Abstract
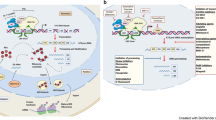
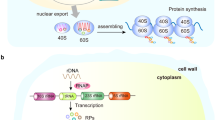
The ribosome is a complex molecular machine composed of numerous distinct proteins and nucleic acids and is responsible for protein synthesis in every living cell. Ribosome biogenesis is one of the most multifaceted and energy- demanding processes in biology, involving a large number of assembly and maturation factors, the functions of which are orchestrated by multiple cellular inputs, including mitogenic signals and nutrient availability. Although causal associations between inherited mutations affecting ribosome biogenesis and elevated cancer risk have been established over the past decade, mechanistic data have emerged suggesting a broader role for dysregulated ribosome biogenesis in the development and progression of most spontaneous cancers. In this Opinion article, we highlight the most recent findings that provide new insights into the molecular basis of ribosome biogenesis in cancer and offer our perspective on how these observations present opportunities for the design of new targeted cancer treatments.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Change history
25 January 2018
In this article a source of grant funding for one of the authors was omitted from the Acknowledgements section. The online version of the article has been corrected to include: “The work of G.T. is also supported by shared funding from the IDIBELL and the Vall d'Hebron Institute of Oncology (VHIO).”
References
Noller, H. F. Evolution of protein synthesis from an RNA world. Cold Spring Harb. Perspect. Biol. 4, a003681 (2012).
Ben-Shem, A. et al. The structure of the eukaryotic ribosome at 3.0 Å resolution. Science 334, 1524–1529 (2011).
Anger, A. M. et al. Structures of the human and Drosophila 80S ribosome. Nature 497, 80–85 (2013).
Wild, T. et al. A protein inventory of human ribosome biogenesis reveals an essential function of exportin 5 in 60S subunit export. PLoS Biol. 8, e1000522 (2010).
Tafforeau, L. et al. The complexity of human ribosome biogenesis revealed by systematic nucleolar screening of pre-rRNA processing factors. Mol. Cell 51, 539–551 (2013).
Badertscher, L. et al. Genome-wide RNAi screening identifies protein modules required for 40S subunit synthesis in human cells. Cell Rep. 13, 2879–2891 (2015).
Orsolic, I. et al. The relationship between the nucleolus and cancer: current evidence and emerging paradigms. Semin. Cancer Biol. 37–38, 36–50 (2016).
Truitt, M. L. & Ruggero, D. New frontiers in translational control of the cancer genome. Nat. Rev. Cancer 16, 288–304 (2016).
Bywater, M. et al. Inhibition of RNA polymerase I as a therapeutic strategy to promote cancer-specific activation of p53. Cancer Cell 22, 51–65 (2012).
Drygin, D. et al. Anticancer activity of CX-3543: a direct inhibitor of rRNA biogenesis. Cancer Res. 69, 7653–7661 (2009).
Bruno, P. M. et al. A subset of platinum-containing chemotherapeutic agents kills cells by inducing ribosome biogenesis stress. Nat. Med. 23, 461–471 (2017).
Ebert, B. L. et al. Identification of RPS14 as a 5q- syndrome gene by RNA interference screen. Nature 451, 335–339 (2008).
Narla, A. & Ebert, B. L. Ribosomopathies: human disorders of ribosome dysfunction. Blood 115, 3196–3205 (2010).
Shen, P. S. et al. Protein synthesis. Rqc2p and 60S ribosomal subunits mediate mRNA-independent elongation of nascent chains. Science 347, 75–78 (2015).
Gentilella, A., Kozma, S. C. & Thomas, G. A liaison between mTOR signaling, ribosome biogenesis and cancer. Biochim. Biophys. Acta 1849, 812–820 (2015).
Schmickel, R. D. Quantitation of human ribosomal DNA: hybridization of human DNA with ribosomal RNA for quantitation and fractionation. Pediatr. Res. 7, 5–12 (1973).
Stults, D. M., Killen, M. W., Pierce, H. H. & Pierce, A. J. Genomic architecture and inheritance of human ribosomal RNA gene clusters. Genome Res. 18, 13–18 (2008).
Gibbons, J. G., Branco, A. T., Godinho, S. A., Yu, S. & Lemos, B. Concerted copy number variation balances ribosomal DNA dosage in human and mouse genomes. Proc. Natl Acad. Sci. USA 112, 2485–2490 (2015).
Caburet, S. et al. Human ribosomal RNA gene arrays display a broad range of palindromic structures. Genome Res. 15, 1079–1085 (2005).
Henderson, A. S., Warburton, D. & Atwood, K. C. Location of ribosomal DNA in the human chromosome complement. Proc. Natl Acad. Sci. USA 69, 3394–3398 (1972).
McStay, B. Nucleolar organizer regions: genomic 'dark matter' requiring illumination. Genes Dev. 30, 1598–1610 (2016).
Goodfellow, S. J. & Zomerdijk, J. C. Basic mechanisms in RNA polymerase I transcription of the ribosomal RNA genes. Subcell. Biochem. 61, 211–236 (2013).
Sorensen, P. D., Lomholt, B., Frederiksen, S. & Tommerup, N. Fine mapping of human 5S rRNA genes to chromosome 1q42.11——q42.13. Cytogenet. Cell Genet. 57, 26–29 (1991).
Uechi, T., Tanaka, T. & Kenmochi, N. A complete map of the human ribosomal protein genes: assignment of 80 genes to the cytogenetic map and implications for human disorders. Genomics 72, 223–230 (2001).
Lafontaine, D. L. Noncoding RNAs in eukaryotic ribosome biogenesis and function. Nat. Struct. Mol. Biol. 22, 11–19 (2015).
