Abstract
Fatty acid amide hydrolase (FAAH) is a promising therapeutic target for the treatment of pain and CNS disorders. However, the development of potent and safe FAAH inhibitors is hindered by their off-target mediated side effect that leads to brain cell death. Its physiological off-targets and their associations with phenotypes may not be characterized using existing experimental and computational techniques as these methods fail to have sufficient proteome coverage and/or ignore native biological assemblies (BAs; i.e., protein quaternary structures). To understand the mechanisms of the side effects from FAAH inhibitors and other drugs, we develop a novel structural phenomics approach to identifying the physiological off-targets binding profile in the cellular context and on a structural proteome scale, and investigate the roles of these off-targets in impacting human physiology and pathology using text mining-based phenomics analysis. Using this integrative approach, we discover that FAAH inhibitors may bind to the dimerization interface of NMDA receptor (NMDAR) and several other BAs, and thus disrupt their cellular functions. Specifically, the malfunction of the NMDAR is associated with a wide spectrum of brain disorders that are directly related to the observed side effects of FAAH inhibitors. This finding is consistent with the existing literature, and provides testable hypotheses for investigating the molecular origin of the side effects of FAAH inhibitors. Thus, the in silico method proposed here, which can for the first time predict proteome-wide drug interactions with cellular BAs and link BA–ligand interaction with clinical outcomes, can be valuable in off-target screening. The development and application of such methods will accelerate the development of more safe and effective therapeutics.
Similar content being viewed by others
Introduction
Fatty acid amide hydrolase (FAAH) is an enzyme involved in the hydrolysis of bioactive lipids such as anandamide, 2-arachidonoylglycerol (2-AG) and oleamide.1 It is distributed in several of the major organs of the human body, but is also regionally distributed in the brain where it is believed to correlate with cannibinoid receptors.2 In particular, it is believed that the overexpression of FAAH reduces the production of known endogenous cannabinoids, specifically, anandamide (AEA) and 2-arachidonoylglycerol (2-AG).3 This makes FAAH an attractive drug target, as inhibition of FAAH would, through the upregulation of AEA and 2-AG, elicit the effects of cannabinoid activation. Thus, FAAH inhibitors may serve as analgesic, anti-inflammatory, anxiolytic and antidepressant therapeutics.4
However, the development of potent and safe FAAH inhibitors is hindered by their possible serious side effects.5 In a recent clinical trial, the FAAH inhibitor BIA 10-2474, caused cerebral hemorrhage and necrosis, leading to the death of a patient.6 It has been suggested that the deadly side effect of this FAAH inhibitor may come from its binding to unidentified off-targets.5,6 In vitro off-target screening is available against a panel of hundreds of proteins including enzymes, receptors, transporters, ion channels and second messages. However, the target space of existing assays is too small to cover the whole human proteome, where many uncharacterized proteins may be responsible for the side effect or the therapeutic effect.7 A large number of computational methods have been developed to predict off-target interactions,8–10 and associate genes with diseases.11 Most of these methods, however, only screen protein monomers or single genes. They are not the functional form of proteins in the cell. Instead, the protein cellular function is dependent on the conformational state of biological assemblies (BAs), i.e., protein quaternary structures. A drug may not only interact with a protein monomer, but interfere with its oligomerization state, leading to the disruption of its normal function. Furthermore, the drug may not bind to the endogenous ligand-binding site in the BA. It hinders the development of reliable experimental and computational methods for the proteome-wide BA screening and disease association. Consequently, few methods exist to predict cellular off-target effects resulting from the drug binding to BAs, and their associations with diseases. Owing to these limitations, it is still unclear what the off-targets of FAAH inhibitors are, and how the off-target affects the system level response.
To address the aforementioned challenges, for the first time, we develop a structural phenomics approach, which integrates heterogeneous data from structural genomics, chemical genomics and the biomedical literature, to reveal the cellular and physiological mechanism of drug–target interactions. First, we screen potential cellular off-targets of FAAH inhibitors on a structural proteome using BAs that represent the functional form of proteins in the cell. Few computational methods that can screen a compound against the structural proteome-wide BAs, including uncharacterized binding sites, are available. To our knowledge, the method in this paper is the first one used for this purpose. Then we use chemical genomics analysis, protein–ligand docking, surface electrostatic potential characterization and molecular dynamics simulation to verify the putative off-targets. Finally, we link the effect of drug off-target binding to its clinical outcome using a deep learning technique to mine the biomedical literature. By applying this integrative approach, we discover that FAAH inhibitors may bind to the dimerization interface of NMDA receptor (NMDAR) and several other BAs, thus disrupting their physiological functions. Our prediction is consistent with the existing experimental evidence. The malfunction of the NMDAR is associated with brain disorders that are directly related to the observed side effects of FAAH inhibitors. This finding suggests that the drug off-target effect is more complicated than our current understanding—which mainly focuses on the study of ligand interactions with protein monomers but not their functionally relevant BAs. Due to the limitation of existing in vitro off-target screening, an in silico method that can predict drug interactions with BAs and link protein–ligand interaction with clinical outcomes, as exampled by the proposed structural phenomics approach in this paper, will be a valuable tool to facilitate drug discovery. Specifically, it may shed new light on the development of safe and effective FAAH inhibitors.
