Key Points
-
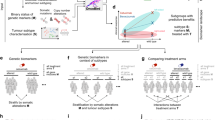
Molecular alterations fostering the progression of colorectal cancers are acquired early in the carcinogenesis process, and there is inter-connectivity among genomic drivers (gene mutations and chromosomal instability), transcriptomic subtypes (microsatellite instability immune, canonical, metabolic or mesenchymal) and immune signatures (highly immunogenic, poorly immunogenic or inflamed and immune tolerant).
-
Primary and metastatic samples display major similarities at the genomic level: novel gene alterations are usually related to chemotherapy or targeted therapy pressure. More studies on inter-metastatic spatial heterogeneity, molecular shifts at the transcriptomic level and changes in microenvironment markers are needed.
-
Until recently, the evolution of biomarkers for targeted therapies in colorectal cancer has been restrictive, with the identification of multiple negative predictive factors determining the response to epidermal growth factor receptor (EGFR) monoclonal antibodies. At progression to these agents, there is convergent reactivation of MAPK pathway
-
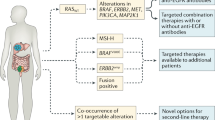
Emerging positive predictive markers for targeted therapies include infrequent genomic events, such as BRAFV600E mutations, ERBB2 amplifications, anaplastic lymphoma kinase (ALK) and neurotrophic receptor tyrosine kinase (NTRK) fusions and alterations in upstream nodes of the WNT pathway, such as ring finger protein 43 (RNF43), zinc and ring finger 3 (ZNRF3) and R-spondin (RSPO) genes. For immune checkpoint inhibitors, promising biomarkers include microsatellite instability and DNA polymerase-ε (POLE) mutations
-
Biomarker–drug co-development has evolved to accommodate a 'multi-molecular, multi-drug' perspective of precision medicine. Novel contexts of vulnerability are likely to be identified, leading to drug-repurposing strategies and combination therapies to halt tumour evolution and tackle minimal residual disease.
Abstract
Critical driver genomic events in colorectal cancer have been shown to affect the response to targeted agents that were initially developed under the 'one gene, one drug' paradigm of precision medicine. Our current knowledge of the complexity of the cancer genome, clonal evolution patterns under treatment pressure and pharmacodynamic effects of target inhibition support the transition from a one gene, one drug approach to a 'multi-gene, multi-drug' model when making therapeutic decisions. Better characterization of the transcriptomic subtypes of colorectal cancer, encompassing tumour, stromal and immune components, has revealed convergent pathway dependencies that mandate a 'multi-molecular' perspective for the development of therapies to treat this disease.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Change history
06 March 2017
In this article a source of grant funding for one of the authors was omitted from the Acknowledgements section. The online version of the article has been corrected to include: "The work of R.D. was supported by the Grant for Oncology Innovation under the project 'Next generation of clinical trials with matched targeted therapies in colorectal cancer'".
References
Ferlay, J. et al. Cancer incidence and mortality patterns in Europe: estimates for 40 countries in 2012. Eur. J. Cancer 49, 1374–1403 (2013).
Siegel, R., Desantis, C. & Jemal, A. Colorectal cancer statistics, 2014. CA Cancer J. Clin. 64, 104–117 (2014).
Welch, H. G. & Robertson, D. J. Colorectal cancer on the decline — why screening can't explain it all. N. Engl. J. Med. 374, 1605–1607 (2016).
Cremolini, C. et al. First-line chemotherapy for mCRC — a review and evidence-based algorithm. Nat. Rev. Clin. Oncol. 12, 607–619 (2015). Evidence for a new patient-oriented algorithm to guide clinicians' decisions on the best choice of upfront therapy for metastatic CRC.
Dienstmann, R., Salazar, R. & Tabernero, J. Overcoming resistance to anti-EGFR therapy in colorectal cancer. Am. Soc. Clin. Oncol. Educ. Book https://dx.doi.org/10.14694/EdBook_AM.2015.35.e149 (2015).
Fearon, E. R. Molecular genetics of colorectal cancer. Ann. Rev. Pathol. 6, 479–507 (2011).
Fearon, E. R. & Vogelstein, B. A genetic model for colorectal tumorigenesis. Cell 61, 759–767 (1990).
Vogelstein, B. et al. Cancer genome landscapes. Science 339, 1546–1558 (2013).
Vogelstein, B. et al. Genetic alterations during colorectal-tumor development. N. Engl. J. Med. 319, 525–532 (1988).
Sjoblom, T. et al. The consensus coding sequences of human breast and colorectal cancers. Science 314, 268–274 (2006).
Cancer Genome Atlas Network. Comprehensive molecular characterization of human colon and rectal cancer. Nature 487, 330–337 (2012). The Cancer Genome Atlas genome-scale analysis of 276 samples, including exome sequencing, and analysis of DNA copy number, promoter methylation and expression of mRNA and microRNA.
Seshagiri, S. et al. Recurrent R-spondin fusions in colon cancer. Nature 488, 660–664 (2012).
Matano, M. et al. Modeling colorectal cancer using CRISPR-Cas9-mediated engineering of human intestinal organoids. Nat. Med. 21, 256–262 (2015). This article uncovers the importance of driver gene mutations and copy number alterations by using genome editing technology in human intestinal epithelium organoids.