Tschochner, H. & Hurt, E. Pre-ribosomes on the road from the nucleolus to the cytoplasm. Trends Cell Biol. 13, 255–263 (2003).
Henras, A. K. et al. The post-transcriptional steps of eukaryotic ribosome biogenesis. Cell. Mol. Life Sci. 65, 2334–2359 (2008).
Woolford, J. L. Jr & Baserga, S. J. Ribosome biogenesis in the yeast Saccharomyces cerevisiae. Genetics 195, 643–681 (2013).
Khatter, H., Myasnikov, A. G., Natchiar, S. K. & Klaholz, B. P. Structure of the human 80S ribosome. Nature 520, 640–645 (2015).
Huh, W. K. et al. Global analysis of protein localization in budding yeast. Nature 425, 686–691 (2003).
Ahmad, Y., Boisvert, F. M., Gregor, P., Cobley, A. & Lamond, A. I. NOPdb: nucleolar proteome database — 2008 update. Nucleic Acids Res. 37, D181–D184 (2009).
Andersen, J. S. et al. Directed proteomic analysis of the human nucleolus. Curr. Biol. 12, 1–11 (2002).
Scherl, A. et al. Functional proteomic analysis of human nucleolus. Mol. Biol. Cell 13, 4100–4109 (2002).
Andersen, J. S. et al. Nucleolar proteome dynamics. Nature 433, 77–83 (2005).
Coute, Y. et al. Deciphering the human nucleolar proteome. Mass Spectrom. Rev. 25, 215–234 (2006).
Shi, Z. et al. Heterogeneous ribosomes preferentially translate dstinct subpools of mRNAs genome-wide. Mol. Cell 67, 71–83.e7 (2017).
Kondrashov, N. et al. Ribosome-mediated specificity in Hox mRNA translation and vertebrate tissue patterning. Cell 145, 383–397 (2011).
Shi, Z. & Barna, M. Translating the genome in time and space: specialized ribosomes, RNA regulons, and RNA-binding proteins. Annu. Rev. Cell Dev. Biol. 31, 31–54 (2015).
Xue, S. et al. RNA regulons in Hox 5′ UTRs confer ribosome specificity to gene regulation. Nature 517, 33–38 (2015).
Komili, S., Farny, N. G., Roth, F. P. & Silver, P. A. Functional specificity among ribosomal proteins regulates gene expression. Cell 131, 557–571 (2007).
Ban, N. et al. A new system for naming ribosomal proteins. Curr. Opin. Struct. Biol. 24, 165–169 (2014).
Gupta, V. & Warner, J. R. Ribosome-omics of the human ribosome. RNA 20, 1004–1013 (2014).
O'Leary, M. N. et al. The ribosomal protein Rpl22 controls ribosome composition by directly repressing expression of its own paralog, Rpl22l1. PLoS Genet. 9, e1003708 (2013).
Slavov, N., Semrau, S., Airoldi, E., Budnik, B. & van Oudenaarden, A. Differential stoichiometry among core ribosomal proteins. Cell Rep. 13, 865–873 (2015).
Simsek, D. et al. The mammalian ribo-interactome reveals ribosome functional diversity and heterogeneity. Cell 169, 1051–1065.e18 (2017).
Pianese, G. Beitrag zur histologie und aetiologie der carcinoma. Histologische und experimentelle untersuchungen [German]. Beitr. Pathol. Anat. Allgem 142, 1–193 (1896).
Serafini, T. et al. Netrin-1 is required for commissural axon guidance in the developing vertebrate nervous system. Cell 87, 1001–1014 (1996).
Delloye-Bourgeois, C. et al. Nucleolar localization of a netrin-1 isoform enhances tumor cell proliferation. Sci. Signal. 5, ra57 (2012).
Justilien, V. et al. Ect2-dependent rRNA synthesis is required for KRAS-TRP53-driven lung adenocarcinoma. Cancer Cell 31, 256–269 (2017).
Albert, B. et al. A molecular titration system coordinates ribosomal protein gene transcription with ribosomal RNA synthesis. Mol. Cell 64, 720–733 (2016).
Barna, M. et al. Suppression of Myc oncogenic activity by ribosomal protein haploinsufficiency. Nature 456, 971–975 (2008).
Mendillo, M. L. et al. HSF1 drives a transcriptional program distinct from heat shock to support highly malignant human cancers. Cell 150, 549–562 (2012).
Santagata, S. et al. Tight coordination of protein translation and HSF1 activation supports the anabolic malignant state. Science 341, 1238303 (2013).
Ellis, S. R. & Gleizes, P. E. Diamond blackfan anemia: ribosomal proteins going rogue. Semin. Hematol. 48, 89–96 (2011).
Ajore, R. et al. Deletion of ribosomal protein genes is a common vulnerability in human cancer, especially in concert with TP53 mutations. EMBO Mol. Med. 9, 498–507 (2017).
Rao, S. et al. Inactivation of ribosomal protein L22 promotes transformation by induction of the stemness factor, Lin28B. Blood 120, 3764–3773 (2012).
Robledo, S. et al. The role of human ribosomal proteins in the maturation of rRNA and ribosome production. RNA 14, 1918–1929 (2008).
O'Donohue, M. F., Choesmel, V., Faubladier, M., Fichant, G. & Gleizes, P. E. Functional dichotomy of ribosomal proteins during the synthesis of mammalian 40S ribosomal subunits. J. Cell Biol. 190, 853–866 (2010).
Karbstein, K. Quality control mechanisms during ribosome maturation. Trends Cell Biol. 23, 242–250 (2013).
Bursac, S., Brdovcak, M. C., Donati, G. & Volarevic, S. Activation of the tumor suppressor p53 upon impairment of ribosome biogenesis. Biochim. Biophys. Acta 1842, 817–830 (2014).