Results
Overview of structural phenomics method
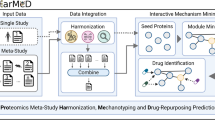
The proposed structural phenomics method aims to screen drug off-targets in the cell. To this end, it is necessary to use protein BAs that represent the cellular functional state of proteins for the study of protein–ligand interactions. Furthermore, it is needed to correlate in vivo molecular interactions with drug response phenotypes. The method used in this study mainly consists of the following steps, as shown in Figure 1.
-
1
Given a drug D and its primary target T (in this case, FAAH), all BAs in the Protein Data Bank (PDB;12 both monomer and oligomer), are compared with the ligand-binding site of T using software SMAP.13–15 If a BA has statistically significantly similar binding site to that of T, it may be the putative off-target of D. In addition, the electrostatic potentials of the binding sites of primary target and off-target are compared to further verify the putative binding promiscuity.
-
2
Known inhibitors of T and their decoys are extracted from ChEMBL,16 and docked into T, and its putative off-targets, respectively, using protein–ligand docking software Autodock Vina.17 The distributions of docking scores of inhibitors and decoys are analyzed to verify the prediction from step 1. Then, molecular dynamics simulation is performed to characterize the conformational dynamics of the putative off-target–ligand interactions.
-
3
The concepts of genes/proteins and diseases in from PubMed abstracts (up to July 2014) are mapped to a vector space using a deep learning technique, Word2Vec.18 Semantic relations between the putative off-target and diseases are determined using the cosine similarity between vectors.
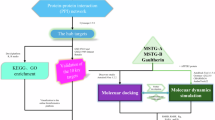
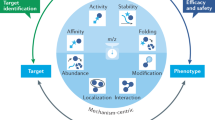
Overview of structural phenomics methodology. (a) The principle of methodology is to build a consistent model of drug–phenotype association, drug–biological assembly (quaternary structure) interaction and biological assembly–phenotype association. (b) An automated pipeline to build the model, which combines tools derived from bioinformatics, systems biology, protein–ligand docking, MD (molecular dynamics) simulation and text mining, and integrates data from structural genomics, chemical genomics and literature.
Structural proteome-wide ligand-binding site similarity of protein BAs
An increasing body of evidence supports the concept that protein fold and binding site spaces are continuous.14,19,20 Thus, ligand-binding cross-reactivity is better characterized by analyzing ligand-binding sites rather than protein sequences or global structures, as shown in many studies.7,21–23 We have developed and proven the success of the ligand-binding site comparison software SMAP.13–15 SMAP has several unique features. First, we represent protein surfaces as a graph, and characterize its geometric and topological properties using Geometric Potential that is robust to conformational variations of protein structure.15 Second, we align two surfaces using a sequence order independent profile–profile alignment (SOIPPA) algorithm.14 The sequence order independent profile–profile alignment can detect the most similar binding surface patch, without requiring predefinition of location/boundary of the binding sites. Thus, we can apply SMAP to uncharacterized binding sites from apo structures, predicted structures and BAs.24,25 Finally, we measure the similarity between aligned binding sites using a scoring function that combines evolutionary, geometric and physico-chemical information. Thus, SMAP is less sensitive to local conformational variances and inaccuracies in predicted structure. Moreover, we have developed an extreme value distribution statistical model that can rapidly estimate the statistical significance of the match between two ligand sites.13 SMAP has been successfully applied to the arena of drug repurposing,26–29 side effects prediction,30,31 polypharmacology24,32 and precision medicine.33
An FAAH structure that is co-crystallized with a drug-like inhibitor PF-750 (PDB id: 2VYA) is compared with 40,491 non-redundant BAs in the PDB using SMAP.13–15 After removing BAs that do not have human homologous proteins, five BA structures have SMAP P value less than 1.0e−3, as shown in Table 1. Most of them are related to neurological functions. The alignment score distribution of all BAs is shown in Supplementary Figure S1. In this study, we will focus on the top-ranked NMDAR. As PDB 4PE5 is not from humans, but rats, the homology model of the human NMDAR is built using I-TASSER,34 mainly to include the missing loops. The sequence alignment generated from ClustalW35 is shown in Supplementary Figures S2 and S3. The sequence identity of the GluN1A and GluN2B subunits between humans and rats is 99.3 and 98.5%, respectively. In addition, all residues in the potential binding pocket of dimerization interface are conserved.
