Abstract
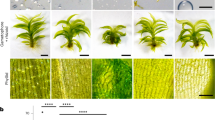
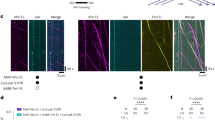
The molecular motors kinesin and dynein drive bidirectional motility along microtubules (MTs) in most eukaryotic cells. Land plants, however, are a notable exception, because they contain a large number of kinesins but lack cytoplasmic dynein, the foremost processive retrograde transporter. It remains unclear how plants achieve retrograde cargo transport without dynein. Here, we have analysed the motility of the six members of minus-end-directed kinesin-14 motors in the moss Physcomitrella patens and found that none are processive as native dimers. However, when artificially clustered into as little as dimer of dimers, the type-VI kinesin-14 (a homologue of Arabidopsis KCBP (kinesin-like calmodulin binding protein)) exhibited highly processive and fast motility (up to 0.6 μm s−1). Multiple kin14-VI dimers attached to liposomes also induced transport of this membrane cargo over several microns. Consistent with these results, in vivo observations of green fluorescent protein-tagged kin14-VI in moss cells revealed fluorescent punctae that moved processively towards the minus-ends of the cytoplasmic MTs. These data suggest that clustering of a kinesin-14 motor serves as a dynein-independent mechanism for retrograde transport in plants.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Shimmen, T. & Yokota, E. Cytoplasmic streaming in plants. Curr. Opin. Cell Biol. 16, 68–72 (2004).
Cai, G. & Cresti, M. Are kinesins required for organelle trafficking in plant cells? Front Plant. Sci. 3, 170 (2012).
Nakaoka, Y., Kimura, A., Tani, T. & Goshima, G. Cytoplasmic nucleation and atypical branching nucleation generate endoplasmic microtubules in Physcomitrella patens. Plant Cell 27, 228–242 (2015).
Miki, T., Nishina, M. & Goshima, G. RNAi screening identifies the armadillo repeat-containing kinesins responsible for microtubule-dependent nuclear positioning in Physcomitrella patens. Plant Cell Physiol. 56, 737–749 (2015).
Hirokawa, N., Noda, Y., Tanaka, Y. & Niwa, S. Kinesin superfamily motor proteins and intracellular transport. Nature Rev. Mol. Cell Biol. 10, 682–696 (2009).
Hancock, W. O. Bidirectional cargo transport: moving beyond tug of war. Nature Rev. Mol. Cell Biol. 15, 615–628 (2014).
Lawrence, C. J., Morris, N. R., Meagher, R. B. & Dawe, R. K. Dyneins have run their course in plant lineage. Traffic 2, 362–363 (2001).
Vale, R. D. The molecular motor toolbox for intracellular transport. Cell 112, 467–480 (2003).
Endow, S. A. Determinants of molecular motor directionality. Nature Cell Biol. 1, E163–E167 (1999).
Mieck, C. et al. Non-catalytic motor domains enable processive movement and functional diversification of the kinesin-14 Kar3. eLife 4, e04489 (2015).
Hepperla, A. J. et al. Minus-end-directed kinesin-14 motors align antiparallel microtubules to control metaphase spindle length. Dev. Cell 31, 61–72 (2014).
Case, R. B., Pierce, D. W., Hom-Booher, N., Hart, C. L. & Vale, R. D. The directional preference of kinesin motors is specified by an element outside of the motor catalytic domain. Cell 90, 959–966 (1997).
Hatsumi, M. & Endow, S. A. Mutants of the microtubule motor protein, nonclaret disjunctional, affect spindle structure and chromosome movement in meiosis and mitosis. J. Cell Sci. 101, 547–559 (1992).
Goshima, G. & Vale, R. D. Cell cycle-dependent dynamics and regulation of mitotic kinesins in Drosophila S2 cells. Mol. Biol. Cell 16, 3896–3907 (2005).
