Abstract
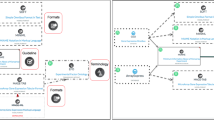
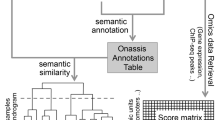
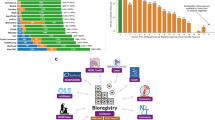
*See also "related poster":http://precedings.nature.com/documents/3144/version/1*Today’s researchers can perform biological and biomedical studies where the same material is run through a wide range of assays, comprising several technologies such as genomics, transcriptomics, proteomics and metabol/nomics (hereafter referred as ‘omics’). To enable others to correctly interpret the complex data sets that result, and the conclusions drawn, it is necessary to provide contextualizing experimental metadata at an appropriate level of granularity.Standards initiatives normally cater to particular domains. However, several synergistic standards activities foster cross-domain harmonization of the three kinds of reporting standard (minimum information checklists, ontologies and file formats). Some 29 groups participate in the "MIBBI":http://www.mibbi.org project, which offers a one-stop shop for those exploring the range of extant ‘minimum information’ checklists, and which fosters integrative development^1^. More than 60 groups participate in the "OBO Foundry":http://www.obofoundry.org ^2^, which coordinates the orthogonal development of ontologies such as "OBI":http://obi-ontology.org for describing experimental (meta)data. And several groups participate in the development of "ISA-Tab":http://isatab.sf.net, a tabular framework for presenting experimental metadata^3^ (analogous to "FuGE":http://fuge.sf.net, a generic data model to underpin various XML file formats^4^).We have developed an infrastructure that leverages the aforementioned synergistic reporting standards to create a common structured representation and storage mechanism for experimental metadata from biological and biomedical investigations ranging from simple single-assay studies to complex, methodologically diverse multi-assay studies.View the "public instance":http://www.ebi.ac.uk/bioinvindex of our ISA-based infrastructure, running at EBI, and/or "download the components":http://isatab.sf.net for your local use.*References*1. Taylor CF, Field D, Sansone SA,… Rocca-Serra P et al. (2008) The MIBBI Project. Nature Biotechnology Aug;26(8):889-896. "http://www.mibbi.org":http://www.mibbi.org2. Smith B, Ashburner M, Rosse C,… Rocca-Serra P, …Sansone SA et al. (2007) The OBO Foundry. Nature Biotechnology Nov;25(11):1251-5. "http://www.obofoundry.org":http://www.obofoundry.org3. Sansone SA, Rocca-Serra P, Brandizi M,… Taylor CF et al. (2008) The First MGED RSBI (ISA-TAB) Workshop. OMICS. Jun;12(2):143-9. "http://isatab.sf.net":http://isatab.sf.net4. Jones AR, Miller M, Aebersold R,… Sansone SA et al. (2007) The Functional Genomics Experiment model (FuGE). Nature Biotechnology Oct;25(10):1127-1133. "http://fuge.sf.net":http://fuge.sf.net
Similar content being viewed by others
Article PDF
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Sansone, SA., Rocca-Serra, P. Standards and infrastructure for managing experimental metadata. Nat Prec (2009). https://doi.org/10.1038/npre.2009.3145.1
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/npre.2009.3145.1