Figure 2 shows the SMAP sequence order independent alignment of the FAAH ligand-binding site with the putative binding pocket of NMDAR, which is located in the dimerization interface of the GluN1A and GluN2B subunits of the NMDAR. The sequence order independent alignment shows the majority of aligned amino acid residues to be conserved or similar, although the sequence order is not conserved. As a similar binding site may bind a similar ligand,21 the NMDAR is a potential off-target of FAAH inhibitors. A full list of amino acid residues involved in the dimerization is shown in Supplementary Table S1.
To verify the similarity between the FAAH ligand-binding site and the putative binding pocket of NMDAR, we calculate and compare the surface electrostatic potentials of both binding pockets using Delphi.36 As shown in Figure 3, in both the pockets, the ligand lies above a relatively negatively charged surface with the more hydrophobic surface on the other side. This result further supports that the dimerization interface of NMDAR is a putative off-target binding pocket of FAAH inhibitors.
Statistical analysis of protein–ligand docking score ensembles and molecular dynamics simulation of BIA 10-2474 bound NMDAR
To verify that NMDAR is the off-target of FAAH inhibitors, we extract active FAAH inhibitors (IC50<10 μ mol/l) and inactive compounds (decoys) with IC50>1,000 μ mol/l from ChEMBL,16 and dock them into the binding pockets of FAAH and the NMDAR, respectively. In each group, the decoy ligand data set acts as a control against the experimental data set of the FAAH inhibitors. The binding score represents the strength of the interaction of the ligand to the binding pocket, in kcal/mol. Therefore, a more negative score indicates a more favorable and stronger interaction.
It is well known that the docking score is not reliable enough to predict whether a molecule is a true or false binder. The correlation analysis of docking scores of a set of molecules between the primary target and the off-target may provide more reliable information on the binding cross-reactivity.30 Although the absolute docking scores themselves are not accurate, they will be relatively similar to each other in similar binding sites. Thus, the docking scores will be linearly correlated if two binding sites are similar. As shown in Figure 4, the docking scores of 624 FAAH inhibitors in the endogenous binding pocket of primary target FAAH (PDB: 2VYA) are indeed linearly correlated with those in the predicted dimer interface of NMDAR. R2 of Pearson’s correlation coefficient is 0.75232. Around 10 molecules within the red circle of Figure 4 are outliers. If these molecules are not included, the R2 of Pearson’s correlation coefficient is 0.83895.
Furthermore, the docking score distribution of active compounds of FAAH is significantly different from that of decoys when docked to the binding site of FAAH (Supplementary Figure S4). An unpaired, two-tailed t-test gave a P value less than 1.0e−5. However, the docking score of BIA 10-2474 is −7.7 when docked to its primary target FAAH. At this score, there is no significant distinction between true and false binders. The score distribution of FAAH inhibitors and their decoys has the same trend when docked to the dimer interface of NMDAR (Supplementary Figure S5). Although the docking score of BIA 10-2474 (−7.6) is not significantly low, the score distribution of FAAH inhibitors is significantly lower than that of the decoys, with a P value less than 1.0e−5. The docking score statistics analysis suggests that the dimer interface of NMDAR has similar binding properties to the binding site of FAAH, thus is possible to bind FAAH inhibitors to this site, including BIA 10-2474.
A predicted binding pose for BIA 10-2474 in the dimerization interface of the GluN1A/GluN2B NMDAR is shown in Figure 5. The ligand is located in a pocket that is mainly formed of three transmembrane helixes, one from the monomer GluN1A (β1), two from the monomer GluN2B (β2 and β3). A total 60 ns molecular dynamics simulation suggests that the complex structure is stable (Supplementary Figure S6). Compared with the unbound structure (gray colored in Figure 5), two helices β2 and β3 had significant shifts. Especially, β3 is positioned in the inner core that acts as an ion channel. The change of conformation of the inner core may impact the normal function of NMDAR as the ion channel.
The endogenous binding site of NMDAR is in its extracellular ligand-binding domain. Docking score of BIA 10-2474 on the endogenous binding site is −5.2, far higher than the predicted binding score on the dimerization interface. Furthermore, the docking score distribution of FAAH inhibitors on this site is statistically significantly higher than that of NMDAR ligands on this site (Supplementary Figure S7). Thus, it is less likely that the FAAH inhibitor binds to the endogenous site.
Phenomics analysis of NMDAR function
Finally, we investigate which phenotypes are associated with the putative off-targets using a deep learning technique Word2Vec. Word2Vec uses a two-layer neural network to reconstruct the semantic context of gene and disease terms from a collection of 22,345,439 PubMed abstracts. In this work, it maps each term in the PubMed abstracts to a 300-dimensional vector, which represents that term’s relation to other terms. It is noted that two terms can be semantically related even if they do not appear in the same document. We extracted 90,645 gene terms and 122,771 terms related to disease phenotypes. Then gene–disease association is evaluated by the statistical significance of cosine similarity between the gene term and disease term.