Pino, M. S. & Chung, D. C. The chromosomal instability pathway in colon cancer. Gastroenterology 138, 2059–2072 (2010).
Vilar, E. & Tabernero, J. Molecular dissection of microsatellite instable colorectal cancer. Cancer Discov. 3, 502–511 (2013).
Rex, D. K. et al. Serrated lesions of the colorectum: review and recommendations from an expert panel. Am. J. Gastroenterol. 107, 1315–1329 (2012).
Giannakis, M. et al. RNF43 is frequently mutated in colorectal and endometrial cancers. Nat. Genet. 46, 1264–1266 (2014).
Amatu, A. et al. Novel CAD-ALK gene rearrangement is drugable by entrectinib in colorectal cancer. Br. J. Cancer. 113, 1730–1734 (2015).
Le Rolle, A. F. et al. Identification and characterization of RET fusions in advanced colorectal cancer. Oncotarget 6, 28929–28937 (2015).
Stransky, N., Cerami, E., Schalm, S., Kim, J. L. & Lengauer, C. The landscape of kinase fusions in cancer. Nat. Commun. 5, 4846 (2014).
Budinska, E. et al. Gene expression patterns unveil a new level of molecular heterogeneity in colorectal cancer. J. Pathol. 231, 63–76 (2013).
De Sousa E Melo, F. et al. Poor-prognosis colon cancer is defined by a molecularly distinct subtype and develops from serrated precursor lesions. Nat. Med. 19, 614–618 (2013).
Marisa, L. et al. Gene expression classification of colon cancer into molecular subtypes: characterization, validation, and prognostic value. PLoS Med. 10, e1001453 (2013).
Perez-Villamil, B. et al. Colon cancer molecular subtypes identified by expression profiling and associated to stroma, mucinous type and different clinical behavior. BMC Cancer 12, 260 (2012).
Roepman, P. et al. Colorectal cancer intrinsic subtypes predict chemotherapy benefit, deficient mismatch repair and epithelial-to-mesenchymal transition. Int. J. Cancer 134, 552–562 (2014).
Sadanandam, A. et al. A colorectal cancer classification system that associates cellular phenotype and responses to therapy. Nat. Med. 19, 619–625 (2013).
Schlicker, A. et al. Subtypes of primary colorectal tumors correlate with response to targeted treatment in colorectal cell lines. BMC Med. Genomics 5, 66 (2012).
Guinney, J. et al. The consensus molecular subtypes of colorectal cancer. Nat. Med. 21, 1350–1356 (2015). Cross-comparison of six independent gene expression-based classification systems revealed marked interconnectivity coalescing into four consensus molecular subtypes with distinguished biological features.
Isella, C. et al. Stromal contribution to the colorectal cancer transcriptome. Nat. Genet. 47, 312–319 (2015).
Dunne, P. D. et al. Challenging the cancer molecular stratification dogma: intratumoral heterogeneity undermines consensus molecular subtypes and potential diagnostic value in colorectal cancer. Clin. Cancer Res. 22, 4095–4104 (2016).
Yun, J. et al. Glucose deprivation contributes to the development of KRAS pathway mutations in tumor cells. Science 325, 1555–1559 (2009).
Son, J. et al. Glutamine supports pancreatic cancer growth through a KRAS-regulated metabolic pathway. Nature 496, 101–105 (2013).
Ying, H. et al. Oncogenic Kras maintains pancreatic tumors through regulation of anabolic glucose metabolism. Cell 149, 656–670 (2012).
Brunelli, L., Caiola, E., Marabese, M., Broggini, M. & Pastorelli, R. Capturing the metabolomic diversity of KRAS mutants in non-small-cell lung cancer cells. Oncotarget 5, 4722–4731 (2014).
Kamphorst, J. J. et al. Hypoxic and Ras-transformed cells support growth by scavenging unsaturated fatty acids from lysophospholipids. Proc. Natl Acad. Sci. USA 110, 8882–8887 (2013).
Kerr, E. M., Gaude, E., Turrell, F. K., Frezza, C. & Martins, C. P. Mutant Kras copy number defines metabolic reprogramming and therapeutic susceptibilities. Nature 531, 110–113 (2016).
Schwitalla, S. et al. Intestinal tumorigenesis initiated by dedifferentiation and acquisition of stem-cell-like properties. Cell 152, 25–38 (2013).
Fessler, E. et al. TGFβ signaling directs serrated adenomas to the mesenchymal colorectal cancer subtype. EMBO Mol. Med. 8, 745–760 (2016). In a model system for sessile serrated adenomas, BRAFV600E mutation and high TGFβ levels are shown to play an important part in the development of mesenchymal CMS4 tumours.
Tosolini, M. et al. Clinical impact of different classes of infiltrating T cytotoxic and helper cells (Th1, Th2, Treg, Th17) in patients with colorectal cancer. Cancer Res. 71, 1263–1271 (2011).
Galon, J. et al. Type, density, and location of immune cells within human colorectal tumors predict clinical outcome. Science 313, 1960–1964 (2006).
Bindea, G. et al. Spatiotemporal dynamics of intratumoral immune cells reveal the immune landscape in human cancer. Immunity 39, 782–795 (2013).
