Key Points
-
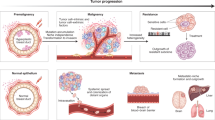
Neoplasms are composed of an ecosystem of evolving clones, competing and cooperating with each other and other cells in their microenvironment, and this has important implications for both neoplastic progression and therapy.
-
Selection at the different levels of genes, cells and organisms might conflict, and have resulted in a legacy of tumour-suppression mechanisms and vulnerability to oncogenesis in our genomes.
-
Most of the dynamics of evolution have not been measured in neoplasms, including mutation rates, fitness effects of mutations, generation times, population structure, the frequency of selective sweeps and the selective effects of our therapies.
-
Many of the genetic and epigenetic alterations observed in neoplasms are evolutionarily neutral.
-
Cancer therapies select for cancer stem cells with resistance mutations, although various evolutionary approaches have been suggested to overcome this problem, including selecting for benign or chemosensitive cells, altering the carrying capacity of the neoplasm and the competitive effects of neoplastic and normal cells on each other.
-
Dispersal theory suggests that high cell mortality and variation of resources and population densities across space might select for metastasis.
-
There is evidence of competition, predation, parasitism and mutualism between co-evolving clones in and around a neoplasm.
-
We will need to interfere with clonal evolution and alter the fitness landscapes of neoplastic cells to prevent or cure cancer. Evolutionary biology should be central to this endeavor.
Abstract
Neoplasms are microcosms of evolution. Within a neoplasm, a mosaic of mutant cells compete for space and resources, evade predation by the immune system and can even cooperate to disperse and colonize new organs. The evolution of neoplastic cells explains both why we get cancer and why it has been so difficult to cure. The tools of evolutionary biology and ecology are providing new insights into neoplastic progression and the clinical control of cancer.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Nowell, P. C. The clonal evolution of tumor cell populations. Science 194, 23–28 (1976). The seminal description of cancer as an evolutionary process. Predicts sequences of clonal expansions, individual variation in response to interventions and therapeutic resistance.
Crespi, B. & Summers, K. Evolutionary Biology of Cancer. Trends Ecol. Evol. 20, 545–552 (2005).
Heppner, G. & Miller, F. The cellular basis of tumor progression. Int. Rev. Cytol. 177, 1–56 (1998).
Cairns, J. Mutation selection and the natural history of cancer. Nature 255, 197–200 (1975). Highlights the interaction between tissue architecture and clonal evolution. Also predicts the retention of a non-recombining, 'immortal' strand of DNA in stem cells.
Tsao, J. L. et al. Genetic reconstruction of individual colorectal tumor histories. Proc. Natl Acad. Sci. USA 97, 1236–1241 (2000). Uses phylogenetic methods to trace the common ancestor of microsatellite-unstable clones in colorectal cancer back to a date before an adenoma was detected.
Tsao, J. L. et al. Colorectal adenoma and cancer divergence. Evidence of multilineage progression. Am. J. Pathol. 154, 1815–1824 (1999).
Michor, F., Iwasa, Y. & Nowak, M. A. Dynamics of cancer progression. Nature Rev. Cancer 4, 197–205 (2004).
Maley, C. C. et al. Genetic clonal diversity predicts progression to esophageal adenocarcinoma. Nature Genet. 38, 468–473 (2006). Shows that genetic diversity measures from ecology and evolution predict progression to malignancy.
Maley, C. C. & Reid, B. J. Natural selection in neoplastic progression of Barrett's esophagus. Semin. Cancer Biol. 15, 474–483 (2005).
Maley, C. C., Reid, B. J. & Forrest, S. Cancer prevention strategies that address the evolutionary dynamics of neoplastic cells: simulating benign cell boosters and selection for chemosensitivity. Cancer Epidemiol. Biomarkers Prev. 13, 1375–1384 (2004). Uses computational models to develop prevention and therapeutic strategies for avoiding the evolution of resistance.
Tomlinson, I. P. M. Game-theory models of interactions between tumour cells. Euro. J. Cancer 33, 1495–1500 (1997).
Gatenby, R. A. & Vincent, T. L. Application of quantitative models from population biology and evolutionary game theory to tumor therapeutic strategies. Mol. Cancer Ther. 2, 919–927 (2003). Describes the use of evolutionary and ecological models to develop new approaches to therapy.