As shown in Figure 6, the probability distribution of cosine similarity between random paired terms can be fitted into a generalized extreme value distribution model with μ=0,0731, σ=0.0453 and ξ=−0.0370, as supported by Bayesian Information Criterion of −30,749.92. The Q–Q plot of fitting is shown in Supplementary Figure S8. Subsequently, we can calculate the e-value of the observed similarity score by multiplying the P value derived from the extreme value distribution model with the number of term clusters (see the 'Materials and Methods' section for details).
We used ‘ndmars’, the frequently appeared term of the NMDAR in the literature, as a query. Table 2 shows top-ranked phenotypes associated with the query gene term. Interestingly, most of these terms are associated with brain disorders that cause brain cell death, which are observed as side effects of BIA 10-2427. The full information of Table 2 is shown in Supplementary Table S2 and References.
The phenotype associations with NMDAR automatically identified from word2vec are supported by the manual search of the literature. As shown in Table 2, the existing experimental evidence suggests that the alternation of NMDAR causes brain disorders in the majority of cases. Liu et al.37 have shown that the composition of the NMDAR is of critical importance to its action as a promoter of either neuronal cell survival or neuronal cell death. Specifically, the NR2B subunit is the one responsible for excitotoxic neuronal death in mature cortical cultures. In addition, NMDARs can cause neuronal cell death through the indirect production of NO (nitric oxide).38 Consistent with these observations, the brain cell death was observed as the side effect of BIA 10-2427. It is noted that the brain disorder leads to the dysregulation of the NMDAR in three cases in Table 2. One of the drawbacks of word2vec is that it cannot determine cause–effect relations but only correlations.
Discussion
As FAAH is localized in the endoplasmic reticulum, the FAAH inhibitor needs to permeate the cell membrane. It is possible for FAAH inhibitors to bind the intracellular region of NMDAR. The predicted associations between the malfunction of the NMDAR and brain disorders are supported by additional evidences from the literatures. The dimer interface is of particular interest as the residues participating in the putative binding site of NMDA (Supplementary Table S1). The dimer interface has been shown by Mcllhinney et al.39 to be critical to the cell surface expression of the NMDA subunits NR1 and NR2A; NR2A is not transported to the cell surface unless already associated with NR1. Furthermore Meddows et al.40 has shown that there is a progressive loss of NR2A surface expression as the N terminus of NR1a is successively shortened ‘…with complete loss when truncated beyond residue 380’. Paoletti and Neyton41 hypothesize that the dimer interface is involved in the allosteric regulation pathway between the Agonist Binding Domains (ABD) of NMDA and its pore domain. The allosteric interaction modulates channel opening (activation). Sun et al.42 provide evidence for the importance of the dimer interface in a different ionotropic glutamate receptor: the AMPA receptor, which is a structural homolog of the NMDAR. They suggest that the stability of the dimer interface controls the coupling of agonist binding and channel gating. Furthermore, the rearrangement of the dimer interface desensitizes the AMPA receptor.42
One of the observed side effects of BIA 10-2474 is hemorrhage. The major cause of brain hemorrhage is high blood pressure. NMDARs have been shown to be associated with the regulation of blood pressure. Glass et al.43 have shown that NMDARs contribute to the slow-pressor response in mice that are injected with Angiotensin II. Furthermore, Glass et al. have shown that the slow-pressor response requires GluN1 gene expression in PVN neurons and that ‘...the subcellular distribution of GluN1 is modified in dendritic profiles of PVN neurons following increases in blood pressure induced by Ang II’. This suggests that ‘..NMDAR plasticity in PVN neurons has a role in the elevated blood pressure mediated by Ang II’. Pyatin et al.44 have suggested that NMDARs in A5 neurons, mediate blood pressure shifts during hypoxia. They show that the blood pressure drop in mice from induced hypoxia is significantly accentuated when NMDARs in A5 neurons are blocked as compared with mice whose receptors are not blocked. Ryu et al.45 have shown that blockage of NMDARs using the selective inhibitor, MK-801, resulted in the blockage of the induced responses in rats, from Endothelin-1, a potent vasoconstrictor that increases the arterial blood pressure. This suggests that the effects of Endothelin-1 are mediated by NMDARs. Li et al.46 have shown that elevated NMDAR activity can increase lead to hypertension. They show that protein kinases and phosphatases are involved in regulating NMDAR activity in hypertensive rates, and that imbalance of NMDAR phosphorylation can ‘...augment the excitability of hypothalamic presympathetic neurons and sympathetic nerve discharges in hypertension’. Their results indicate that reducing NMDAR phosphorylation levels may be effective in treating neurogenic hypertension. Finally, Gören et al.47 have suggested that NMDARs located in the paraventricular nucleus of the hypothalamus in rats may participate in the regulation of cardiovascular homeostasis, specifically, arterial blood pressure and heart rate. They show that the blockade of NMDARs in the paraventricular nucleus, through a competitive NMDAR antagonist, results in the blunting of increases in arterial blood pressure induced by intracerebroventricular injections of NMDA.