Galon, J. et al. Cancer classification using the Immunoscore: a worldwide task force. J. Transl Med. 10, 205 (2012).
Galon, J. et al. Validation of the Immunoscore (IM) as a prognostic marker in stage I/II/III colon cancer: results of a worldwide consortium-based analysis of 1,336 patients. J. Clin. Oncol. 34, abstr.3500 (2016).
Saito, T. et al. Two FOXP3+CD4+ T cell subpopulations distinctly control the prognosis of colorectal cancers. Nat. Med. 22, 679–684 (2016).
Llosa, N. J. et al. The vigorous immune microenvironment of microsatellite instable colon cancer is balanced by multiple counter-inhibitory checkpoints. Cancer Discov. 5, 43–51 (2015).
Angelova, M. et al. Characterization of the immunophenotypes and antigenomes of colorectal cancers reveals distinct tumor escape mechanisms and novel targets for immunotherapy. Genome Biol. 16, 64 (2015). Comprehensive analyses of immunophenotypes and the anti-genome of CRC identify non-hypermutated tumours as being enriched with immunosuppressive cells.
Becht, E. et al. Immune and stromal classification of colorectal cancer is associated with molecular subtypes and relevant for precision immunotherapy. Clin. Cancer Res. 22, 4057–4066 (2016). This study shows that immunocharacterization of consensus molecular subtypes associates immunosuppressive CRC with a mesenchymal gene-expression signature.
Gatalica, Z. et al. Programmed cell death 1 (PD-1) and its ligand (PD-L1) in common cancers and their correlation with molecular cancer type. Cancer Epidemiol. Biomarkers Prev. 23, 2965–2970 (2014).
Maby, P. et al. Correlation between density of CD8+ T-cell infiltrate in microsatellite unstable colorectal cancers and frameshift mutations: a rationale for personalized immunotherapy. Cancer Res. 75, 3446–3455 (2015).
Chun, E. et al. CCL2 promotes colorectal carcinogenesis by enhancing polymorphonuclear myeloid-derived suppressor cell population and function. Cell Rep. 12, 244–257 (2015).
Grivennikov, S. I. et al. Adenoma-linked barrier defects and microbial products drive IL-23/IL-17-mediated tumour growth. Nature 491, 254–258 (2012). Barrier deterioration induced by initiating genomic alterations in CRC results in adenoma invasion by microbial products that trigger tumour-elicited inflammation.
Mlecnik, B. et al. The tumor microenvironment and Immunoscore are critical determinants of dissemination to distant metastasis. Sci. Transl Med. 8, 327ra26 (2016). Supervised analyses of metastatic versus primary CRC samples show microenvironment factors (immunoscore and lymphatic invasion) as key determinants of relapse.
Yashiro, M. Ulcerative colitis-associated colorectal cancer. World J. Gastroenterol. 20, 16389–16397 (2014).
Luke, J. J. Correlation of WNT/β-catenin pathway activation with immune exclusion across most human cancers. J. Clin. Oncol. 34, abstr. 3004 (2016).
Reilley, M. et al. Immunologic profiling of consensus molecular subtype stratified colorectal cancer primary and liver metastectomy specimens: implications for immune targeting of proficient mismatch repair CRC. J. Clin. Oncol. 34, abstr. 3520 (2016).
Brannon, A. R. et al. Comparative sequencing analysis reveals high genomic concordance between matched primary and metastatic colorectal cancer lesions. Genome Biol. 15, 454 (2014).
Kopetz, S. et al. Mutation and copy number discordance in primary versus metastatic colorectal cancer (mCRC). J. Clin. Oncol. 32, abstr. 3509 (2014).
Arena, S. et al. Emergence of multiple EGFR extracellular mutations during cetuximab treatment in colorectal cancer. Clin. Cancer Res. 21, 2157–2166 (2015).
Bettegowda, C. et al. Detection of circulating tumor DNA in early- and late-stage human malignancies. Sci. Transl Med. 6, 224ra24 (2014). This article shows that circulating tumour DNA has high accuracy for detection of emerging KRAS -mutated clones during anti-EGFR therapy in advanced CRC.
Misale, S. et al. Emergence of KRAS mutations and acquired resistance to anti-EGFR therapy in colorectal cancer. Nature 486, 532–536 (2012).
Morelli, M. P. et al. Characterizing the patterns of clonal selection in circulating tumor DNA from patients with colorectal cancer refractory to anti-EGFR treatment. Ann. Oncol. 26, 731–736 (2015).
Siravegna, G. et al. Clonal evolution and resistance to EGFR blockade in the blood of colorectal cancer patients. Nat. Med. 21, 795–801 (2015). This study identifies novel resistance mechanisms to EGFR mAbs and clonal dynamics during and after therapy.
Russo, M. et al. Tumor heterogeneity and lesion-specific response to targeted therapy in colorectal cancer. Cancer Discov. 6, 147–153 (2016).
Mohan, S. et al. Changes in colorectal carcinoma genomes under anti-EGFR therapy identified by whole-genome plasma DNA sequencing. PLoS Genet. 10, e1004271 (2014).