De Keersmaecker, K. et al. Exome sequencing identifies mutation in CNOT3 and ribosomal genes RPL5 and RPL10 in T-cell acute lymphoblastic leukemia. Nat. Genet. 45, 186–190 (2013).
Sulima, S. O. et al. Bypass of the pre-60S ribosomal quality control as a pathway to oncogenesis. Proc. Natl Acad. Sci. USA 111, 5640–5645 (2014).
Belin, S. et al. Dysregulation of ribosome biogenesis and translational capacity is associated with tumor progression of human breast cancer cells. PLoS ONE 4, e7147 (2009).
Marcel, V. et al. p53 acts as a safeguard of translational control by regulating fibrillarin and rRNA methylation in cancer. Cancer Cell 24, 318–330 (2013).
Marcel, V., Catez, F. & Diaz, J. J. p53, a translational regulator: contribution to its tumour-suppressor activity. Oncogene 34, 5513–5523 (2015).
Krogh, N. et al. Profiling of 2′-O-Me in human rRNA reveals a subset of fractionally modified positions and provides evidence for ribosome heterogeneity. Nucleic Acids Res. 44, 7884–7895 (2016).
Stumpf, C. R. & Ruggero, D. The cancerous translation apparatus. Curr. Opin. Genet. Dev. 21, 474–483 (2011).
Yoon, A. et al. Impaired control of IRES-mediated translation in X-linked dyskeratosis congenita. Science 312, 902–906 (2006).
Jack, K. et al. rRNA pseudouridylation defects affect ribosomal ligand binding and translational fidelity from yeast to human cells. Mol. Cell 44, 660–666 (2011).
Bellodi, C. et al. H/ACA small RNA dysfunctions in disease reveal key roles for noncoding RNA modifications in hematopoietic stem cell differentiation. Cell Rep. 3, 1493–1502 (2013).
French, S. L., Osheim, Y. N., Cioci, F., Nomura, M. & Beyer, A. L. In exponentially growing Saccharomyces cerevisiae cells, rRNA synthesis is determined by the summed RNA polymerase I loading rate rather than by the number of active genes. Mol. Cell. Biol. 23, 1558–1568 (2003).
Kobayashi, T. Ribosomal RNA gene repeats, their stability and cellular senescence. Proc. Jpn Acad. Ser. B Phys. Biol. Sci. 90, 119–129 (2014).
Durkin, S. G. & Glover, T. W. Chromosome fragile sites. Annu. Rev. Genet. 41, 169–192 (2007).
Ide, S., Miyazaki, T., Maki, H. & Kobayashi, T. Abundance of ribosomal RNA gene copies maintains genome integrity. Science 327, 693–696 (2010).
Helmrich, A., Ballarino, M. & Tora, L. Collisions between replication and transcription complexes cause common fragile site instability at the longest human genes. Mol. Cell 44, 966–977 (2011).
El Hage, A., French, S. L., Beyer, A. L. & Tollervey, D. Loss of topoisomerase I leads to R-loop-mediated transcriptional blocks during ribosomal RNA synthesis. Genes Dev. 24, 1546–1558 (2010).
Chan, Y. A. et al. Genome-wide profiling of yeast DNA:RNA hybrid prone sites with DRIP-chip. PLoS Genet. 10, e1004288 (2014).
Santos-Pereira, J. M. & Aguilera, A. R loops: new modulators of genome dynamics and function. Nat. Rev. Genet. 16, 583–597 (2015).
Ginno, P. A., Lott, P. L., Christensen, H. C., Korf, I. & Chedin, F. R-Loop formation is a distinctive characteristic of unmethylated human CpG island promoters. Mol. Cell 45, 814–825 (2012).
Aguilera, A. & Garcia-Muse, T. R loops: from transcription byproducts to threats to genome stability. Mol. Cell 46, 115–124 (2012).
Karanam, K., Kafri, R., Loewer, A. & Lahav, G. Quantitative live cell imaging reveals a gradual shift between DNA repair mechanisms and a maximal use of HR in mid S phase. Mol. Cell 47, 320–329 (2012).
Murray, J. M., Stiff, T. & Jeggo, P. A. DNA double-strand break repair within heterochromatic regions. Biochem. Soc. Trans. 40, 173–178 (2012).
Aymard, F. et al. Transcriptionally active chromatin recruits homologous recombination at DNA double-strand breaks. Nat. Struct. Mol. Biol. 21, 366–374 (2014).
Kobayashi, T. Regulation of ribosomal RNA gene copy number and its role in modulating genome integrity and evolutionary adaptability in yeast. Cell. Mol. Life Sci. 68, 1395–1403 (2011).
van Sluis, M. & McStay, B. A localized nucleolar DNA damage response facilitates recruitment of the homology-directed repair machinery independent of cell cycle stage. Genes Dev. 29, 1151–1163 (2015).
Warmerdam, D. O., van den Berg, J. & Medema, R. H. Breaks in the 45S rDNA lead to recombination-mediated loss of repeats. Cell Rep. 14, 2519–2527 (2016).
Lawrence, M. S. et al. Discovery and saturation analysis of cancer genes across 21 tumour types. Nature 505, 495–501 (2014).
Kruhlak, M. et al. The ATM repair pathway inhibits RNA polymerase I transcription in response to chromosome breaks. Nature 447, 730–734 (2007).
Harding, S. M., Boiarsky, J. A. & Greenberg, R. A. ATM dependent silencing links nucleolar chromatin reorganization to DNA damage recognition. Cell Rep. 13, 251–259 (2015).
Larsen, D. H. et al. The NBS1-treacle complex controls ribosomal RNA transcription in response to DNA damage. Nat. Cell Biol. 16, 792–803 (2014).