Shen, Z., Collatos, A. R., Bibeau, J. P., Furt, F. & Vidali, L. Phylogenetic analysis of the kinesin superfamily from Physcomitrella. Front. Plant Sci. 3, 230 (2012).
Reddy, A. S. & Day, I. S. Kinesins in the Arabidopsis genome: a comparative analysis among eukaryotes. BMC Genomics 2, 2 (2001).
Oppenheimer, D. G. et al. Essential role of a kinesin-like protein in Arabidopsis trichome morphogenesis. Proc. Natl Acad. Sci. USA 94, 6261–6266 (1997).
Suetsugu, N. et al. The KAC family of kinesin-like proteins is essential for the association of chloroplasts with the plasma membrane in land plants. Plant Cell Physiol. 53, 1854–1865 (2012).
Suetsugu, N. et al. Two kinesin-like proteins mediate actin-based chloroplast movement in Arabidopsis thaliana. Proc. Natl Acad. Sci. USA 107, 8860–8865 (2010).
Song, H., Golovkin, M., Reddy, A. S. & Endow, S. A. In vitro motility of AtKCBP, a calmodulin-binding kinesin protein of Arabidopsis. Proc. Natl Acad. Sci. USA 94, 322–327 (1997).
Pechatnikova, E. & Taylor, E. W. Kinetic mechanism of monomeric non-claret disjunctional protein (Ncd) ATPase. J. Biol. Chem. 272, 30735–30740 (1997).
Endres, N. F., Yoshioka, C., Milligan, R. A. & Vale, R. D. A lever-arm rotation drives motility of the minus-end-directed kinesin Ncd. Nature 439, 875–878 (2006).
Verhey, K. J. & Hammond, J. W. Traffic control: regulation of kinesin motors. Nature Rev. Mol. Cell Biol. 10, 765–777 (2009).
Harbury, P. B., Zhang, T., Kim, P. S. & Alber, T. A switch between two-, three-, and four-stranded coiled coils in GCN4 leucine zipper mutants. Science 262, 1401–1407 (1993).
Ulbrich, M. H. & Isacoff, E. Y. Subunit counting in membrane-bound proteins. Nature Methods 4, 319–321 (2007).
Hines, K. E. Inferring subunit stoichiometry from single molecule photobleaching. J. Gen. Physiol. 141, 737–746 (2013).
Vale, R. D. et al. Direct observation of single kinesin molecules moving along microtubules. Nature 380, 451–453 (1996).
McKenney, R. J., Huynh, W., Tanenbaum, M. E., Bhabha, G. & Vale, R. D. Activation of cytoplasmic dynein motility by dynactin-cargo adapter complexes. Science 345, 337–341 (2014).
Cove, D. The moss Physcomitrella patens. Annu. Rev. Genet. 39, 339–358 (2005).
Choi, Y. K., Liu, P., Sze, S. K., Dai, C. & Qi, R. Z. CDK5RAP2 stimulates microtubule nucleation by the gamma-tubulin ring complex. J. Cell Biol. 191, 1089–1095 (2010).
Kollman, J. M., Polka, J. K., Zelter, A., Davis, T. N. & Agard, D. A. Microtubule nucleating gamma-TuSC assembles structures with 13-fold microtubule-like symmetry. Nature 466, 879–882 (2010).
Vos, J. W., Safadi, F., Reddy, A. S. & Hepler, P. K. The kinesin-like calmodulin binding protein is differentially involved in cell division. Plant Cell 12, 979–990 (2000).
Lazzaro, M. D., Marom, E. Y. & Reddy, A. S. Polarized cell growth, organelle motility, and cytoskeletal organization in conifer pollen tube tips are regulated by KCBP, the calmodulin-binding kinesin. Planta 238, 587–597 (2013).
Furuta, K. et al. Measuring collective transport by defined numbers of processive and nonprocessive kinesin motors. Proc. Natl Acad. Sci. USA 110, 501–506 (2013).