In summary, the results from our structural phenomics prediction are consistent with existing experimental and clinical results, indicating that the dimer interface of the NMDAR may be critical to its function. Because the residues of our putative binding sites are those that primarily participate in the dimer interface of NMDA, we hypothesize that the binding of the FAAH inhibitors (in particular BIA 10-2427) may alter the conformation of NMDAR, leading to potentially deadly side effects. In addition to NMDAR, several other BAs are also potential off-targets of FAAH inhibitors. Further computational and experimental validations of this prediction may provide new insights into the design of safe and potent FAAH inhibitors.
Materials and methods
Structural proteome-wide off-target search
A FAAH structure that is co-crystallized with a drug-like inhibitor PF-750 (PDB id: 2VYA) was compared with 40,491 non-redundant BAs in PDB48 using SMAP v2.1.13–15 Default parameters were used in the computation. For the top-ranked hits with P value <1.0e−3, proteins that were not human homologous were removed. The remaining proteins were selected for further analysis. In this work, only the highest ranked protein, GluN1A/GluN2B (PDB id: 4PE5), was studied.
Homology modeling of the GluN1A/GluN2B NMDAR
The PDB structure 4PE5 was not of a human protein, but a rat protein; in addition, there were missing loops in the structure. The human sequence of GluN1A and GluN2B was separately sent to the protein structure prediction server, I-TASSER34 to build a homology model with reconstructed loops. The resulting two models were combined as a single complex structure by aligning with corresponding chains in 4PE5 using FATCAT structural alignment server with the rigid alignment option.49
Docking of FAAH inhibitors, decoys and BIA 10-2474
Next, a list of 663 known FAAH inhibitors was compiled from ChEMBL16 for docking with the main target, FAAH (PDB id: 2VYA) as well as the off-target GlnN1A/Gln2B NMDAR homology model. A total 624 FAAH inhibitors were iteratively docked to each target. Autodock Vina,17 through the PyRx GUI interface, was used to perform the docking simulations. The centers for the binding box were approximated by averaging the positions of the conjugated co-crystal ligand’s (PF7) atoms in the main target; PF7 was also conjugated to the off-target (as part of the prediction of the protein structure of the off-target, detailed above) and a similar averaging was carried out to determine its binding center. The PDB files of the protein targets were also cleaned; DS Visualizer was used to remove the co-crystallized ligand and the surrounding water molecules for proper docking with Autodock Vina. The FAAH inhibitor in question, BIA 10-2474, was then docked to both targets and the binding scores determined.
As a control, 97 decoy ligands that were extracted from ChEMBL were also docked to each target. Once the binding scores of inhibitors and decoys for both targets were obtained, a paired, two-tailed t-test was done to determine whether the difference in the scores were statistically significant.
Molecular dynamics simulation
The membrane of POPC was first constructed with 334 POPC molecules. The membrane was 120 Å in the x direction and 100 Å in the y direction. The transmembrane region (TMD) of NMDAR complex is positioned in the membrane manually using VMD with pseudo-four-fold symmetrical axis (z axis) at TMD perpendicular to the membrane. The system was solvated in explicit water molecules with the margin of solvent from the solute molecule 12 Å, which led to an initial simulation box of 148×125×155 Å. The ion concentration (ionic strength 0.2 mol/l), 150 Na+ and 136 Cl− were added. ACEMD software50 was used for all molecular dynamics simulations with the CGenFF36 force field for small molecules, CHARMM27 force field for protein, CHARMM36 lipid force field for membrane molecules and TIP3P for water molecules. Langevin thermostat and Berendsen pressure were used for temperature control at 300 K and pressure maintenance of 1 atmosphere. Periodic boundary conditions were applied in the simulation.
Calculation of surface electrostatic potential
DelPhi (v5.1) software36 was used to calculate electrostatic potentials. The AMBER partial atomic charges and atomic radii were used, and the internal dielectric constant value was set as 4. External dielectric constant was set as 2 and 80 for transmembrane and solvent-exposed domain, respectively. The solvent and ionic probe radius was 1.4 and 2.0 Å, respectively. The electrostatic potential surface was shown by Pymol.51 The negatively and positively charged surfaces are shown in red and blue, respectively.
