Abstract
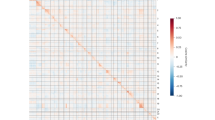
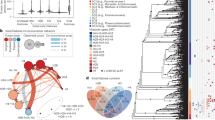
Genomes maybe organized as networks where protein-protein association plays the role of network links. The resulting networks are far from being random and their topological properties are a consequence of the underlying mechanisms for genome evolution. Considering data on protein-protein association networks from STRING database, we present experimental evidence that degree distribution is not scale free, presenting an increased probability for high degree nodes. We also show that the degree distribution approaches a scale invariant state as the number of genes in the network increases, although real genomes still present finite size effects. Based on the experimental evidence unveiled by these data analyses, we propose a simulation model for genome evolution, where genes in a network are either acquired de novo using a preferential attachment rule, or duplicated, with a duplication probability that linearly grows with gene degree and decreases with its clustering coefficient. The results show that topological distributions are better described than in previous genome evolution models. This model correctly predicts that, in order to produce protein-protein association networks with number of links and number of nodes in the observed range, it is necessary 90% of gene duplication and 10% of de novo gene acquisition. If this scenario is true, it implies a universal mechanism for genome evolution.
Similar content being viewed by others
Article PDF
Author information
Authors and Affiliations
Corresponding authors
Rights and permissions
About this article
Cite this article
Dalmolin, R., de Almeida, R., Brunnet, L. et al. Evolution signatures in genome network properties. Nat Prec (2011). https://doi.org/10.1038/npre.2011.6653.1
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/npre.2011.6653.1