Tomishige, M., Klopfenstein, D. R. & Vale, R. D. Conversion of Unc104/KIF1A kinesin into a processive motor after dimerization. Science 297, 2263–2267 (2002).
Klopfenstein, D. R., Tomishige, M., Stuurman, N. & Vale, R. D. Role of phosphatidylinositol(4,5)bisphosphate organization in membrane transport by the Unc104 kinesin motor. Cell 109, 347–358 (2002).
Sivaramakrishnan, S. & Spudich, J. A. Coupled myosin VI motors facilitate unidirectional movement on an F-actin network. J. Cell Biol. 187, 53–60 (2009).
Aitken, C. E., Marshall, R. A. & Puglisi, J. D. An oxygen scavenging system for improvement of dye stability in single-molecule fluorescence experiments. Biophys. J. 94, 1826–1835 (2008).
Edelstein, A., Amodaj, N., Hoover, K., Vale, R. & Stuurman, N. Computer control of microscopes using μManager. Curr. Protoc. Mol. Biol. 92, 14.20.1–14.20.17 (2010).
Bieling, P. et al. Reconstitution of a microtubule plus-end tracking system in vitro. Nature 450, 1100–1105 (2007).
Nakaoka, Y. et al. An inducible RNA interference system in Physcomitrella patens reveals a dominant role of augmin in phragmoplast microtubule generation. Plant Cell 24, 1478–1493 (2012).
Miki, T., Naito, H., Nishina, M. & Goshima, G. Endogenous localizome identifies 43 mitotic kinesins in a plant cell. Proc. Natl Acad. Sci. USA 111, E1053–E1061 (2014).
Yamada, M., Miki, T. & Goshima, G. Imaging mitosis in the moss Physcomitrella patens. Methods Mol. Biol. (in the press).
Acknowledgements
We are grateful to S. Ross and L. Chang (Nikon USA) for providing microscopes at MBL. W. Huynh provided invaluable advice in establishing the liposome assay, and A. Jain assisted with the photobleaching experiment. We also thank M. Bezanilla (University of Massachusetts Amherst) and Tomoko Nishiyama (Nagoya University) for help with microscopy and helpful discussion, Y. Nakaoka for moss lines, and M. Nishina and R. Inaba for technical assistance. This work was supported by the Human Frontier Science Program, the James A. and Faith Miller Memorial Fund (MBL), the Laura and Arthur Colwin Endowed Summer Research Fellowship Fund (MBL), the TORAY Science Foundation, Grants-in-Aid for Scientific Research (15K14540, MEXT) (G.G.), and the NIH (38499) (R.D.V.).
Author information
Authors and Affiliations
Contributions
E.J., M.Y., R.D.V. and G.G. designed the research; E.J., M.Y. and G.G. performed experiments; E.J., M.Y., R.D.V. and G.G. analysed data; E.J., R.D.V. and G.G. wrote the paper.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Rights and permissions
About this article
Cite this article
Jonsson, E., Yamada, M., Vale, R. et al. Clustering of a kinesin-14 motor enables processive retrograde microtubule-based transport in plants. Nature Plants 1, 15087 (2015). https://doi.org/10.1038/nplants.2015.87
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/nplants.2015.87
This article is cited by
-
Armadillo repeat-containing kinesin represents the versatile plus-end-directed transporter in Physcomitrella
Nature Plants (2023)
-
Modeling specific aneuploidies: from karyotype manipulations to biological insights
Chromosome Research (2023)
-
A legume kinesin controls vacuole morphogenesis for rhizobia endosymbiosis
Nature Plants (2022)
-
Self-repair protects microtubules from destruction by molecular motors
Nature Materials (2021)
-
Molecular chaperone Hsp90 protects KCBP from degradation by proteasome in Dunaliella salina cells
Folia Microbiologica (2021)