Abstract
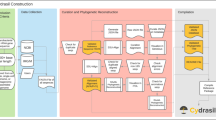
The CloVR-16S pipeline employs several well-known phylogenetic tools and protocols for the analysis of 16S rRNA sequence datasets:A) Mothur – a C++-based software package used for clustering 16S rRNA sequences into operational taxonomic units (OTUs). Mothur creates OTUs using a matrix that describes pairwise distances between representative aligned sequences and subsequently estimates within-sample diversity (alpha diversity);B) The Ribosomal Database (RDP) naïve Bayesian classifier assigns each 16S sequence to a reference taxonomy with associated empirical probabilities based on oligonucleotide frequencies;C) Qiime – a python-based workflow package, allowing for sequence processing and phylogenetic analysis using different methods including phylogenetic distance (UniFrac) for within-(alpha diversity) and between-(beta diversity) sample analysis;D) Metastats and custom R scripts used to generate additional statistical and graphical evaluations.Though some of the different protocols used in CloVR-16S overlap in purpose (e.g. OTU clustering), the end-user benefits from their overall complementary nature as they focus on different aspects of the phylogenetic analysis. CloVR-16S accepts as input raw multiplex 454-pyrosequencer output, i.e. pooled pyrotagged sequences from multiple samples, or alternatively, pre-processed sequences from multiple samples in separate files. This protocol is available in CloVR beta versions 0.5 and 0.6.
Similar content being viewed by others
Article PDF
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Fricke, W., White, J., Arze, C. et al. CloVR-16S: Phylogenetic microbial community composition analysis based on 16S ribosomal RNA amplicon sequencing – standard operating procedure, version 1.0. Nat Prec (2011). https://doi.org/10.1038/npre.2011.5888.2
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/npre.2011.5888.2