Ciccia, A. et al. Treacher Collins syndrome TCOF1 protein cooperates with NBS1 in the DNA damage response. Proc. Natl Acad. Sci. USA 111, 18631–18636 (2014).
Stults, D. M. et al. Human rRNA gene clusters are recombinational hotspots in cancer. Cancer Res. 69, 9096–9104 (2009).
Grummt, I. & Langst, G. Epigenetic control of RNA polymerase I transcription in mammalian cells. Biochim. Biophys. Acta 1829, 393–404 (2013).
Dobbelstein, M. & Sorensen, C. S. Exploiting replicative stress to treat cancer. Nat. Rev. Drug Discov. 14, 405–423 (2015).
Gaillard, H., Garcia-Muse, T. & Aguilera, A. Replication stress and cancer. Nat. Rev. Cancer 15, 276–289 (2015).
Flach, J. et al. Replication stress is a potent driver of functional decline in ageing haematopoietic stem cells. Nature 512, 198–202 (2014).
Guetg, C. et al. The NoRC complex mediates the heterochromatin formation and stability of silent rRNA genes and centromeric repeats. EMBO J. 29, 2135–2146 (2010).
Killen, M. W., Stults, D. M., Adachi, N., Hanakahi, L. & Pierce, A. J. Loss of Bloom syndrome protein destabilizes human gene cluster architecture. Hum. Mol. Genet. 18, 3417–3428 (2009).
Grummt, I. Life on a planet of its own: regulation of RNA polymerase I transcription in the nucleolus. Genes Dev. 17, 1691–1702 (2003).
Murayama, A. et al. Epigenetic control of rDNA loci in response to intracellular energy status. Cell 133, 627–639 (2008).
Yang, L. et al. Regulation of SirT1-nucleomethylin binding by rRNA coordinates ribosome biogenesis with nutrient availability. Mol. Cell. Biol. 33, 3835–3848 (2013).
Song, T. et al. The NAD+ synthesis enzyme nicotinamide mononucleotide adenylyltransferase (NMNAT1) regulates ribosomal RNA transcription. J. Biol. Chem. 288, 20908–20917 (2013).
Yang, L., Song, T., Chen, L., Soliman, H. & Chen, J. Nucleolar repression facilitates initiation and maintenance of senescence. Cell Cycle 14, 3613–3623 (2015).
Saxton, R. A. & Sabatini, D. M. mTOR signaling in growth, metabolism, and disease. Cell 169, 361–371 (2017).
Iadevaia, V., Liu, R. & Proud, C. G. mTORC1 signaling controls multiple steps in ribosome biogenesis. Semin. Cell Dev. Biol. 36, 113–120 (2014).
Ben-Sahra, I., Hoxhaj, G., Ricoult, S. J., Asara, J. M. & Manning, B. D. mTORC1 induces purine synthesis through control of the mitochondrial tetrahydrofolate cycle. Science 351, 728–733 (2016).
Ben-Sahra, I., Howell, J. J., Asara, J. M. & Manning, B. D. Stimulation of de novo pyrimidine synthesis by growth signaling through mTOR and S6K1. Science 339, 1323–1328 (2013).
Robitaille, A. M. et al. Quantitative phosphoproteomics reveal mTORC1 activates de novo pyrimidine synthesis. Science 339, 1320–1323 (2013).
Hannan, K. et al. mTOR-dependent regulation of ribosomal gene transcription requires S6K1 and is mediated by phosphorylation of the carboxy-terminal activation domain of the nucleolar transcription factor UBF. Mol. Cell. Biol. 23, 8862–8877 (2003).
Mayer, C., Zhao, J., Yuan, X. & Grummt, I. mTOR-dependent activation of the transcription factor TIF-IA links rRNA synthesis to nutrient availability. Genes Dev. 18, 423–434 (2004).
Kantidakis, T., Ramsbottom, B. A., Birch, J. L., Dowding, S. N. & White, R. J. mTOR associates with TFIIIC, is found at tRNA and 5S rRNA genes, and targets their repressor Maf1. Proc. Natl Acad. Sci. USA 107, 11823–11828 (2010).
Shor, B. et al. Requirement of the mTOR kinase for the regulation of Maf1 phosphorylation and control of RNA polymerase III-dependent transcription in cancer cells. J. Biol. Chem. 285, 15380–15392 (2010).
Gentilella, A. & Thomas, G. Cancer biology: the director's cut. Nature 485, 50–51 (2012).
Hsieh, A. C. et al. The translational landscape of mTOR signalling steers cancer initiation and metastasis. Nature 485, 55–61 (2012).
Thoreen, C. C. et al. A unifying model for mTORC1-mediated regulation of mRNA translation. Nature 485, 109–113 (2012).
Gentilella, A. et al. Autogenous control of 5′TOP mRNA stability by 40S ribosomes. Mol. Cell 67, 55–70.e4 (2017).
Hsu, P. P. et al. The mTOR-regulated phosphoproteome reveals a mechanism of mTORC1-mediated inhibition of growth factor signaling. Science 332, 1317–1322 (2011).
Yu, Y. et al. Phosphoproteomic analysis identifies Grb10 as an mTORC1 substrate that negatively regulates insulin signaling. Science 332, 1322–1326 (2011).
Chauvin, C. et al. Ribosomal protein S6 kinase activity controls the ribosome biogenesis transcriptional program. Oncogene 33, 474–483 (2013).
Pourdehnad, M. et al. Myc and mTOR converge on a common node in protein synthesis control that confers synthetic lethality in Myc-driven cancers. Proc. Natl Acad. Sci. USA 110, 11988–11993 (2013).