Gonzalez-Garcia, I., Sole, R. V. & Costa, J. Metapopulation dynamics and spatial heterogeneity in cancer. Proc. Natl Acad. Sci. USA 99, 13085–13089 (2002). Shows genetic heterogeneity within a neoplasm, and how clones can be intertwined in complex patterns.
Brash, D. E., Zhang, W., Grossman, D. & Takeuchi, S. Colonization of adjacent stem cell compartments by mutant keratinocytes. Semin. Cancer Biol. 15, 97–102 (2005).
Braakhuis, B. J., Leemans, C. R. & Brakenhoff, R. H. Expanding fields of genetically altered cells in head and neck squamous carcinogenesis. Semin. Cancer Biol. 15, 113–120 (2005).
Maley, C. C. et al. Selectively advantageous mutations and hitchhikers in neoplasms: p16 lesions are selected in Barrett's esophagus. Cancer Res. 64, 3414–3427 (2004).
Franklin, W. A. et al. Widely dispersed p53 mutation in respiratory epithelium. A novel mechanism for field car-cinogenesis. J. Clin. Invest. 100, 2133–2137 (1997). Shows that neoplastic clones can spread over large surface areas.
Castro, M. A., Onsten, T. T., de Almeida, R. M. & Moreira, J. C. Profiling cytogenetic diversity with entropy-based karyotypic analysis. J. Theor. Biol. 234, 487–495 (2005).
Keller, L. K. Levels of Selection in Evolution (Princeton University Press, Princeton, New Jersey, 1999).
Leroi, A. M., Koufopanou, V. & Burt, A. Cancer selection. Nature Rev. Cancer 3, 226–231 (2003).
Weinstein, B. S. & Ciszek, D. The reserve-capacity hypothesis: evolutionary origins and modern implications of the trade-off between tumor-suppression and tissue-repair. Exp. Gerontol. 37, 615–627 (2002).
Frank, S. A. & Nowak, M. A. Problems of somatic mutation and cancer. Bioessays 26, 291–299 (2004).
Campisi, J. Aging, tumor suppression and cancer: high wire-act! Mech. Ageing Dev. 126, 51–58 (2005).
Summers, K., da Silva, J. & Farwell, M. Intragenomic conflict and cancer. Med. Hypotheses 59, 170–179 (2002).
Roth, M. J. et al. Genetic progression and heterogeneity associated with the development of esophageal squamous cell carcinoma. Cancer Res. 61, 4098–4104 (2001).
Hanahan, D. & Weinberg, R. A. The hallmarks of cancer. Cell 100, 57–70 (2000).
Stoler, D. L. et al. The onset and extent of genomic instability in sporadic colorectal tumor progression. Proc. Natl Acad. Sci. USA 96, 15121–15126 (1999). Measures the frequency of genomic alterations in colorectal cancer at approximately 11,000 per clone.
Sole, R. V. & Deisboeck, T. S. An error catastrophe in cancer? J. Theor. Biol. 228, 47–54 (2004).
Breivik, J. The evolutionary origin of genetic instability in cancer development. Semin. Cancer Biol. 15, 51–60 (2005).
Michor, F., Iwasa, Y., Vogelstein, B., Lengauer, C. & Nowak, M. A. Can chromosomal instability initiate tumorigenesis? Semin. Cancer Biol. 15, 43–49 (2005).
Rajagopalan, H., Nowak, M. A., Vogelstein, B. & Lengauer, C. The significance of unstable chromosomes in colorectal cancer. Nature Rev. Cancer 3, 695–701 (2003).
Gray, M. W., Burger, G. & Lang, B. F. Mitochondrial evolution. Science 283, 1476–1481 (1999).
Baylin, S. B. & Herman, J. G. DNA hypermethylation in tumorigenesis- epigenetics joins genetics. Trends Genet. 16, 168–174 (2000).
Feinberg, A. P., Ohlsson, R. & Henikoff, S. The epigenetic progenitor origin of human cancer. Nature Rev. Genet. 7, 21–33 (2006).
Weisenberger, D. J. et al. CpG island methylator phenotype underlies sporadic microsatellite instability and is tightly associated with BRAF mutation in colorectal cancer. Nature Genet. 38, 787–793 (2006).