Price, T. J. et al. Prevalence and outcomes of patients (pts) with EGFR S492R ectodomain mutations in ASPECCT: panitumumab (pmab) versus cetuximab (cmab) in pts with chemorefractory wild-type KRAS exon 2 metastatic colorectal cancer (mCRC). J. Clin. Oncol. 33, abstr. 740 (2015).
Bertotti, A. et al. The genomic landscape of response to EGFR blockade in colorectal cancer. Nature 526, 263–267 (2015). This study shows that therapeutic resistance to EGFR blockade can be overcome in tumour graft models through combinatorial therapies targeting actionable genes.
Esposito, C. et al. The S492R EGFR ectodomain mutation is never detected in KRAS wild-type colorectal carcinoma before exposure to EGFR monoclonal antibodies. Cancer Biol. Ther. 14, 1143–1146 (2013).
Roessler, S. et al. Integrative genomic and transcriptomic characterization of matched primary and metastatic liver and colorectal carcinoma. Int. J. Biol. Sci. 11, 88–98 (2015).
Stange, D. E. et al. Expression of an ASCL2 related stem cell signature and IGF2 in colorectal cancer liver metastases with 11p15.5 gain. Gut 59, 1236–1244 (2010).
Sun, Y. L. et al. Expression of HGF and Met in human tissues of colorectal cancers: biological and clinical implications for synchronous liver metastasis. Int. J. Med. Sci. 10, 548–559 (2013).
Munoz-Bellvis, L. et al. Unique genetic profile of sporadic colorectal cancer liver metastasis versus primary tumors as defined by high-density single-nucleotide polymorphism arrays. Mod. Pathol. 25, 590–601 (2012).
Bertotti, A. et al. A molecularly annotated platform of patient-derived xenografts (“xenopatients”) identifies HER2 as an effective therapeutic target in cetuximab-resistant colorectal cancer. Cancer Discov. 1, 508–523 (2011).
Bardelli, A. et al. Amplification of the MET receptor drives resistance to anti-EGFR therapies in colorectal cancer. Cancer Discov. 3, 658–673 (2013).
Dienstmann, R. et al. Safety and activity of the first-in-class Sym004 anti-EGFR antibody mixture in patients with refractory colorectal cancer. Cancer Discov. 5, 598–609 (2015).
Sartore-Bianchi, A. et al. Dual-targeted therapy with trastuzumab and lapatinib in treatment-refractory, KRAS codon 12/13 wild-type, HER2-positive metastatic colorectal cancer (HERACLES): a proof-of-concept, multicentre, open-label, phase 2 trial. Lancet Oncol. 17, 738–746 (2016).
Sveen, A. et al. Intra-patient inter-metastatic genetic heterogeneity in colorectal cancer as a key determinant of survival after curative liver resection. PLoS Genet. 12, e1006225 (2016).
Sottoriva, A. et al. A Big Bang model of human colorectal tumor growth. Nat. Genet. 47, 209–216 (2015). In this study, genomic profiling of individual glands from colorectal tumours uncovers a new model for intra-tumour heterogeneity.
Kopetz, S. et al. Phase II pilot study of vemurafenib in patients with metastatic BRAF-mutated colorectal cancer. J. Clin. Oncol. 33, 4032–4038 (2015).
Zimmer, L. et al. Phase I expansion and pharmacodynamic study of the oral MEK inhibitor RO4987655 (CH4987655) in selected patients with advanced cancer with RAS–RAF mutations. Clin. Cancer Res. 20, 4251–4261 (2014).
Van Cutsem, E. et al. Updated results of the MEK inhibitor trametinib (T), BRAF inhibitor dabrafenib (D), and anti-EGFR antibody panitumumab (P) in patients (pts) with BRAF V600E mutated (BRAFm) metastatic colorectal cancer (mCRC). Ann. Oncol. 26, iv119 (2015).
Guinney, J. et al. Modeling RAS phenotype in colorectal cancer uncovers novel molecular traits of RAS dependency and improves prediction of response to targeted agents in patients. Clin. Cancer Res. 20, 265–272 (2014).
Tian, S. et al. A combined oncogenic pathway signature of BRAF, KRAS and PI3KCA mutation improves colorectal cancer classification and cetuximab treatment prediction. Gut 62, 540–549 (2013).
Van Cutsem, E. et al. Fluorouracil, leucovorin, and irinotecan plus cetuximab treatment and RAS mutations in colorectal cancer. J. Clin. Oncol. 33, 692–700 (2015).
Douillard, J. Y. et al. Panitumumab–FOLFOX4 treatment and RAS mutations in colorectal cancer. N. Engl. J. Med. 369, 1023–1034 (2013).
Sorich, M. J. et al. Extended RAS mutations and anti-EGFR monoclonal antibody survival benefit in metastatic colorectal cancer: a meta-analysis of randomized, controlled trials. Ann. Oncol. 26, 13–21 (2015).
Dono, M. et al. Low percentage of KRAS mutations revealed by locked nucleic acid polymerase chain reaction: implications for treatment of metastatic colorectal cancer. Mol. Med. 18, 1519–1526 (2012).
Laurent-Puig, P. et al. Clinical relevance of KRAS-mutated subclones detected with picodroplet digital PCR in advanced colorectal cancer treated with anti-EGFR therapy. Clin. Cancer Res. 21, 1087–1097 (2015).