Chan, J. et al. AKT promotes rRNA synthesis and cooperates with c-MYC to stimulate ribosome biogenesis in cancer. Sci. Signal. 4, ra56 (2011).
Kim, L. C., Cook, R. S. & Chen, J. mTORC1 and mTORC2 in cancer and the tumor microenvironment. Oncogene 36, 2191–2201 (2017).
Deshaies, R. J. Proteotoxic crisis, the ubiquitin-proteasome system, and cancer therapy. BMC Biol. 12, 94 (2014).
Bhat, M. et al. Targeting the translation machinery in cancer. Nat. Rev. Drug Discov. 14, 261–278 (2015).
Harper, J. W. & Bennett, E. J. Proteome complexity and the forces that drive proteome imbalance. Nature 537, 328–338 (2016).
Brandman, O. & Hegde, R. S. Ribosome-associated protein quality control. Nat. Struct. Mol. Biol. 23, 7–15 (2016).
Conn, C. S. & Qian, S. B. Nutrient signaling in protein homeostasis: an increase in quantity at the expense of quality. Sci. Signal. 6, ra24 (2013).
Koplin, A. et al. A dual function for chaperones SSB-RAC and the NAC nascent polypeptide-associated complex on ribosomes. J. Cell Biol. 189, 57–68 (2010).
Bublik, D. R. et al. Regulatory module involving FGF13, miR-504, and p53 regulates ribosomal biogenesis and supports cancer cell survival. Proc. Natl Acad. Sci. USA 114, E496–E505 (2016).
Zhang, Y. et al. Coordinated regulation of protein synthesis and degradation by mTORC1. Nature 513, 440–443 (2014).
Topalian, S. L., Taube, J. M., Anders, R. A. & Pardoll, D. M. Mechanism-driven biomarkers to guide immune checkpoint blockade in cancer therapy. Nat. Rev. Cancer 16, 275–287 (2016).
Volarevic, S. et al. Proliferation, but not growth, blocked by conditional deletion of 40S ribosomal protein S6. Science 288, 2045–2047 (2000).
Pestov, D. G., Strezoska, Z. & Lau, L. F. Evidence of p53-dependent cross-talk between ribosome biogenesis and the cell cycle: effects of nucleolar protein Bop1 on G1/S transition. Mol. Cell. Biol. 21, 4246–4255 (2001).
Lohrum, M., Ludwig, R., Kubbutat, M., Hanlon, M. & Vousden, K. Regulation of HDM2 activity by the ribosomal protein L11. Cancer Cell 3, 577–587 (2003).
Zhang, Y. et al. Ribosomal protein L11 negatively regulates oncoprotein MDM2 and mediates a p53-dependent ribosomal-stress checkpoint pathway. Mol. Cell. Biol. 23, 8902–8912 (2003).
Bhat, K., Itahana, K., Jin, A. & Zhang, Y. Essential role of ribosomal protein L11 in mediating growth inhibition-induced p53 activation. EMBO J. 23, 2402–2412 (2004).
Dai, M.-S. & Lu, H. Inhibition of MDM2-mediated p53 ubiquitination and degradation by ribosomal protein L5. J. Biol. Chem. 279, 44475–44482 (2004).
Donati, G., Peddigari, S., Mercer, C. A. & Thomas, G. 5S ribosomal RNA is an essential component of a nascent ribosomal precursor complex that regulates the Hdm2-p53 checkpoint. Cell Rep. 4, 87–98 (2013).
Bursac, S. et al. Mutual protection of ribosomal proteins L5 and L11 from degradation is essential for p53 activation upon ribosomal biogenesis stress. Proc. Natl Acad. Sci. USA 109, 20467–20472 (2012).
Boisvert, F. M., van Koningsbruggen, S., Navascues, J. & Lamond, A. I. The multifunctional nucleolus. Nat. Rev. Mol. Cell Biol. 8, 574–585 (2007).
Bode, A. M. & Dong, Z. Post-translational modification of p53 in tumorigenesis. Nat. Rev. Cancer 4, 793–805 (2004).
Zhang, Y. & Lu, H. Signaling to p53: ribosomal proteins find their way. Cancer Cell 16, 369–377 (2009).
Fumagalli, S., Ivanenkov, V. V., Teng, T. & Thomas, G. Suprainduction of p53 by disruption of 40S and 60S ribosome biogenesis leads to the activation of a novel G2/M checkpoint. Genes Dev. 26, 1028–1040 (2012).
Horn, H. F. & Vousden, K. H. Cooperation between the ribosomal proteins L5 and L11 in the p53 pathway. Oncogene 27, 5774–5784 (2008).
Sloan, K. E., Bohnsack, M. T. & Watkins, N. J. The 5S RNP couples p53 homeostasis to ribosome biogenesis and nucleolar stress. Cell Rep. 5, 237–247 (2013).
Zheng, J. et al. Structure of human MDM2 complexed with RPL11 reveals the molecular basis of p53 activation. Genes Dev. 29, 1524–1534 (2015).
Rubbi, C. P. & Milner, J. Disruption of the nucleolus mediates stabilization of p53 in response to DNA damage and other stresses. EMBO J. 22, 6068–6077 (2003).
Fumagalli, S. et al. Absence of nucleolar disruption after impairment of 40S ribosome biogenesis reveals an rpL11-translation-dependent mechanism of p53 induction. Nat. Cell Biol. 11, 501–508 (2009).
Nicolas, E. et al. Involvement of human ribosomal proteins in nucleolar structure and p53-dependent nucleolar stress. Nature Commun. 7, 11390 (2016).
Zemp, I. & Kutay, U. Nuclear export and cytoplasmic maturation of ribosomal subunits. FEBS Lett. 581, 2783–2793 (2007).