Horie-Inoue, K. & Inoue, S. Epigenetic and proteolytic inactivation of 14–3-3sigma in breast and prostate cancers. Semin. Cancer Biol. 16, 235–239 (2006).
Albertini, R. J., Nicklas, J. A., O'Neill, J. P. & Robison, S. H. In vivo somatic mutations in humans: measurement and analysis. Annu. Rev. Genet. 24, 305–326 (1990).
Araten, D. J. et al. A quantitative measurement of the human somatic mutation rate. Cancer Res. 65, 8111–8117 (2005).
Sniegowski, P. D., Gerrish, P. J., Johnson, T. & Shaver, A. The evolution of mutation rates: separating causes from consequences. Bioessays 22, 1057–1066 (2000).
Sniegowski, P. D., Gerrish, P. J. & Lenski, R. E. Evolution of high mutation rates in experimental populations of E. coli. Nature 387, 703–705 (1997).
Dyer, M. A. & Bremner, R. The search for the retinoblastoma cell of origin. Nature Rev. Cancer 5, 91–101 (2005).
Renan, M. J. How many mutations are required for tumorigenesis? Implications from human cancer data. Mol. Carcinogenesis 7, 139–146 (1993).
Nunney, L. The population genetics of multistage carcinogenesis. Proc. Biol. Sci. 270, 1183–1191 (2003).
Loeb, L. A. Mutator phenotype may be required for multistage carcinogenesis. Cancer Res. 51, 3075–3079 (1991). Shows that the background mutation rate is not adequate to explain carcinogenesis, and proposes that genetic instability might be necessary for cancer to develop.
Tomlinson, I. & Bodmer, W. Selection, the mutation rate and cancer: ensuring that the tail does not wag the dog. Nature Med. 5, 11–12 (1999).
Moolgavkar, S. H. & Luebeck, E. G. Multistage carcinogenesis and the incidence of human cancer. Genes Chromosomes Cancer 38, 302–306 (2003).
Loeb, K. R. & Loeb, L. A. Significance of multiple muta-tions in cancer. Carcinogenesis 21, 379–385 (2000).
Maley, C. C. et al. The combination of genetic instability and clonal expansion predicts progression to esophageal adenocarcinoma. Cancer Res. 64, 7629–7633 (2004).
Ohta, T. Slightly deleterious mutant substitutions in evolution. Nature 246, 96–98 (1973).
Huntly, B. J. & Gilliland, D. G. Leukaemia stem cells and the evolution of cancer-stem-cell research. Nature Rev. Cancer 5, 311–321 (2005).
Al-Hajj, M., Wicha, M. S., Benito-Hernandez, A., Morrison, S. J. & Clarke, M. F. Prospective identification of tumorigenic breast cancer cells. Proc. Natl Acad. Sci. USA 100, 3983–3988 (2003).
Collins, A. T., Berry, P. A., Hyde, C., Stower, M. J. & Maitland, N. J. Prospective identification of tumorigenic prostate cancer stem cells. Cancer Res. 65, 10946–10951 (2005).
Sell, S. Cellular origin of cancer: dedifferentiation or stem cell maturation arrest? Environ. Health Perspect. 101, 15–26 (1993).
Potten, C. S., Booth, C. & Pritchard, D. M. The intestinal epithelial stem cell: the mucosal governor. Int. J. Exp. Pathol. 78, 219–243 (1997).
Michor, F., Frank, S. A., May, R. M., Iwasa, Y. & Nowak, M. A. Somatic selection for and against cancer. J. Theor. Biol. 225, 377–382 (2003).
Frank, S. A. & Nowak, M. A. Cell biology: developmental predisposition to cancer. Nature 422, 494 (2003).
Meza, R., Luebeck, E. G. & Moolgavkar, S. H. Gestational mutations and carcinogenesis. Math. Biosci. 197, 188–210 (2005).
Maynard Smith, J. & Haigh, J. The hitch-hiking effect of a favorable gene. Genet. Res. 231, 1114–1116 (1974).
Lewontin, R. C. The Genetic Basis of Evolutionary Change (Columbia University Press, New York, 1970).
Braakhuis, B. J., Tabor, M. P., Kummer, J. A., Leemans, C. R. & Brakenhoff, R. H. A genetic explanation of Slaughter's concept of field cancerization: evidence and clinical implications. Cancer Res. 63, 1727–1730 (2003).