Dienstmann, R. & Cervantes, A. Heterogeneity of driver genes and therapeutic implications in colorectal cancer. Ann. Oncol. 26, 1523–1525 (2015).
Kavuri, S. M. et al. HER2 activating mutations are targets for colorectal cancer treatment. Cancer Discov. 5, 832–841 (2015).
De Roock, W. et al. Effects of KRAS, BRAF, NRAS, and PIK3CA mutations on the efficacy of cetuximab plus chemotherapy in chemotherapy-refractory metastatic colorectal cancer: a retrospective consortium analysis. Lancet Oncol. 11, 753–762 (2010).
Weickhardt, A. J. et al. Dual targeting of the epidermal growth factor receptor using the combination of cetuximab and erlotinib: preclinical evaluation and results of the phase II DUX study in chemotherapy-refractory, advanced colorectal cancer. J. Clin. Oncol. 30, 1505–1512 (2012).
Kearns, J. D. et al. Enhanced targeting of the EGFR network with MM-151, an oligoclonal anti-EGFR antibody therapeutic. Mol. Cancer Ther. 14, 1625–1636 (2015).
Montagut, C. et al. Identification of a mutation in the extracellular domain of the Epidermal Growth Factor Receptor conferring cetuximab resistance in colorectal cancer. Nat. Med. 18, 221–223 (2012).
Juric, D. et al. Safety and pharmacokinetics/pharmacodynamics of the first-in-class dual action HER3/EGFR antibody MEHD7945A in locally advanced or metastatic epithelial tumors. Clin. Cancer Res. 21, 2462–2470 (2015).
Sclafani, F. et al. A randomized phase II/III study of dalotuzumab in combination with cetuximab and irinotecan in chemorefractory, KRAS wild-type, metastatic colorectal cancer. J. Natl Cancer Inst. 107, djv258 (2015).
Van Cutsem, E. et al. Randomized phase Ib/II trial of rilotumumab or ganitumab with panitumumab versus panitumumab alone in patients with wild-type KRAS metastatic colorectal cancer. Clin. Cancer Res. 20, 4240–4250 (2014).
Rimassa, L. et al. Phase II study of tivantinib (ARQ 197) in combination with cetuximab in EGFR inhibitor-resistant, MET-high, KRAS wild-type (KRASwt) metastatic colorectal cancer (mCRC). Ann. Oncol. 26, iv108–iv116 (2015).
Iida, M. et al. Sym004, a novel EGFR antibody mixture, can overcome acquired resistance to cetuximab. Neoplasia 15, 1196–1206 (2013).
Hobor, S. et al. TGFα and amphiregulin paracrine network promotes resistance to EGFR blockade in colorectal cancer cells. Clin. Cancer Res. 20, 6429–6438 (2014).
Kawakami, H. et al. The anti-HER3 antibody patritumab abrogates cetuximab resistance mediated by heregulin in colorectal cancer cells. Oncotarget 5, 11847–11856 (2014).
Luraghi, P. et al. MET signaling in colon cancer stem-like cells blunts the therapeutic response to EGFR inhibitors. Cancer Res. 74, 1857–1869 (2014).
Misale, S. et al. Vertical suppression of the EGFR pathway prevents onset of resistance in colorectal cancers. Nat. Commun. 6, 8305 (2015). EGFR–MEK combined blockade prevents development of resistance in 'all wild-type' advanced CRC by triggering apoptosis in minor KRAS -mutated clones.
Deming, D. A. et al. A phase I study of selumetinib (AZD6244/ARRY-142866), a MEK1/2 inhibitor, in combination with cetuximab in refractory solid tumors and KRAS mutant colorectal cancer. Invest. New Drugs 34, 168–175 (2015).
Brule, S. Y. et al. Location of colon cancer (right-sided versus left-sided) as a prognostic factor and a predictor of benefit from cetuximab in NCIC CO.17. Eur. J. Cancer 51, 1405–1414 (2015).
Venook, A. P. et al. Impact of primary tumor location on overall survival and progression-free survival in patients with metastatic colorectal cancer: analysis of CALGB/SWOG 80405 (Alliance). J. Clin. Oncol. 34, abstr. 3504 (2016).
Lee, M. S. et al. Association of primary site and molecular features with progression-free survival and overall survival of metastatic colorectal cancer after anti-epidermal growth factor receptor therapy. J. Clin. Oncol. 34, abstr. 3506 (2016).
Missiaglia, E. et al. Distal and proximal colon cancers differ in terms of molecular, pathological, and clinical features. Ann. Oncol. 25, 1995–2001 (2014).
Laurent-Puig, P. et al. Analysis of PTEN, BRAF, and EGFR status in determining benefit from cetuximab therapy in wild-type KRAS metastatic colon cancer. J. Clin. Oncol. 27, 5924–5930 (2009).
Zanella, E. R. et al. IGF2 is an actionable target that identifies a distinct subpopulation of colorectal cancer patients with marginal response to anti-EGFR therapies. Sci. Transl Med. 7, 272ra12 (2015).