Morgado-Palacin, L. et al. Partial loss of Rpl11 in adult mice recapitulates diamond-blackfan anemia and promotes lymphomagenesis. Cell Rep. 13, 712–722 (2015).
Macias, E. et al. An ARF-independent c-MYC-activated tumor suppression pathway mediated by ribosomal protein-Mdm2 Interaction. Cancer Cell 18, 231–243 (2010).
Nie, Z. et al. c-Myc is a universal amplifier of expressed genes in lymphocytes and embryonic stem cells. Cell 151, 68–79 (2012).
Lin, C. Y. et al. Transcriptional amplification in tumor cells with elevated c-Myc. Cell 151, 56–67 (2012).
van Riggelen, J., Yetil, A. & Felsher, D. MYC as a regulator of ribosome biogenesis and protein synthesis. Nat. Rev. Cancer 10, 301–309 (2010).
Lowe, S. W., Cepero, E. & Evan, G. Intrinsic tumour suppression. Nature 432, 307–315 (2004).
Dai, M.-S., Arnold, H., Sun, X.-X., Sears, R. & Lu, H. Inhibition of c-Myc activity by ribosomal protein L11. EMBO J. 26, 3332–3345 (2007).
Dai, M.-S., Sun, X.-X. & Lu, H. Ribosomal protein L11 associates with c-Myc at 5 S rRNA and tRNA genes and regulates their expression. J. Biol. Chem. 285, 12587–12594 (2010).
Challagundla, K. et al. Ribosomal protein L11 recruits miR-24/miRISC to repress c-Myc expression in response to ribosomal stress. Mol. Cell. Biol. 31, 4007–4021 (2011).
Liao, J. M., Zhou, X., Gatignol, A. & Lu, H. Ribosomal proteins L5 and L11 co-operatively inactivate c-Myc via RNA-induced silencing complex. Oncogene 33, 4916–4923 (2013).
Perez-Mancera, P. A., Young, A. R. & Narita, M. Inside and out: the activities of senescence in cancer. Nat. Rev. Cancer 14, 547–558 (2014).
Nishimura, K. et al. Perturbation of ribosome biogenesis drives cells into senescence through 5S RNP-mediated p53 activation. Cell Rep. 10, 1310–1323 (2015).
Halazonetis, T. D., Gorgoulis, V. G. & Bartek, J. An oncogene-induced DNA damage model for cancer development. Science 319, 1352–1355 (2008).
Fancello, L., Kampen, K. R. & Hofman, I. J., Verbeeck, J. & De Keersmaecker, K. The ribosomal protein gene RPL5 is a haploinsufficient tumor suppressor in multiple cancer types. Oncotarget 8, 14462–14478 (2017).
Woods, S. J., Hannan, K. M., Pearson, R. B. & Hannan, R. D. The nucleolus as a fundamental regulator of the p53 response and a new target for cancer therapy. Biochim. Biophys. Acta 1849, 821–829 (2015).
Golomb, L. Volarevic, S. & Oren, M. p53 and ribosome biogenesis stress: the essentials. FEBS Lett. 588, 2571–2579 (2014).
Peltonen, K. et al. A targeting modality for destruction of RNA polymerase I that possesses anticancer activity. Cancer Cell 25, 77–90 (2014).
Perry, R. P. & Kelley, D. E. Inhibition of RNA synthesis by actinomycin D: characteristic dose-response of different RNA species. J. Cell. Physiol. 76, 127–139 (1970).
Morgado-Palacin, L. et al. Non-genotoxic activation of p53 through the RPL11-dependent ribosomal stress pathway. Carcinogenesis 35, 2822–2830 (2014).
Yang, D. & Okamoto, K. Structural insights into G-quadruplexes: towards new anticancer drugs. Future Med. Chem. 2, 619–646 (2010).
Drygin, D. et al. Targeting RNA polymerase I with an oral small molecule CX-5461 inhibits ribosomal RNA synthesis and solid tumor growth. Cancer Res. 71, 1418–1430 (2011).
US National Library of Medicine. ClinicalTrials.gov https://www.clinicaltrials.gov/ct2/show/NCT00955786?term=CX-3543&rank=1 (2009).
Australian New Zealand Clinical Trials Registry. Trial review. Australian New Zealand Clinical Trials Registry https://www.anzctr.org.au/Trial/Registration/TrialReview.aspx?id=364713 (2013)
Harrison, S. J. et al. A phase 1, open-label, dose escalation, safety, PK and PD study of a first in class Pol1 inhibitor (CX-5461) in patients with advanced hematologic malignancies (HM). J. Clin. Oncol. 33, e22212 (2015).
US Natonal Library of Medicine. ClinicalTrials.gov https://www.clinicaltrials.gov/ct2/show/NCT00780663?term=NCT00780663&rank=1 (2010).
Xu, H. et al. CX-5461 is a DNA G-quadruplex stabilizer with selective lethality in BRCA1/2 deficient tumours. Nat. Commun. 8, 14432 (2017).
US National Library of Medicine. ClinicalTrials.gov https://www.clinicaltrials.gov/ct2/show/NCT02719977?term=NCT02719977&rank=1 (2017).
Raymond, E., Faivre, S., Chaney, S., Woynarowski, J. & Cvitkovic, E. Cellular and molecular pharmacology of oxaliplatin. Mol. Cancer Ther. 1, 227–235 (2002).
Machover, D. et al. Two consecutive phase II studies of oxaliplatin (L-OHP) for treatment of patients with advanced colorectal carcinoma who were resistant to previous treatment with fluoropyrimidines. Ann. Oncol. 7, 95–98 (1996).
Andre, T. et al. Adjuvant fluorouracil, leucovorin, and oxaliplatin in stage II to III colon cancer: updated 10-year survival and outcomes according to BRAF mutation and mismatch repair status of the MOSAIC study. J. Clin. Oncol. 33, 4176–4187 (2015).