Chao, E. C. & Lipkin, S. M. Molecular models for the tissue specificity of DNA mismatch repair-deficient carcinogenesis. Nucleic Acids Res. 34, 840–852 (2006).
Fearon, E. R. & Vogelstein, B. A genetic model for colorectal tumorigenesis. Cell 61, 759–767 (1990).
Desper, R. et al. Inferring tree models for oncogenesis from comparative genome hybridization data. J. Comput. Biol. 37–51 (1999).
Smith, G. et al. Mutations in APC, Kirsten-ras, and p53 — alternative genetic pathways to colorectal cancer. Proc. Natl Acad. Sci. USA 99, 9433–9438 (2002).
Kobayashi, S. et al. EGFR mutation and resistance of non-small-cell lung cancer to gefitinib. N. Engl. J. Med. 352, 786–792 (2005).
Gorre, M. E. et al. Clinical resistance to STI-571 cancer therapy caused by BCR-ABL gene mutation or amplification. Science 293, 876–880 (2001).
Wang, T. L. et al. Digital karyotyping identifies thymidylate synthase amplification as a mechanism of resistance to 5-fluorouracil in metastatic colorectal cancer patients. Proc. Natl Acad. Sci. USA 101, 3089–3094 (2004).
Donnenberg, V. S. & Donnenberg, A. D. Multiple drug resistance in cancer revisited: the cancer stem cell hypothesis. J. Clin. Pharmacol. 45, 872–877 (2005).
Michor, F. et al. Dynamics of chronic myeloid leukaemia. Nature 435, 1267–1270 (2005).
Luria, S. E. & Delbruck, M. Mutations of bacteria from virus sensitivity to virus resistance. Genetics 28, 491–511 (1943).
Komarova, N. Stochastic modeling of drug resistance in cancer. J. Theor. Biol. 239, 351–366 (2006).
Roche-Lestienne, C. & Preudhomme, C. Mutations in the ABL kinase domain pre-exist the onset of imatinib treatment. Semin. Hematol. 40, 80–82 (2003).
Iwasa, Y., Nowak, M. A. & Michor, F. Evolution of resistance during clonal expansion. Genetics 172, 2557–2566 (2006).
Knudson, A. G. Jr. Mutation and cancer: statistical study of retinoblastoma. Proc. Natl Acad. Sci. USA 68, 820–823 (1971). Uses a Poisson model to deduce the requirement of two mutations to generate retinoblastoma.
Etzioni, R. et al. The case for early detection. Nature Rev. Cancer 3, 1–10 (2003).
Chabner, B. A. & Roberts, T. G. Jr. Timeline: chemotherapy and the war on cancer. Nature Rev. Cancer 5, 65–72 (2005).
Komarova, N. L. & Wodarz, D. Evolutionary dynamics of mutator phenotypes in cancer: implications for chemotherapy. Cancer Res. 63, 6635–6642 (2003).
Suiter, A. M., Banziger, O. & Dean, A. M. Fitness consequences of a regulatory polymorphism in a seasonal environment. Proc. Natl Acad. Sci. USA 100, 12782–12786 (2003).
Kim, J. J. & Tannock, I. F. Repopulation of cancer cells during therapy: an important cause of treatment failure. Nature Rev. Cancer 5, 516–525 (2005).
Kern, W. & Estey, E. H. High-dose cytosine arabinoside in the treatment of acute myeloid leukemia: review of three randomized trials. Cancer 107, 116–124 (2006).
Dolken, G. Detection of minimal residual disease. Adv. Cancer Res. 82, 133–185 (2001).
Rubin, C. E. et al. DNA aneuploidy in colonic biopsies predicts future development of dysplasia in ulcerative colitis. Gastroenterology 103, 1611–1620 (1992).
Brentnall, T. A. et al. Mutations in the p53 gene: an early marker of neoplastic progression in ulcerative colitis. Gastroenterology 107, 369–378 (1994).
van Oijen, M. G. & Slootweg, P. J. Oral field cancerization: carcinogen-induced independent events or micrometastatic deposits? Cancer Epidemiol. Biomarkers Prev. 9, 249–256 (2000).
Hunter, K. W. Allelic diversity in the host genetic background may be an important determinant in tumor metastatic dissemination. Cancer Lett. 200, 97–105 (2003).
