Abstract
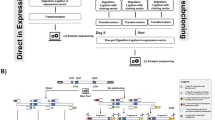
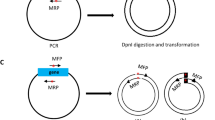
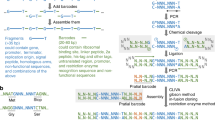
We developed an efficient "Combinatorial Strategy” for cloning different vectors with various clone sites. 1) Using originally existed clone sites from circular vectors to prepare the inserts, and if no appropriate sites are available, performing SDM to create compatible sites, to achieve maximal correct digestion of the inserts. 2) Different vectors were digested with various restriction endonucleases, and then dephosphorylated with CIP. 3) Top10 competent cells were used for transformation to increase the transformant colonies. Our results showed that, when blunt sites, or a Xba I site was adopted for ligation, the percentages of positive clones were about 50%. Whereas, when different sites, including one blunt and another Pst I sites, Not I and Xho I sites, the percentages of positive clones were nearly 100%. Using this strategy, most vectors could be successfully cloned through “one ligation, one transformation, three to five minipreps."
Similar content being viewed by others
Article PDF
Author information
Authors and Affiliations
Corresponding authors
Rights and permissions
About this article
Cite this article
Zhang, G., Tandon, A. Quantitative Models for Efficient Cloning of Different Vectors with Various Clone sites. Nat Prec (2012). https://doi.org/10.1038/npre.2012.6965.1
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/npre.2012.6965.1