Trinh, A. et al. Practical and robust identification of molecular subtypes in colorectal cancer by immunohistochemistry. Clin. Cancer Res. http://dx.doi.org/10.1158/1078-0432.ccr-16-0680 (2016). Development and validation of a panel of five immunohistochemical markers to identify CRC gene-expression subtypes. This panel showed a poor prognosis for patients with mesenchymal-like tumours, as well as the benefit of adding cetuximab to bevacizumab and chemotherapy in patients with RAS wild-type cancers of the epithelial-like subtypes.
Oliveras-Ferraros, C. et al. Stem cell property epithelial-to-mesenchymal transition is a core transcriptional network for predicting cetuximab (Erbitux™) efficacy in KRAS wild-type tumor cells. J. Cell. Biochem. 112, 10–29 (2011).
Elez, E. et al. Abituzumab combined with cetuximab plus irinotecan versus cetuximab plus irinotecan alone for patients with KRAS wild-type metastatic colorectal cancer: the randomised phase I/II POSEIDON trial. Ann. Oncol. 26, 132–140 (2015).
Do, K. et al. Biomarker-driven phase 2 study of MK-2206 and selumetinib (AZD6244, ARRY-142886) in patients with colorectal cancer. Invest. New Drugs 33, 720–728 (2015).
Liu, Y. et al. Metabolic and functional genomic studies identify deoxythymidylate kinase as a target in LKB1-mutant lung cancer. Cancer Discov. 3, 870–879 (2013).
Kim, H. S. et al. Systematic identification of molecular subtype-selective vulnerabilities in non-small-cell lung cancer. Cell 155, 552–566 (2013).
Skoulidis, F. et al. Co-occurring genomic alterations define major subsets of KRAS-mutant lung adenocarcinoma with distinct biology, immune profiles, and therapeutic vulnerabilities. Cancer Discov. 5, 860–877 (2015).
Yun, J. et al. Vitamin C selectively kills KRAS and BRAF mutant colorectal cancer cells by targeting GAPDH. Science 350, 1391–1396 (2015).
Gross, M. I. et al. Antitumor activity of the glutaminase inhibitor CB-839 in triple-negative breast cancer. Mol. Cancer Ther. 13, 890–901 (2014).
Ventura, R. et al. Inhibition of de novo palmitate synthesis by fatty acid synthase induces apoptosis in tumor cells by remodeling cell membranes, inhibiting signaling pathways, and reprogramming gene expression. EBioMedicine 2, 806–822 (2015).
Straussman, R. et al. Tumour micro-environment elicits innate resistance to RAF inhibitors through HGF secretion. Nature 487, 500–504 (2012).
Song, N. et al. Clinical outcome from oxaliplatin treatment in stage II/III colon cancer according to intrinsic subtypes: secondary analysis of NSABP C-07/NRG oncology randomized clinical trial. JAMA Oncol. 2, 1162–1169 (2016). This retrospective biomarker analysis of a randomized clinical trial (NSABP C-07) finds that patients with tumours harbouring an 'enterocyte' subtype have the highest benefit from oxaliplatin as adjuvant chemotherapy in stage III disease. An adapted version of the CMS classifier robustly identifies CMS4 mesenchymal patients with poor prognosis.
Calon, A. et al. Stromal gene expression defines poor-prognosis subtypes in colorectal cancer. Nat. Genet. 47, 320–329 (2015). TGFβ derived from cancer-associated fibroblasts increases the frequency of tumour-initiating cells, and TGFβ-signalling inhibitors block the crosstalk between cancer cells and the microenvironment, halting disease progression in preclinical models.
MoTriColor. Molecularly guided trials with specific treatment strategies in patients with advanced newly molecular defined subtypes of colorectal cancer. CORDIS http://cordis.europa.eu/project/rcn/193328_es.html (2015).
Sonoshita, M. et al. Promotion of colorectal cancer invasion and metastasis through activation of NOTCH-DAB1-ABL-RHOGEF protein TRIO. Cancer Discov. 5, 198–211 (2015).
Dunne, P. D. et al. AXL is a key regulator of inherent and chemotherapy-induced invasion and predicts a poor clinical outcome in early-stage colon cancer. Clin. Cancer Res. 20, 164–175 (2014).
Schaer, D. et al. Targeting the TGFb pathway with galunisertib, a TGFbRI SMI, promotes antitumor immunity leading to durable, complete responses, as monotherapy and in combination with checkpoint inhibition. Cancer Immunol. Res. 4, A091 (2016).
Triplett, T. A. et al. STAT3 signaling is required for optimal regression of large established tumors in mice treated with anti-OX40 and TGFβ receptor blockade. Cancer Immunol. Res. 3, 526–535 (2015).
Young, K. H. et al. TGFβ inhibition prior to hypofractionated radiation enhances efficacy in preclinical models. Cancer Immunol. Res. 2, 1011–1022 (2014).
Wang, G. et al. Targeting YAP-dependent MDSC infiltration impairs tumor progression. Cancer Discov. 6, 80–95 (2016).
Smyth, M. J., Ngiow, S. F., Ribas, A. & Teng, M. W. Combination cancer immunotherapies tailored to the tumour microenvironment. Nat. Rev. Clin. Oncol. 13, 143–158 (2015).
Zheng, H. et al. HDAC inhibitors enhance T-cell chemokine expression and augment response to PD-1 immunotherapy in lung adenocarcinoma. Clin. Cancer Res. 22, 4119–4132 (2016).