Bernards, R. & Weinberg, R. A. A progression puzzle. Nature 418, 823 (2002).
Bernards, R. & Weinberg, R. A. Bernards and Weinberg reply. Nature 419, 560 (2002).
Fidler, I. J. & Kripke, M. L. Metastasis results from preexisting variant cells within a malignant tumor. Science 197, 893–895 (1977).
Kirkwood, T. B. Evolution of ageing. Mech. Ageing Dev. 123, 737–745 (2002).
Futuyma, D. J. Evolutionary Biology (Sinauer Associates Inc., Sunderland, Massachusetts, 1998).
Paszek, M. J. et al. Tensional homeostasis and the malignant phenotype. Cancer Cell 8, 241–254 (2005).
Vaupel, P. & Mayer, A. Hypoxia and anemia: effects on tumor biology and treatment resistance. Transfus. Clin. Biol. 12, 5–10 (2005).
Cadet, C., Ferriere, R., Metz, J. A. & van Baalen, M. The evolution of dispersal under demographic stochasticity. Am. Nat. 162, 427–441 (2003).
Fidler, I. J. The pathogenesis of cancer metastasis: the 'seed and soil' hypothesis revisited. Nature Rev. Cancer 3, 453–458 (2003).
Kolar, C. S. & Lodge, D. M. Progress in invasion biology: predicting invaders. Trends Ecol. Evol. 16, 199–204 (2001).
Peterson, A. T. Predicting the geography of species' invasions via ecological niche modeling. Q. Rev. Biol. 78, 419–433 (2003).
Shea, K. & Chesson, P. Community ecology theory as a framework for biological invasions. Trends Ecol. Evol. 17, 170–176 (2002).
Mable, B. K. Breaking down taxonomic barriers in polyploidy research. Trends Plant Sci. 8, 582–590 (2003).
Otto, S. P. & Whitton, J. Polyploid incidence and evolution. Annu. Rev. Genet. 34, 401–437 (2000).
Imhof, M. & Schlotterer, C. E. coli microcosms indicate a tight link between predictability of ecosystem dynamics and diversity. PLoS Genet. 2, e103 (2006).
Heppner, G., Miller, B., Cooper, D. N. & Miller, F. R. in Cell Biology of Breast Cancer (eds McGrath, C. M., Brennan, M. J. & Rich, M. A.) 161–172 (Academic Press, New York, 1980).
Guba, M. et al. A primary tumor promotes dormancy of solitary tumor cells before inhibiting angiogenesis. Cancer Res. 61, 5575–5579 (2001).
Miller, B. E., Miller, F. R., Leith, J. & Heppner, G. H. Growth interaction in vivo between tumor subpopulations derived from a single mouse mammary tumor. Cancer Res. 40, 3977–3981 (1980). Shows that different clones in a neoplasm can affect each other's growth in complex ways.
Caignard, A., Martin, M. S., Michel, M. F. & Martin, F. Interaction between two cellular subpopulations of a rat colonic carcinoma when inoculated to the syngeneic host. Int. J. Cancer 36, 273–279 (1985).
Bach, L. A., Bentzen, S. M., Alsner, J. & Christiansen, F. B. An evolutionary-game model of tumour-cell interactions: possible relevance to gene therapy. Eur. J. Cancer 37, 2116–2120 (2001).
Gatenby, R. A. & Vincent, T. L. An evolutionary model of carcinogenesis. Cancer Res. 63, 6212–6220 (2003).
Nagy, J. D. Competition and natural selection in a mathematical model of cancer. Bull. Math. Biol. 66, 663–687 (2004).
Seliger, B. Strategies of tumor immune evasion. BioDrugs 19, 347–354 (2005).
Uhr, J. W., Scheuermann, R. H., Street, N. E. & Vitetta, E. S. Cancer dormancy: opportunities for new therapeutic approaches. Nature Med. 3, 505–509 (1997).
Mitteldorf, J. Chaotic population dynamics and the evolution of aging. Evol. Ecol. Res. 8, 561–574 (2006).
Rundhaug, J. E. Matrix metalloproteinases and angiogenesis. J. Cell. Mol. Med. 9, 267–285 (2005).
Travisano, M. & Velicer, G. J. Strategies of microbial cheater control. Trends Microbiol. 12, 72–78 (2004).