Burger, K. et al. Chemotherapeutic drugs inhibit ribosome biogenesis at various levels. J. Biol. Chem. 285, 12416–12425 (2010).
Cancer Genome Atlas Network. Comprehensive molecular characterization of human colon and rectal cancer. Nature 487, 330–337 (2012).
Zhang, Q. et al. The role of IMP dehydrogenase 2 in Inauhzin-induced ribosomal stress. eLife 3, e03077 (2014).
Kawashima, S. A. et al. Potent, reversible, and specific chemical inhibitors of eukaryotic ribosome biogenesis. Cell 167, 512–524.e14 (2016).
Guinney, J. et al. The consensus molecular subtypes of colorectal cancer. Nat. Med. 21, 1350–1356 (2015).
Loke, P. & Allison, J. P. PD-L1 and PD-L2 are differentially regulated by Th1 and Th2 cells. Proc. Natl Acad. Sci. USA 100, 5336–5341 (2003).
Lastwika, K. J. et al. Control of PD-L1 expression by oncogenic activation of the AKT-mTOR pathway in non-small cell lung cancer. Cancer Res. 76, 227–238 (2016).
Devlin, J. R. et al. Combination therapy targeting ribosome biogenesis and mRNA translation synergistically extends survival in MYC-driven lymphoma. Cancer Discov. 6, 59–70 (2015).
Crawford, L. J. & Irvine, A. E. Targeting the ubiquitin proteasome system in haematological malignancies. Blood Rev. 27, 297–304 (2013).
Anderson, D. J. et al. Targeting the AAA ATPase p97 as an approach to treat cancer through disruption of protein homeostasis. Cancer Cell 28, 653–665 (2015).
Ingolia, N. T., Brar, G. A., Rouskin, S., McGeachy, A. M. & Weissman, J. S. The ribosome profiling strategy for monitoring translation in vivo by deep sequencing of ribosome-protected mRNA fragments. Nat. Protoc. 7, 1534–1550 (2012).
Gorski, S. A., Snyder, S. K., John, S., Grummt, I. & Misteli, T. Modulation of RNA polymerase assembly dynamics in transcriptional regulation. Mol. Cell 30, 486–497 (2008).
Klein, J. & Grummt, I. Cell cycle-dependent regulation of RNA polymerase I transcription: the nucleolar transcription factor UBF is inactive in mitosis and early G1. Proc. Natl Acad. Sci. USA 96, 6096–6101 (1999).
Berger, C., Horlebein, A., Gogel, E. & Grummt, F. Temporal order of replication of mouse ribosomal RNA genes during the cell cycle. Chromosoma 106, 479–484 (1997).
Li, J., Santoro, R., Koberna, K. & Grummt, I. The chromatin remodeling complex NoRC controls replication timing of rRNA genes. EMBO J. 24, 120–127 (2005).
Smirnov, E. et al. Separation of replication and transcription domains in nucleoli. J. Struct. Biol. 188, 259–266 (2014).
Dalgaard, J. Z., Godfrey, E. L. & MacFarlane, R. J. in DNA Replication-Current Advances Ch. 13 (ed. Seligmann, H.) (InTech, 2011).
Akamatsu, Y. & Kobayashi, T. The human RNA polymerase I transcription terminator complex acts as a replication fork barrier that coordinates the progress of replication with rRNA transcription activity. Mol. Cell. Biol. 35, 1871–1881 (2015).
McStay, B. & Grummt, I. The epigenetics of rRNA genes: from molecular to chromosome biology. Annu. Rev. Cell Dev. Biol. 24, 131–157 (2008).
Jossen, R. & Bermejo, R. The DNA damage checkpoint response to replication stress: a game of forks. Front. Genet. 4, 26 (2013).
Klusmann, I. et al. p53 activity results in dna replication fork processivity. Cell Rep. 17, 1845–1857 (2016).
Wang, M., Herrmann, C. J., Simonovic, M., Szklarczyk, D. & von Mering, C. Version 4.0 of PaxDb: protein abundance data, integrated across model organisms, tissues, and cell-lines. Proteomics 15, 3163–3168 (2015).
Turowski, T. W. & Tollervey, D. Cotranscriptional events in eukaryotic ribosome synthesis. Wiley Interdiscip. Rev. RNA 6, 129–139 (2015).
Wilson, D. N. & Doudna Cate, J. H. The structure and function of the eukaryotic ribosome. Cold Spring Harb. Perspect. Biol. 4, a011536 (2012).
Lane, A. N. & Fan, T. W. Regulation of mammalian nucleotide metabolism and biosynthesis. Nucleic Acids Res. 43, 2466–2485 (2015).
Henras, A. K., Plisson-Chastang, C., O'Donohue, M. F., Chakraborty, A. & Gleizes, P. E. An overview of pre-ribosomal RNA processing in eukaryotes. Wiley Interdiscip. Rev. RNA 6, 225–242 (2015).
Strunk, B. S. & Karbstein, K. Powering through ribosome assembly. RNA 15, 2083–2104 (2009).
Granneman, S. & Tollervey, D. Building ribosomes: even more expensive than expected? Curr. Biol. 17, R415–R417 (2007).
Nagaraj, N. et al. Deep proteome and transcriptome mapping of a human cancer cell line. Mol. Syst. Biol. 7, 548 (2011).
Beck, M. et al. The quantitative proteome of a human cell line. Mol. Syst. Biol. 7, 549 (2011).
Geiger, T., Wehner, A., Schaab, C., Cox, J. & Mann, M. Comparative proteomic analysis of eleven common cell lines reveals ubiquitous but varying expression of most proteins. Mol. Cell. Proteomics 11, M111.014050 (2012).
