Abstract
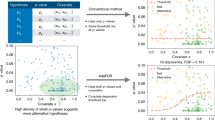
An application of great interest in microarray data analysis is the identification of a group of genes that show very similar patterns of expression in a data set, and are expected to represent groups of genes that perform common/similar functions, also known as functional modules. Although clustering offers a natural solution to this problem, it suffers from the limitation that it uses all the conditions to compare two genes, whereas only a subset of them may be relevant. Association analysis offers an alternative route for finding such groups of genes that may be co-expressed only over a subset of the experimental conditions used to prepare the data set. The techniques in this field attempt to find groups of data objects that contain coherent values across a set of attributes, in an exhaustive and efficient manner. In this paper, we illustrate how a generalization of the techniques in association analysis for real-valued data can be utilized to extract coherent functional modules from large microarray data sets.
Similar content being viewed by others
Article PDF
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Pandey, G., Atluri, G., Steinbach, M. et al. Association Analysis Techniques for Discovering Functional Modules from Microarray Data. Nat Prec (2008). https://doi.org/10.1038/npre.2008.2184.1
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/npre.2008.2184.1
Keywords
This article is cited by
-
Frequent itemset mining using FP-tree: a CLA-based approach and its extended application in biodiversity data
Innovations in Systems and Software Engineering (2022)