Abstract
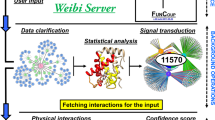
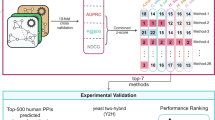
In recent years, in-silico literature-based mammalian protein-protein interaction network datasets have been developed. These datasets contain binary interactions extracted manually from legacy experimental biomedical research literature. Placing lists of genes or proteins identified as significantly changing in multivariate experiments, in the context of background knowledge about binary interactions, can be used to place these genes or proteins in the context of pathways and protein complexes.Genes2Networks is a software system that integrates the content of ten mammalian literature-based interaction network datasets. Filtering to prune low-confidence interactions was implemented. Genes2Networks is delivered as a web-based service using AJAX. The system can be used to extract relevant subnetworks created from “seed” lists of human Entrez gene names. The output includes a dynamic linkable three color web-based network map, with a statistical analysis report that identifies significant intermediate nodes used to connect the seed list. Genes2Networks is available at http://actin.pharm.mssm.edu/genes2networks.Genes2Network is a powerful web-based software application tool that can help experimental biologists to interpret high-throughput experimental results used in genomics and proteomics studies where the output of these experiments is a list of significantly changing genes or proteins. The system can be used to find relationships between nodes from the seed list, and predict novel nodes that play a key role in a common function.
Similar content being viewed by others
Article PDF
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Posner, J., Berger, S. & Ma'ayan, A. Genes2Networks: Connecting Lists of Proteins by Using Background Literature-based Mammalian Networks. Nat Prec (2007). https://doi.org/10.1038/npre.2007.35.2
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/npre.2007.35.2