Tesniere, A. et al. Immunogenic death of colon cancer cells treated with oxaliplatin. Oncogene 29, 482–491 (2010).
Motz, G. T. et al. Tumor endothelium FasL establishes a selective immune barrier promoting tolerance in tumors. Nat. Med. 20, 607–615 (2014).
Ebert, P. J. et al. MAP kinase inhibition promotes T cell and anti-tumor activity in combination with PD-L1 checkpoint blockade. Immunity 44, 609–621 (2016).
Bendell, J. C. et al. Clinical activity and safety of cobimetinib and atezolizumab in colorectal cancer. J. Clin. Oncol. 34, abstr. 3502 (2016).
Perez, E. A. et al. Genomic analysis reveals that immune function genes are strongly linked to clinical outcome in the North Central Cancer Treatment Group N9831 Adjuvant Trastuzumab Trial. J. Clin. Oncol. 33, 701–708 (2015).
Dienstmann, R., Rodon, J. & Tabernero, J. Optimal design of trials to demonstrate the utility of genomically-guided therapy: putting precision cancer medicine to the test. Mol. Oncol. 9, 940–950 (2015).
Alberts, S. R. et al. Effect of oxaliplatin, fluorouracil, and leucovorin with or without cetuximab on survival among patients with resected stage III colon cancer: a randomized trial. JAMA 307, 1383–1393 (2012).
Taieb, J. et al. Oxaliplatin, fluorouracil, and leucovorin with or without cetuximab in patients with resected stage III colon cancer (PETACC-8): an open-label, randomised phase 3 trial. Lancet Oncol. 15, 862–873 (2014).
Allegra, C. J. et al. Bevacizumab in stage II–III colon cancer: 5-year update of the National Surgical Adjuvant Breast and Bowel Project C-08 trial. J. Clin. Oncol. 31, 359–364 (2013).
de Gramont, A. et al. Bevacizumab plus oxaliplatin-based chemotherapy as adjuvant treatment for colon cancer (AVANT): a phase 3 randomised controlled trial. Lancet Oncol. 13, 1225–1233 (2012).
Corcoran, R. B. et al. Combined BRAF and MEK inhibition with dabrafenib and trametinib in BRAF V600-mutant colorectal cancer. J. Clin. Oncol. 33, 4023–4031 (2015).
Ahronian, L. G. et al. Clinical acquired resistance to RAF inhibitor combinations in BRAF-mutant colorectal cancer through MAPK pathway alterations. Cancer Discov. 5, 358–367 (2015).
Oddo, D. et al. Molecular landscape of acquired resistance to targeted therapy combinations in BRAF-mutant colorectal cancer. Cancer Res. 76, 4504–4515 (2016).
Corcoran, R. B. et al. EGFR-mediated re-activation of MAPK signaling contributes to insensitivity of BRAF mutant colorectal cancers to RAF inhibition with vemurafenib. Cancer Discov. 2, 227–235 (2012).
Prahallad, A. et al. Unresponsiveness of colon cancer to BRAF(V600E) inhibition through feedback activation of EGFR. Nature 483, 100–103 (2012). BRAFV600E inhibition in colorectal cancer cells causes a rapid feedback activation of EGFR, supporting the continued proliferation of cancer cells in the presence of the inhibitor. This proliferation can be abrogated if BRAFV600E inhibitors are combined with anti-EGFR therapy.
Tabernero, J. et al. Phase 2 results: encorafenib (ENCO) and cetuximab (CETUX) with or without alpelisib (ALP) in patients with advanced BRAF-mutant colorectal cancer (BRAFm CRC). J. Clin. Oncol. 34, abstr. 3544 (2016).
Ramanathan, R. K. et al. Low overexpression of HER-2/neu in advanced colorectal cancer limits the usefulness of trastuzumab (Herceptin) and irinotecan as therapy. A phase II trial. Cancer Invest. 22, 858–865 (2004).
Medico, E. et al. The molecular landscape of colorectal cancer cell lines unveils clinically actionable kinase targets. Nat. Commun. 6, 7002 (2015).
Sartore-Bianchi, A. et al. Sensitivity to entrectinib associated with a novel LMNA-NTRK1 gene fusion in metastatic colorectal cancer. J. Natl Cancer Inst. 108, djv306 (2016).
van de Wetering, M. et al. Prospective derivation of a living organoid biobank of colorectal cancer patients. Cell 161, 933–945 (2015). Tumour organoids from patients with CRC are amenable to high-throughput drug screens, allowing detection of gene–drug interactions such as RNF43 mutations and porcupine inhibitors.
Madan, B. et al. Wnt addiction of genetically defined cancers reversed by PORCN inhibition. Oncogene 35, 2197–2207 (2015).
Janku, F. et al. Phase I study of WNT974, a first-in-class Porcupine inhibitor, in advanced solid tumors. Mol. Cancer Ther. 14, abstr.C45 (2015).
Storm, E. E. et al. Targeting PTPRK-RSPO3 colon tumours promotes differentiation and loss of stem-cell function. Nature 529, 97–100 (2015).
Le, D. T. et al. PD-1 blockade in tumors with mismatch-repair deficiency. N. Engl. J. Med. 372, 2509–2520 (2015).