Text mining of gene–phenotype association
Data preprocessing
A total 22,345,439 PubMed abstracts dated before July 2014 were downloaded from the National Library of Medicine. BioLemmatizer v1.2 was used to transform a word to a lemma.52 Part-of-speech tagger was carried out using RDRPostagger v1.13.53 Disease, gene and chemical name entities were recognized using DNorm v0.0.6;54 Gimli v1.0.2;55 and tmChem,56 respectively. A total 90,645 gene terms and 122,771 terms related to disease phenotypes were extracted. In this study, chemical terms were not involved. Phrases were recognized from words except those recognized as disease, gene and chemical terms, using the Word2Vec utility tool.18
Vector representation of terms
Word2Vec18 was used to map each processed term in the PubMed abstracts to a 300-dimensional vector, which represent that term’s relation to other terms. Instead of individual sentences, whole abstracts were used as the training sample. The rationale was that the terms in the different sentences could be related if they were in the same abstract. A Skip-gram model with a widow size of 10 was used for the training.
Calculation of term–term relations
Gene–disease associations were then evaluated by the cosine similarity between the vector representation of gene term and that of disease term. One of features of the Word2Vec representation is that it could capture analog relations between terms. For example, if gene G1 is similar to gene G2, and disease D1 is similar to disease D2 and G1 is associated with D1, then G2 may be associated with D2.
Markov Cluster Algorithm MCL57 was used to cluster diseases based on disease–disease cosine similarity. A total 9,787 disease clusters were formed when the MCL-I option was set to 4.0.
The cosine similarity score of randomly selected gene–disease pairs were fitted into a generalized extreme value distribution model. The P value was calculated for each top-ranked association using the fitted extreme value distribution model when querying a gene term against the disease terms. The P value was converted to an e-value by multiplying the number of disease clusters.
Conclusion
In this paper, we introduce a structural phenomics method to identify the cellular off-target of drugs and to link them to human physiology and pathology. We apply this method to identify cellular off-targets of FAAH inhibitors that are promising therapeutics for the treatment of pain and CNS disorders, but may cause deadly side effects. We propose that the FAAH inhibitor may bind to the dimerization interface of the NMDAR, and thus disrupt its physiological function. Moreover, the elucidated cellular off-target effects from the NMDAR are strongly correlated to observed phenotypes. This study demonstrates that a structural phenomics approach, which systematically explores cellular protein–ligand interactions on a genome-wide scale and then links them to phenotypes, is complementary to existing methods for the prediction of drug–target interactions that, not only has limited proteome coverage, but also focuses on protein monomers that may not represent the functional form of proteins in the cell. It is expected that the structural phenomics approach will provide us with valuable clues as to the cellular basis of drug actions. At the same time, it will help to transform the conventional one-drug-one-target drug discovery process to a new paradigm of polypharmacology.
References
Ueda, N., Puffenbarger, R. A., Yamamoto, S. & Deutsch, D. G. The fatty acid amide hydrolase (FAAH). Chem. Phys. Lipids 108, 107–121 (2000).
Seierstad, M. & Breitenbucher, J. G. Discovery and development of fatty acid amide hydrolase (FAAH) inhibitors. J. Med. Chem. 51, 7327–7343 (2008).
Devane, W. et al. Isolation and structure of a brain constituent that binds to the cannabinoid receptor. Science 258, 1946–1949 (1992).
Ahn, K., Johnson, D. S. & Cravatt, B. F. Fatty acid amide hydrolase as a potential therapeutic target for the treatment of pain and CNS disorders. Expert Opin. Drug Discov. 4, 763–784 (2009).
Butler, D. & Callaway, E. Scientists in the dark after French clinical trial proves fatal. Nature 529, 263–264 (2016).
Everts, S. in C&E News, Vol. 94 (2016).
Xie, L., Kinnings, S. L. & Bourne, P. E. Novel computational approaches to polypharmacology as a means to define responses to individual drugs. Annu. Rev. Pharmacol. Toxicol. 52, 361–379 (2012).
Woo, J. H. et al. Elucidating compound mechanism of action by network perturbation analysis. Cell 162, 441–451 (2015).
Cichonska, A., Rousu, J. & Aittokallio, T. Identification of drug candidates and repurposing opportunities through compound-target interaction networks. Expert Opin. Drug Discov. 10, 1333–1345 (2015).
Hart, T. & Xie, L. Providing data science support for systems pharmacology and its implications to drug discovery. Expert Opin. Drug Discov. 11, 241–256 (2016).
Gill, N., Singh, S. & Aseri, T. C. Computational disease gene prioritization: an appraisal. J. Comput. Biol. 21, 456–465 (2014).
Deshpande, N. et al. The RCSB Protein Data Bank: a redesigned query system and relational database based on the mmCIF schema. Nucleic Acids Res. 33, D233–D237 (2005).
Xie, L. & Bourne, P. E. A unified statistical model to support local sequence order independent similarity searching for ligand-binding sites and its application to genome-based drug discovery. Bioinformatics 25, i305–i312 (2009).