Turner, P. E. Parasitism between co-infecting bacteriophages. Adv. Ecol. Res. 37, 309–332 (2005).
Velicer, G. J., Kroos, L. & Lenski, R. E. Developmental cheating in the social bacterium Myxococcus xanthus. Nature 404, 598–601 (2000).
Jouanneau, J., Moens, G., Bourgeois, Y., Poupon, M. F. & Thiery, J. P. A minority of carcinoma cells producing acidic fibroblast growth factor induces a community effect for tumor progression. Proc. Natl Acad. Sci. USA 91, 286–290 (1994).
Axelrod, R., Axelrod, D. E. & Pienta, K. J. Evolution of cooperation among tumor cells. Proc. Natl Acad. Sci. USA 103, 13474–13479 (2006).
Mueller, M. M. & Fusenig, N. E. Friends or foes- bipolar effects of the tumour stroma in cancer. Nature Rev. Cancer 4, 839–849 (2004).
Shao, Z. M., Nguyen, M. & Barsky, S. H. Human breast carcinoma desmoplasia is PDGF initiated. Oncogene 19, 4337–4345 (2000).
Tlsty, T. D. Stromal cells can contribute oncogenic signals. Semin. Cancer Biol. 11, 97–104 (2001).
Fukino, K. et al. Combined total genome loss of heterozygosity scan of breast cancer stroma and epithelium reveals multiplicity of stromal targets. Cancer Res. 64, 7231–7236 (2004). Shows that tumour stroma also has genetic lesions, and so must be co-evolving with the epithelium.
Paterson, R. F. et al. Molecular genetic alterations in the laser-capture-microdissected stroma adjacent to bladder carcinoma. Cancer 98, 1830–1836 (2003).
Ishiguro, K., Yoshida, T., Yagishita, H., Numata, Y. & Okayasu, T. Epithelial and stromal genetic instability contributes to genesis of colorectal adenomas. Gut 55, 695–702 (2006).
Kenny, P. A. & Bissell, M. J. Tumor reversion: correction of malignant behavior by microenvironmental cues. Int. J. Cancer 107, 688–695 (2003).
Mintz, B. & Illmensee, K. Normal genetically mosaic mice produced from malignant teratocarcinoma cells. Proc. Natl Acad. Sci. USA 72, 3585–3589 (1975).
Miller, F. R. & Heppner, G. H. Cellular interactions in metastasis. Cancer Metastasis Rev. 9, 21–34 (1990).
Kuperwasser, C. et al. Reconstruction of functionally normal and malignant human breast tissues in mice. Proc. Natl Acad. Sci. USA 101, 4966–4971 (2004).
Bhowmick, N. A. et al. TGF-β signaling in fibroblasts modulates the oncogenic potential of adjacent epithelia. Science 303, 848–851 (2004).
Orimo, A. et al. Stromal fibroblasts present in invasive human breast carcinomas promote tumor growth and angiogenesis through elevated SDF-1/CXCL12 secretion. Cell 121, 335–348 (2005).
Roxburgh, S. H., Shea, K. & Wilson, J. B. The intermediate disturbance hypothesis: patch dynamics and mechanisms of species coexistence. Ecology 85, 359–371 (2004).
Bjerkvig, R., Tysnes, B. B., Aboody, K. S., Najbauer, J. & Terzis, A. J. Opinion: the origin of the cancer stem cell: current controversies and new insights. Nature Rev. Cancer 5, 899–904 (2005).
Boyer, B., Valles, A. M. & Edme, N. Induction and regulation of epithelial-mesenchymal transitions. Biochem. Pharmacol. 60, 1091–1099 (2000).
Elena, S. F. & Lenski, R. E. Evolution experiments with microorganisms: the dynamics and genetic bases of adaptation. Nature Rev. Genet. 4, 457–469 (2003).
Spencer, S. L., Gerety, R. A., Pienta, K. J. & Forrest, S. Modeling somatic evolution in tumorigenesis. PLoS Comput. Biol. 2, e108 (2006).
Drummond, A., Forsberg, R. & Rodrigo, A. G. The inference of stepwise changes in substitution rates using serial sequence samples. Mol. Biol. Evol. 18, 1365–1371 (2001).