Overman, M. J. et al. Nivolumab ± ipilimumab in treatment of patients with metastatic colorectal cancer with and without high microsatellite instability: CheckMate-142 interim results. J. Clin. Oncol. 34, abstr. 3501 (2016).
Giannakis, M. et al. Genomic correlates of immune-cell infiltrates in colorectal cancer. Cell Rep. 15, 857–865 (2016).
Tie, J. K. et al. Circulating tumor DNA as an early marker of therapeutic response in patients with metastatic colorectal cancer. Ann. Oncol. 26, 1715–1722 (2015).
Tie, J. K. et al. Circulating tumor DNA analysis detects minimal residual disease and predicts recurrence in patients with stage II colon cancer. Sci. Transl Med. 8, 346ra92 (2016). This prospective trial implementing massively parallel sequencing-based assays in stage II colon cancer shows the ability of circulating tumour DNA to accurately detect minimal residual disease after surgery.
Acknowledgements
R.D. and J.T. acknowledge the FERO Foundation for supporting research in gastrointestinal malignancies and the Cellex Foundation for providing research facilities and equipment. The work of R.D., S.T. and J.T. was supported by the European Union's Horizon 2020 research and innovation programme 2014–2020 under grant agreement number 635342 (MoTriColor project). The work of S.T. was supported by the University of Leuven (grant GOA/12/2106), the Research Foundation Flanders and the Belgian National Cancer Plan. The work of L.V. was supported by grants from KWF Kankerbestrijding (Dutch Cancer Society) (UVA2014-7245), the European Research Council (ERG-StG 638193) and the Netherlands Organisation for Health Research and Development (Vidi 016.156.308). The work of J.G. was supported by grant U54CA149237 from the US National Cancer Institute. The work of S.C. was supported by the generous philanthropic contributions to the University of Texas MD Anderson Moon Shots Program and Cancer Center Sipport Grants grant 3 P30 CA016672 41. The work of R.D. was supported by the Grant for Oncology Innovation under the project 'Next generation of clinical trials with matched targeted therapies in colorectal cancer'.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Glossary
- Early-stage
-
Tumours that remain localized and have not spread to other parts of the body.
- Advanced-stage
-
Tumours that became metastatic and have spread to other parts of the body.
- Sporadic background
-
Tumours that have no identifiable inherited gene involved in the carcinogenesis process.
- Sessile serrated adenomas
-
Pre-malignant flat (or sessile) polyps, predominantly seen in the right side of the colon. They have been identified as the main precursor lesion in the serrated carcinogenesis process.
- Desmoplastic reaction
-
At histopathological examination, pervasive growth of dense fibrous connective tissue around the tumour.
- Trabecular
-
At histopathological examination, tumours composed of cells structured in a nested pattern.
- Mucinous
-
At histopathological examination, tumours characterized by abundant extracellular accumulation of mucus bound to neoplastic epithelium or stroma.
- Papillary
-
At histopathological examination, tumours demonstrating prominent papillae with fibrovascular cores.
- Late-stage neoplasms
-
Localized tumours that have grown more deeply into nearby tissue or have spread to regional lymph nodes.
- Rechallenge
-
Reintroduction of the same therapy after a drug holiday following disease progression during therapy.
- Metachronous disease
-
Tumours that became metastatic after the diagnosis and treatment of localized disease (usually later than 6 months).
- Selective sweeps
-
These occur when a rare or previously non-existing allele that increases the fitness of the cell — relative to other clonal populations — expands rapidly in frequency as a result of natural selection.
- Underpowered
-
A study with low statistical power of detecting a true effect of practical importance.
- Caecum
-
The first segment of the right colon, an intraperitoneal pouch that connects the ileum with the ascending colon.
- Splenic flexure
-
The first segment of the left colon, a sharp bend under the spleen where the transverse colon joins the descending colon.
- Adjuvant
-
Therapy applied after initial surgical treatment for cancer to avoid tumour relapse.
Rights and permissions
About this article
Cite this article
Dienstmann, R., Vermeulen, L., Guinney, J. et al. Consensus molecular subtypes and the evolution of precision medicine in colorectal cancer. Nat Rev Cancer 17, 79–92 (2017). https://doi.org/10.1038/nrc.2016.126
Published:
Issue Date:
DOI: https://doi.org/10.1038/nrc.2016.126
This article is cited by
-
An integrated framework for prognosis prediction and drug response modeling in colorectal liver metastasis drug discovery
Journal of Translational Medicine (2024)
-
Towards precision oncology discovery: four less known genes and their unknown interactions as highest-performed biomarkers for colorectal cancer
npj Precision Oncology (2024)
-
Phosphorylation-dependent deubiquitinase OTUD3 regulates YY1 stability and promotes colorectal cancer progression
Cell Death & Disease (2024)
-
Discovery of a nitroaromatic nannocystin with potent in vivo anticancer activity against colorectal cancer by targeting AKT1
Acta Pharmacologica Sinica (2024)
-
Sidedness is not a prognostic factor in an unselected cohort of patients with colon cancer but prognosis for caecal carcinoma is worse – A multivariate analysis of a large single institution database
International Journal of Colorectal Disease (2024)