Abstract
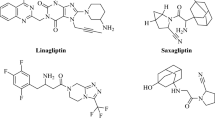
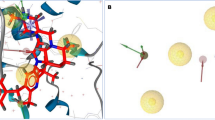
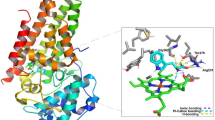
Design of potential drug-like candidates for cardiovascular therapy is of interest in recent years. Elevated level of IGFBP-6 leads to perilous diseases like atherosclerosis, high output cardiac failure and myocardial infarction. The current IGFBP-6 inhibitors available in clinical practice are not having satisfactory anti-cardiovascular effects, leaving room for further improvement. Therefore, with an aim to propose a potent inhibitor to arrest the cardiovascular disease caused by IGFBP-6, tools of computer-aided drug designing were used for virtual screening from small molecule databases. Accordingly, IGFBP-6 tertiary structure was solved through nuclear magnetic resonance techniques retrieved from the protein data bank. Fifteen IGFBP-6 inhibitors were acquired from literature databases and subsequent 2D searching protocol of Ligand.Info yielded 5759 structural analogs. The 3D structural conversion and multiple conformations for each compound were generated using LigPrep with constraints of ADME evaluation and toxicity assessments. The docking and scoring calculations were performed using Glide v5.7. The Glide extra precision (XP) docking had reported 138 leads and ranked based on XP Gscore. Six leads having better XP Gscore compared to current IGFBP-6 inhibitors were proposed as potential inhibitors. Chelidamic acid showed the highest XP Gscore (-7.094 Kcal/mol) with good pharmacological properties and molecular interaction with IGFBP-6. Therefore, chelidamic acid is proposed for rational drug design towards cardiovascular therapy.
Similar content being viewed by others
Article PDF
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Sravani, B., Pradhan, D. & Umamaheswari, A. Docking-based virtual screening for the exploration of potential antagonists for human IGFBP6. Nat Prec (2011). https://doi.org/10.1038/npre.2011.6525.1
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/npre.2011.6525.1