Harper, D. M. et al. Sustained efficacy up to 4.5 years of a bivalent L1 virus-like particle vaccine against human papillomavirus types 16 and 18: follow-up from a randomised control trial. Lancet 367, 1247–1255 (2006).
Jakobisiak, M., Lasek, W. & Golab, J. Natural mechanisms protecting against cancer. Immunol. Lett. 90, 103–122 (2003).
Sherr, C. J. Tumor surveillance via the ARF-p53 pathway. Genes Dev. 12, 2984–2991 (1998).
Michor, F., Nowak, M. A., Frank, S. A. & Iwasa, Y. Stochastic elimination of cancer cells. Proc. Biol. Sci. 270, 2017–2024 (2003).
Torres-Montaner, A. & Hughes, D. A hypothetical anti-neoplastic mechanism associated to reserve cells. J. Theor. Biol. 231, 239–248 (2004).
Frank, S. A., Iwasa, Y. & Nowak, M. A. Patterns of cell division and the risk of cancer. Genetics 163, 1527–1532 (2003).
Komarova, N. L. & Cheng, P. Epithelial tissue architecture protects against cancer. Math. Biosci. 200, 90–117 (2006).
Nowak, M. A., Michor, F. & Iwasa, Y. The linear process of somatic evolution. Proc. Natl Acad. Sci. USA 100, 14966–14969 (2003). Uses a mathematical model to analyse the evolutionary dynamics of cells in structured tissues.
Frank, S. A. Genetic predisposition to cancer — insights from population genetics. Nature Rev. Genet. 5, 764–772 (2004).
Fleming, M. A., Potter, J. D., Ramirez, C. J., Ostrander, G. K. & Ostrander, E. A. Understanding missense mutations in the BRCA1 gene: an evolutionary approach. Proc. Natl Acad. Sci. USA 100, 1151–1156 (2003).
Slatkin, M. & Rannala, B. Estimating allele age. Annu. Rev. Genomics Hum. Genet. 1, 225–249 (2000).
Deng, C. X. BRCA1: cell cycle checkpoint, genetic instability, DNA damage response and cancer evolution. Nucleic Acids Res. 34, 1416–1426 (2006).
Jernstrom, H., Johannsson, O., Borg, A., Ivarsson, H. & Olsson, H. BRCA1-positive patients are small for gestational age compared with their unaffected relatives. Eur. J. Cancer 34, 368–371 (1998).
Smith, T. M. et al. Complete genomic sequence and analysis of 117 kb of human DNA containing the gene BRCA1. Genome Res. 6, 1029–1049 (1996).
Pavlicek, A. et al. Evolution of the tumor suppressor BRCA1 locus in primates: implications for cancer pre-disposition. Hum. Mol. Genet. 13, 2737–2751 (2004).
Summers, K. & Crespi, B. Cadherins in maternal–foetal interactions: red queen with a green beard? Proc. Biol. Sci. 272, 643–649 (2005).
Nielsen, R. et al. A scan for positively selected genes in the genomes of humans and chimpanzees. PLoS Biol. 3, e170 (2005).
Crespi, B. J. & Summers, K. Positive selection in the evolution of cancer. Biol. Rev. Camb. Philos. Soc. 81, 407–424 (2006).
Hernandez, L., Kozlov, S., Piras, G. & Stewart, C. L. Paternal and maternal genomes confer opposite effects on proliferation, cell-cycle length, senescence, and tumor formation. Proc. Natl Acad. Sci. USA 100, 13344–13349 (2003).
Moran, P. A. P. Random processes in genetics. Proc. Camb. Phil. Soc. 54, 60–71 (1958).
Ewens, W. J. Mathematical Population Genetics (Springer-Verlag, New York, 2004).
Reid, B. J. et al. Barrett's esophagus: ordering the events that lead to cancer. Euro. J. Cancer Prev. 5 (Suppl. 2), 57–65 (1996). Shows how spatial information can be used to infer dependencies between genetic alterations in carcinogenesis.
Wang, G. Q. et al. Histological precursors of oesophageal squamous cell carcinoma: results from a 13 year prospective follow up study in a high risk population. Gut 54, 187–192 (2005).
Reid, B. J. et al. Predictors of progression in Barrett's esophagus II: baseline 17p (p53) loss of heterozygosity identifies a patient subset at increased risk for neoplastic progression. Am. J. Gastroenterol. 96, 2839–2848 (2001).