Xie, L. & Bourne, P. E. Detecting evolutionary relationships across existing fold space, using sequence order-independent profile-profile alignments. Proc. Natl Acad. Sci. USA 105, 5441–5446 (2008).
Xie, L. & Bourne, P. E. A robust and efficient algorithm for the shape description of protein structures and its application in predicting ligand binding sites. BMC Bioinformatics 8 (Suppl 4): S9 (2007).
Gaulton, A. et al. ChEMBL: a large-scale bioactivity database for drug discovery. Nucleic Acids Res. 40, D1100–D1107 (2012).
Trott, O. & Olson, A. J. AutoDock Vina: improving the speed and accuracy of docking with a new scoring function, efficient optimization and multithreading. J. Comput. Chem. 31, 455–461 (2010).
Mikolov, T., Sutskever, I., Chen, K., Corrado, G. S. & Dean, J. in NIPS, 3111–3119 (2013).
Petrey, D., Fischer, M. & Honig, B. Structural relationships among proteins with different global topologies and their implications for function annotation strategies. Proc. Natl Acad. Sci. USA 106, 17377–17382 (2009).
Kolodny, R., Petrey, D. & Honig, B. Protein structure comparison: implications for the nature of ‘fold space’, and structure and function prediction. Curr. Opin. Struct. Biol. 16, 393–398 (2006).
Xie, L., Xie, L. & Bourne, P. E. Structure-based systems biology for analyzing off-target binding. Curr. Opin. Struct. Biol. 21, 189–199 (2011).
Xie, L. et al. Towards structural systems pharmacology to study complex diseases and personalized medicine. PLoS Comput. Biol. 10, e1003554 (2014).
Hart, T. & Xie, L. Providing data science support for systems pharmacology and its implications to drug discovery. Expert Opin. Drug Discov. 11, 241–256 (2016).
Kinnings, S. L., Xie, L., Fung, K., Xie, L. & Bourne, P. E. The Mycobacterium tuberculosis Drugome and its polypharmacological implications. PLoS Comput. Biol. 6, e100976 (2010).
Chang, R. L., Xie, L., Bourne, P. E. & Palsson, B. O. Antibacterial mechanisms identified through structural systems pharmacology. BMC Syst. Biol. 7, 102 (2013).
Xie, L., Evangelidis, T., Xie, L. & Bourne, P. E. Drug discovery using chemical systems biology: weak inhibition of multiple kinases may contribute to the anti-cancer effect of Nelfinavir. PLoS Comput. Biol. 7, e1002037 (2011).
Kinnings, S. L. et al. Drug discovery using chemical systems biology: repositioning the safe medicine Comtan to treat multi-drug and extensively drug resistant tuberculosis. PLoS Comput. Biol. 5, e1000423 (2009).
Kinnings, S. L. et al. A machine learning-based method to improve docking scoring functions and its application to drug repurposing. J. Chem. Inf. Model. 51, 408–419 (2011).
Han, W. & Xie, L. Bioinformatics and Biomedicine Workshops (BIBMW), 2012 in IEEE International Conference on Bioinformatics and Biomedicine ( BIBM , 2012) 28–31 (IEEE, 2012).
Xie, L., Wang, J. & Bourne, P. E. In silico elucidation of the molecular mechanism defining the adverse effect of selective estrogen receptor modulators. PLoS Comput. Biol. 3, e217 (2007).
Xie, L., Li, J., Xie, L. & Bourne, P. E. Drug discovery using chemical systems biology: identification of the protein-ligand binding network to explain the side effects of CETP inhibitors. PLoS Comput. Biol. 5, e1000387 (2009).
Durrant, J. D. et al. A multidimensional strategy to detect polypharmacological targets in the absence of structural and sequence homology. PLoS Comput. Biol. 6, e1000648 (2010).
Hart, T. et al. Toward repurposing metformin as a precision anti-cancer therapy using structural systems pharmacology. Sci. Rep. 6, 20441 (2016).
Zhang, Y. I-TASSER server for protein 3D structure prediction. BMC Bioinformatics 9, 40 (2008).
Thompson, J. D., Higgins, D. G. & Gibson, T. J. CLUSTAL W: improving the sensitivity of progressive multiple sequence alignment through sequence weighting, position-specific gap penalties and weight matrix choice. Nucleic Acids Res. 22, 4673–4680 (1994).
Honig, B. & Nicholls, A. Classical electrostatics in biology and chemistry. Science 268, 1144 (1995).
Liu, Y. et al. NMDA receptor subunits have differential roles in mediating excitotoxic neuronal death both in vitro and in vivo. J. Neurosci. 27, 2846–2857 (2007).
Madajka M. H. Nitroxidative Stress Induced Neurodegeneration in Intracerebral Hemorrhagic Stroke—A Nanomedical Approach (Ohio University, 2007).
