001).
Ioannidis, P. et al. 8q24 Copy number gains and expression of the c-myc mRNA stabilizing protein CRD-BP in primary breast carcinomas. Int. J. Cancer 104, 54–59 (2003).
Smith, D. R., Myint, T. & Goh, H. S. Over-expression of the c-myc proto-oncogene in colorectal carcinoma. Br. J. Cancer 68, 407–413 (1993).
Eberhart, C. G. et al. Histopathological and molecular prognostic markers in medulloblastoma: c-myc, N-myc, TrkC, and anaplasia. J. Neuropathol. Exp. Neurol. 63, 441–449 (2004).
Chen, C. H., Shen, J., Lee, W. J. & Chow, S. N. Overexpression of cyclin D1 and c-Myc gene products in human primary epithelial ovarian cancer. Int. J. Gynecol. Cancer 15, 878–883 (2005).
Schleger, C., Verbeke, C., Hildenbrand, R., Zentgraf, H. & Bleyl, U. c-MYC activation in primary and metastatic ductal adenocarcinoma of the pancreas: incidence, mechanisms, and clinical significance. Mod. Pathol. 15, 462–469 (2002).
Frost, M. et al. Comparative immunohistochemical analysis of pediatric Burkitt lymphoma and diffuse large B-cell lymphoma. Am. J. Clin. Path. 121, 384–392 (2004).
Davies, A. J. et al. Transformation of follicular lymphoma to diffuse large B-cell lymphoma proceeds by distinct oncogenic mechanisms. Br. J. Haematol. 136, 286–293 (2007).
Nagy, B. et al. Abnormal expression of apoptosis-related genes in haematological malignancies: overexpression of MYC is poor prognostic sign in mantle cell lymphoma. Br. J. Haematol. 120, 434–441 (2003).
Acknowledgements
This research has been supported by grants from the US National Institutes of Health (CA95060, GM085585 and CA122952) and the Leukemia & Lymphoma Society (6225-08). We are grateful to D. Bishop for help with the manuscript.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
Laurence H. Hurley was the scientific founder of Cylene Pharmaceutics, which has the first-in-class G-quadruplex drug Quarfloxin in Phase II clinical trials.
Related links
Glossary
- Negative supercoiling
-
The reduction of twist or unwinding. Because the transcriptional complex associated with RNAP is large, it is unable to rotate around the DNA and instead the DNA rotates to allow forward movement. The DNA twist ahead of the transcriptional complex increases and the twist behind the complex decreases.
- B-DNA
-
DNA with the normal right-handed double helix structure that makes a turn every 3.4 Å.
- Bidirectional transcription
-
Transcription that occurs when there are closely placed divergent promoters in the genome. The dynamic supercoils generated reinforce each other, which increases the likelihood of DNA melting owing to the additive negative supercoiling.
- Melting
-
The process whereby a double helix dissociates into two single strands.
- Stress-induced duplex destabilization
-
The propensity of a duplex site to melt is dependent on the precise sequence and sequence context. Torsional stress induced by negative supercoiling will then open up those sequences that have the lowest energy barrier for melting and this can be calculated using an algorithm.
- Slipped mismatched structures
-
Structures that can occur in regions of DNA with direct repeat symmetry. In the case of the CT element, the repetition of the five repeats of (C/T)C(C/T)TCCCCA allows considerable residual base pairing even when there is an unpaired region. For realignment back to the fully duplex DNA the slipped structure must be melted, which potentially allows the formation of other secondary DNA structures.
- Z-DNA element
-
An element found in DNA molecules with alternating purine and pyrimidine sequences. It is left-handed, with a zigzag pattern of the sugar–phosphate backbone.
- Cruise control
-
The maintenance of an optimum level of promoter firing by using two counter mechanisms to control transcriptional run-off speed.
Rights and permissions
About this article
Cite this article
Brooks, T., Hurley, L. The role of supercoiling in transcriptional control of MYC and its importance in molecular therapeutics. Nat Rev Cancer 9, 849–861 (2009). https://doi.org/10.1038/nrc2733
Published:
Issue Date:
DOI: https://doi.org/10.1038/nrc2733
This article is cited by
-
Downregulation of HINFP induces senescence-associated secretory phenotype to promote metastasis in a non-cell-autonomous manner in bladder cancer
Oncogene (2022)
-
Recent advances in bioprobes and biolabels based on cyanine dyes
Analytical and Bioanalytical Chemistry (2022)
-
Parallel G-quadruplexes recruit the HSV-1 transcription factor ICP4 to promote viral transcription in herpes virus-infected human cells
Communications Biology (2021)
-
Myc-mediated SDHA acetylation triggers epigenetic regulation of gene expression and tumorigenesis
Nature Metabolism (2020)
-
Identification of Hub Genes in Atypical Teratoid/Rhabdoid Tumor by Bioinformatics Analyses
Journal of Molecular Neuroscience (2020)