Abstract
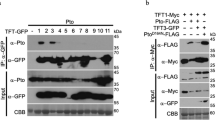
Nucleotide-binding domain and leucine-rich repeat domain-containing (NLR) proteins are sentinels of plant immunity that monitor host proteins for perturbations induced by pathogenic effector proteins. Here we show that the Arabidopsis ZAR1 NLR protein requires the ZRK3 kinase to recognize the Pseudomonas syringae type III effector (T3E) HopF2a. These results support the hypothesis that ZAR1 associates with an expanded ZRK protein family to broaden its effector recognition spectrum.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
References
Büttner, D. & He, S. Y. Plant Physiol. 150, 1656–1664 (2009).
Jones, J. D. & Dangl, J. L. Nature 444, 323–329 (2006).
Collier, S. M. & Moffett, P. Trends Plant Sci. 14, 521–529 (2009).
Ntoukakis, V. et al. PLoS Pathog. 9, e1003123 (2013).
Kim, S. H., Qi, D., Ashfield, T., Helm, M. & Innes, R. W. Science 351, 684–687 (2016).
Lewis, J. D., Wu, R., Guttman, D. S. & Desveaux, D. PLoS Genet. 6, e1000894 (2010).
Lee, A. H.-Y. et al. PLoS Pathog. 8, e1002523 (2012).
Lewis, J. D. et al. Proc. Natl Acad. Sci. USA 110, 18722–18727 (2013).
Khan, M., Subramaniam, R. & Desveaux, D. Curr. Opin. Microbiol. 29, 49–55 (2016).
Wang, G. et al. Cell Host Microbe 18, 285–295 (2015).
Lo, T., Koulena, N., Seto, D., Guttman, D. S. & Desveaux, D. Mol. Plant Pathol. (2016).
Singer, A. U. et al. Structure 12, 1669–1681 (2004).
Wang, Y. et al. Plant Cell 22, 2033–2044 (2010).
Hinsch, M. & Staskawicz, B. Mol. Plant Microbe Interact. 9, 55–61 (1996).
Gassmann, W. Mol. Plant Microbe Interact. 18, 1054–1060 (2005).
Menna, A., Nguyen, D., Guttman, D. S. & Desveaux, D. Front. Plant Sci. 6, 995 (2015).
Huard-Chauveau, C. et al. PLoS Genet. 9, e1003766 (2013).
Acknowledgements
We thank members of the Desveaux and Guttman laboratories for valuable input on the project. This work was supported by Ontario Graduate Scholarships and the Natural Sciences and Engineering Research Council of Canada (NSERC) postgraduate awards (D.S. and T.L.), NSERC Discovery Grants (D.D. and D.S.G.), a Canada Research Chair in Plant-Microbe Systems Biology (D.D.) or Comparative Genomics (D.S.G.) and the Centre for the Analysis of Genome Evolution and Function (D.D. and D.S.G.). T-DNA lines were obtained from the ABRC.
Author information
Authors and Affiliations
Contributions
D.S., N.K., T.L., A.M. and D.D. designed experiments. D.S., N.K., T.L. and A.M. conducted experiments. D.S., N.K., T.L., A.M., D.S.G. and D.D. analysed data. D.S. and D.D. wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Information
Supplementary Methods, Supplementary References, Supplementary Figure Legends 1–4 and Supplementary Figures 1–4. (PDF 10830 kb)
Rights and permissions
About this article
Cite this article
Seto, D., Koulena, N., Lo, T. et al. Expanded type III effector recognition by the ZAR1 NLR protein using ZED1-related kinases. Nature Plants 3, 17027 (2017). https://doi.org/10.1038/nplants.2017.27
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/nplants.2017.27
This article is cited by
-
R gene-mediated resistance in the management of plant diseases
Journal of Plant Biochemistry and Biotechnology (2024)
-
The Arabidopsis effector-triggered immunity landscape is conserved in oilseed crops
Scientific Reports (2022)
-
Tackling multiple bacterial diseases of Solanaceae with a handful of immune receptors
Horticulture, Environment, and Biotechnology (2022)
-
A Maize (Zea mays L.) BIK1-Like Receptor-Like Cytoplasmic Kinase Contributes to Disease Resistance
Plant Molecular Biology Reporter (2022)
-
Regulation of plant responses to biotic and abiotic stress by receptor-like cytoplasmic kinases
Stress Biology (2022)