McIlhinney, R. A. J. et al. Assembly intracellular targeting and cell surface expression of the human N-methyl-d-aspartate receptor subunits NR1a and NR2A in transfected cells. Neuropharmacology 37, 1355–1367 (1998).
Meddows, E. et al. Identification of molecular determinants that are important in the assembly of N-methyl-D-aspartate receptors. J. Biol. Chem. 276, 18795–18803 (2001).
Paoletti, P. & Neyton, J. NMDA receptor subunits: function and pharmacology. Curr. Opin. Pharmacol. 7, 39–47 (2007).
Sun, Y. et al. Mechanism of glutamate receptor desensitization. Nature 417, 245–253 (2002).
Glass, M. J. et al. NMDA receptor plasticity in the hypothalamic paraventricular nucleus contributes to the elevated blood pressure produced by angiotensin II. J. Neurosci. 35, 9558–9567 (2015).
Pyatin, V. & Tatarnikov, V. NMDA receptors of A5 area in the regulation of blood pressure and respiratory activity during hypoxia in rats. Bull. Exp. Biol. Med. 159, 420–423 (2015).
Ryu, J. S. et al. NMDA receptor and NO mediate ET-1-induced behavioral and cardiovascular effects in periaqueductal gray matter of rats. Arch. Pharm. Res. 24, 64–68 (2001).
Li, D. P., Zhou, J. J. & Pan, H. L. Endogenous casein kinase‐1 modulates NMDA receptor activity of hypothalamic presympathetic neurons and sympathetic outflow in hypertension. J. Physiol. 593, 4439–4452 (2015).
Gören, M. Z., Onat, F. & Berkman, K. Participation of NMDA and kainate receptors of paraventricular nucleus in cardiovascular responses to glutamate receptor agonist. Eur. J. Pharmacol. 388, 77–84 (2000).
Berman, H. M. et al. The Protein Data Bank. Nucleic Acids Res. 28, 235–242 (2000).
Ye Y. & Godzik, A. Flexible structure alignment by chaining aligned fragment pairs allowing twists. Bioinformatics 9, ii246–ii255 (2003).
Harvey, M. J., Giupponi, G. & De Fabritiis, G. ACEMD: Accelerated biomolecular dynamics simulations in the microsecond timescale. J. Chem. Theory Comput. 5, 1632–1639 (2009).
PyMOL Molecular Graphics System, version 1.8. (Schrödinger, 2015).
Liu, H., Christiansen, T., Baumgartner, W. A. Jr. & Verspoor, K. BioLemmatizer: a lemmatization tool for morphological processing of biomedical text. J. Biomed. Semantics 3, 3 (2012).
Nguyen, D. Q., Dai Quoc Nguyen, D. D. P. & Pham, S. B. RDRPOSTagger: A Ripple Down Rules-based Part-Of-Speech Tagger in Proceedings of the Demonstrations at the 14th Conference of the European Chapter of the Association for Computational Linguistics (EACL) (2014).
Leaman, R., Islamaj Dogan, R. & Lu, Z. DNorm: disease name normalization with pairwise learning to rank. Bioinformatics 29, 2909–2917 (2013).
Campos, D., Matos, S. & Oliveira, J. L. Gimli: open source and high-performance biomedical name recognition. BMC Bioinformatics 14, 54 (2013).
Leaman, R., Wei, C. H. & Lu, Z. tmChem: a high performance approach for chemical named entity recognition and normalization. J. Cheminform. 7, S3 (2015).
Enright, A. J., Van Dongen, S. & Ouzounis, C. A. An efficient algorithm for large-scale detection of protein families. Nucleic Acids Res. 30, 1575–1584 (2002).
Acknowledgements
We thank reviewers’ constructive suggestions in revising the manuscript. We thank Dr Zhiyong Lu for providing software DNorm and tmChem. This research was supported by the National Library of Medicine of the National Institute of Health under the award number R01LM011986; and National Science Foundation under the award number CNS-0958379, CNS-0855217, ACI-1126113; and the City University of New York High Performance Computing Center at the College of Staten Island.
Author information
Authors and Affiliations
Contributions
LX conceived the project; LX designed the experiments; SD, JJ, ZZ and LX performed the experiments and analyzed the results; SD, ZZ and LX wrote the manuscript; All the authors reviewed the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies the paper on the npj Systems Biology and Applications website
Rights and permissions
This work is licensed under a Creative Commons Attribution-NonCommercial-ShareAlike 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-sa/4.0/
About this article
Cite this article
Dider, S., Ji, J., Zhao, Z. et al. Molecular mechanisms involved in the side effects of fatty acid amide hydrolase inhibitors: a structural phenomics approach to proteome-wide cellular off-target deconvolution and disease association. npj Syst Biol Appl 2, 16023 (2016). https://doi.org/10.1038/npjsba.2016.23
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1038/npjsba.2016.23