Dolan, K., Morris, A. I., Gosney, J. R., Field, J. K. & Sutton, R. Loss of heterozygosity on chromosome 17p predicts neoplastic progression in Barrett's esophagus. J. Gastroenterol. Hepatol. 18, 683–689 (2003).
Teodori, L. et al. DNA/protein flow cytometry as a predictive marker of malignancy in dysplasia-free Barrett's esophagus: thirteen-year follow-up study on a cohort of patients. Cytometry 34, 257–263 (1998).
Lee, J. J. et al. Predicting cancer development in oral leukoplakia: ten years of translational research. Clin. Cancer Res. 6, 1702–1710 (2000).
Rosin, M. P. et al. Use of allelic loss to predict malignant risk for low-grade oral epithelial dysplasia. Clin. Cancer Res. 6, 357–362 (2000).
Befrits, R., Hammarberg, C., Rubio, C., Jaramillo, E. & Tribukait, B. DNA aneuploidy and histologic dysplasia in long-standing ulcerative colitis. A 10-year follow-up study. Dis. Colon. Rectum 37, 313–319; discussion 319–320 (1994).
Lofberg, R., Brostrom, O., Karlen, P., Tribukait, B. & Ost, A. Colonoscopic surveillance in long-standing total ulcerative colitis-a 15-year follow-up study. Gastroenterology 4, 1021–1021 (1990).
Wong, B. C. et al. Helicobacter pylori eradication to prevent gastric cancer in a high-risk region of China: a randomized controlled trial. JAMA 291, 187–194 (2004).
Thompson, I. M. et al. The influence of finasteride on the development of prostate cancer. N. Engl. J. Med. 349, 215–224 (2003).
Abrams, P. A. On classifying interactions between populations. Oecologia 73, 272–281 (1987).
Heppner, G. H., Miller, B. E. & Miller, F. R. Tumor subpopulation interactions in neoplasms. Biochim. Biophys. Acta 695, 215–226 (1983).
Acknowledgements
This work was supported by the US National Institutes of Health, the Commonwealth of Pennsylvania, and the Pew Charitable Trust, and initiated by the Santa Fe Institute. We thank W. Ewens, M. Carroll and J. Radich for helpful comments. We apologize to our peers whose work we were unable to cite owing to space limitations.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Related links
Related links
DATABASES
National Cancer Institute
FURTHER INFORMATION
Glossary
- Clone
-
A set of cells that share a common genotype owing to descent from a common ancestor. In some contexts a clone is more restrictively defined as a set of genetically identical cells.
- Fitness
-
The average contribution of a genotype to future generations. Fitness is generally a function of both survival and reproduction.
- Genetic drift
-
Random changes in allele frequencies over generations. This dynamic of random sampling has a greater effect in smaller populations.
- Neutral mutation
-
A mutation that has no fitness effect (survival or reproductive effect).
- Fixation
-
When an allele (or in this case a clone) reaches 100% frequency in a population.
- Hitchhiker mutation
-
An effectively neutral mutation that expands in a population because it is linked to a selectively advantageous allele. Sometimes called a 'passenger mutation' in cancer biology.
- Molecular clock
-
When mutations occur at a constant rate, the number of mutations that have accumulated between two different lineages is representative of the time since the lineages diverged.
- Selective sweep
-
The process of an adaptive mutation spreading through a population, typically ending in fixation.
- Amensal
-
An interaction between individuals that decreases the fitness of one party but has no effect on the other.
- Lotka–Volterra competition equations
-
The Lotka–Volterra model of competition is based on logistic growth equations of two populations that negatively affect each other's growth.
- Mutualistic
-
An interaction between individuals that increases the fitness of both parties.
- Commensal
-
An interaction between individuals that increases the fitness of one party and has no fitness effect on the other.
- Fitness landscape
-
A multi-dimensional space in which every point represents the genotype or phenotype of a cell and its fitness value. Points are connected if a mutational event can transform one genotype (or phenotype) into the other.
Rights and permissions
About this article
Cite this article
Merlo, L., Pepper, J., Reid, B. et al. Cancer as an evolutionary and ecological process. Nat Rev Cancer 6, 924–935 (2006). https://doi.org/10.1038/nrc2013
Published:
Issue Date:
DOI: https://doi.org/10.1038/nrc2013