Warner, J. R. In the absence of ribosomal RNA synthesis, the ribosomal proteins of HeLa cells are synthesized normally and degraded rapidly. J. Mol. Biol. 115, 315–333 (1977).
Lam, Y. W., Lamond, A. I., Mann, M. & Andersen, J. S. Analysis of nucleolar protein dynamics reveals the nuclear degradation of ribosomal proteins. Curr. Biol. 17, 749–760 (2007).
Boisvert, F. M. et al. A quantitative spatial proteomics analysis of proteome turnover in human cells. Mol. Cell. Proteomics 11, M111.011429 (2012).
Acknowledgements
The authors thank A. Gentilella, R. Loewith, C. Munoz-Pinedo, D. Ruggero, J. Warner and R. White for providing insights and comments during the preparation of this manuscript. G.T.'s laboratory is funded by grants from the Asociación Española Contra el Cáncer (AECC) (grant GCB14-2035), Instituto de Salud Carlos III (ISCIII) (grants IIS10/00015), the Spanish Ministry of Science and Innovation (grant SAF2011-24967), the Career Integration Grants (CIG) European Commission (grant PCIG10-GA-2011-304160), the US National Institutes of Health (NIH)/National Cancer Institute (NCI) (grant R01-CA158768), ISCIII-Red Temática de Investigación Cooperativa en Cáncer (RTICC) (grants RD12/0036/0049), Agència de Gestió d'Ajuts Universitaris i de Recerca (AGAUR) (grant SGR 870) and Ministerio de Economía y Competitividad, ISCIII (grant PIE13/00022). The work of G.T. is also supported by shared funding from the IDIBELL and the Vall d'Hebron Institute of Oncology (VHIO). S.V.'s laboratory is funded by the Croatian Science Foundation (CSF) (grant 2079), and J.P. is supported by a Juan de la Cierva Fellowship (FJCI-2014-20422). The authors apologize to those whose work could not be cited because of space limitations.
Author information
Authors and Affiliations
Contributions
The authors contributed equally to this manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Glossary
- 5′-Terminal oligopyrimidine tract
-
(5′TOP). A specific mRNA sequence that contains a cytosine at the transcriptional start site followed by a stretch of 4 to 13 pyrimidine nucleotides.
- 7-Methylguanylate-cap-dependent mRNA translation
-
The prevalent mode of translation initiation in eukaryotic cells, mediated by binding of the eukaryotic translation initiation factor 4E (eIF4E) to the 5′ end of an mRNA via the 7-methylguanylate-cap structure (m7GpppN, where N is any nucleotide), which recruits the 40S ribosome to mRNAs.
- Common fragile sites
-
(CFS). Unstable chromosomal regions that preferentially break or form gaps during metaphase.
- G-quadruplex DNA
-
(G4 DNA). A stable secondary helical structure formed in G-rich nucleic acid sequences and consisting of one or more square planar structures (called tetrads) formed by guanine hydrogen bonds. The G4 structures can form intramolecularly or intermolecularly from DNA strands in parallel or in antiparallel positions.
- Homologous recombination
-
A DNA double-strand break repair process that uses a homologous DNA sequence as a template for repair.
- Internal ribosome entry site
-
(IRES). An RNA structural element that facilitates translation initiation by recruiting 40S ribosomes to an internal position in eukaryotic mRNAs in a cap-independent manner.
- I-PpoI
-
A homing endonuclease from Physarum polycephalum that recognizes and cleaves a 15-base pair sequence present in 47S and 5S ribosomal DNA genes as well as 12 other sites in the human genome.
- Non-homologous end joining
-
(NHEJ). A DNA double-strand break repair process that modifies broken ends and ligates them without the participation of a homologous DNA sequence.
- Polysomes
-
Complexes of an mRNA with two or more ribosomes.
- Ribosomal DNA
-
(rDNA). A region in the nucleolus containing genes whose transcription by RNA polymerase I gives rise to the precursor 47S ribosomal RNA.
- Ribosome profiling
-
Also known as ribosome footprinting or Ribo-Seq. A genome-wide, quantitative technique that uses deep sequencing of mRNA fragments protected by ribosomes from nuclease digestion.
- Ribosomopathies
-
Inherited human disorders caused by mutations in genes encoding ribosome components, or ribosome biogenesis factors, leading to impaired ribosome biogenesis or function.
- Unequal sister chromatid exchange
-
The non-reciprocal exchange of a nucleotide sequence that occurs between repeated sequences present at non-equivalent locations on sister chromatids.
Rights and permissions
About this article
Cite this article
Pelletier, J., Thomas, G. & Volarević, S. Ribosome biogenesis in cancer: new players and therapeutic avenues. Nat Rev Cancer 18, 51–63 (2018). https://doi.org/10.1038/nrc.2017.104
Published:
Issue Date:
DOI: https://doi.org/10.1038/nrc.2017.104
This article is cited by
-
Quantified pathway mutations associate epithelial-mesenchymal transition and immune escape with poor prognosis and immunotherapy resistance of head and neck squamous cell carcinoma
BMC Medical Genomics (2024)
-
Refining mutanome-based individualised immunotherapy of melanoma using artificial intelligence
European Journal of Medical Research (2024)
-
METTL3 promotes osteoblast ribosome biogenesis and alleviates periodontitis
Clinical Epigenetics (2024)
-
CTPS1 is a novel therapeutic target in multiple myeloma which synergizes with inhibition of CHEK1, ATR or WEE1
Leukemia (2024)
-
The heterogeneity of signaling pathways and drug responses in intrahepatic cholangiocarcinoma with distinct genetic mutations
Cell Death & Disease (2024)