Abstract
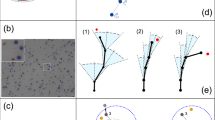
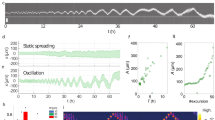
Cell migration paths are generally described as random walks, associated with both intrinsic and extrinsic noise. However, complex cell locomotion is not merely related to such fluctuations, but is often determined by the underlying machinery. Cell motility is driven mechanically by actin and myosin, two molecular components that generate contractile forces. Other cell functions make use of the same components and, therefore, will compete with the migratory apparatus. Here, we propose a physical model of such a competitive system, namely dendritic cells whose antigen capture function and migratory ability are coupled by myosin II. The model predicts that this coupling gives rise to a dynamic instability, whereby cells switch from persistent migration to unidirectional self-oscillation, through a Hopf bifurcation. Cells can then switch to periodic polarity reversals through a homoclinic bifurcation. These predicted dynamic regimes are characterized by robust features that we identify through in vitro trajectories of dendritic cells over long timescales and distances. We expect that competition for limited resources in other migrating cell types can lead to similar deterministic migration modes.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Selmeczi, D. et al. Cell motility as random motion: a review. Eur. Phys. J. Spec. Top. 157, 1–15 (2008).
Harris, T. H. et al. Generalized lévy walks and the role of chemokines in migration of effector cd8+ t cells. Nature 486, 545–548 (2012).
Bénichou, O., Loverdo, C., Moreau, M. & Voituriez, R. Intermittent search strategies. Rev. Mod. Phys. 83, 81–129 (2011).
Blanch-Mercader, C. & Casademunt, J. Spontaneous motility of actin lamellar fragments. Phys. Rev. Lett. 110, 078102 (2013).
Mitchison, T. & Cramer, L. Actin-based cell motility and cell locomotion. Cell 84, 371–379 (1996).
Verkhovsky, A. B., Svitkina, T. M. & Borisy, G. G. Self-polarization and directional motility of cytoplasm. Curr. Biol. 9, 11–20 (1999).
Nelson, W. J. Adaptation of core mechanisms to generate cell polarity. Nature 422, 766–774 (2003).
Kruse, K., Joanny, J., Jülicher, F. & Prost, J. Contractility and retrograde flow in lamellipodium motion. Phys. Biol. 3, 130–137 (2006).
Yam, P. T. et al. Actin-myosin network reorganization breaks symmetry at the cell rear to spontaneously initiate polarized cell motility. J. Cell Biol. 178, 1207–1221 (2007).
Kozlov, M. M. & Mogilner, A. Model of polarization and bistability of cell fragments. Biophys. J. 93, 3811–3819 (2007).
Bendix, P. M. et al. A quantitative analysis of contractility in active cytoskeletal protein networks. Biophys. J. 94, 3126–3136 (2008).
Rubinstein, B., Fournier, M. F., Jacobson, K., Verkhovsky, A. B. & Mogilner, A. Actin-myosin viscoelastic flow in the keratocyte lamellipod. Biophys. J. 97, 1853–1863 (2009).
Wilson, C. A. et al. Myosin II contributes to cell-scale actin network treadmilling through network disassembly. Nature 465, 373–377 (2010).
Fournier, M. F., Sauser, R., Ambrosi, D., Meister, J.-J. & Verkhovsky, A. B. Force transmission in migrating cells. J. Cell Biol. 188, 287–297 (2010).
Wolgemuth, C. W., Stajic, J. & Mogilner, A. Redundant mechanisms for stable cell locomotion revealed by minimal models. Biophys. J. 101, 545–553 (2011).
Hawkins, R. J. et al. Spontaneous contractility-mediated cortical flow generates cell migration in three-dimensional environments. Biophys. J. 101, 1041–1045 (2011).
Asnacios, A. & Hamant, O. The mechanics behind cell polarity. Trends Cell Biol. 22, 584–591 (2012).
Callan-Jones, A. & Voituriez, R. Active gel model of amoeboid cell motility. New J. Phys. 15, 025022 (2013).
Danuser, G., Allard, J. & Mogilner, A. Mathematical modeling of eukaryotic cell migration: insights beyond experiments. Annu. Rev. Cell Dev. Biol. 29, 501–528 (2013).
Recho, P., Putelat, T. & Truskinovsky, L. Contraction-driven cell motility. Phys. Rev. Lett. 111, 108102 (2013).
Liu, Y.-J. et al. Confinement and low adhesion induce fast amoeboid migration of slow mesenchymal cells. Cell 160, 659–672 (2015).
Maiuri, P. et al. Actin flows mediate a universal coupling between cell speed and cell persistence. Cell 161, 374–386 (2015).
Chabaud, M. et al. Cell migration and antigen capture are antagonistic processes coupled by myosin II in dendritic cells. Nature Commun. 6, 7526 (2015).
Vargas, P. et al. Innate control of actin nucleation determines two distinct migration behaviours in dendritic cells. Nature Cell Biol. 18, 43–53 (2016).
Austyn, J. Lymphoid dendritic cells. Immunology 62, 161–170 (1987).
Banchereau, J. & Steinman, R. M. Dendritic cells and the control of immunity. Nature 392, 245–252 (1998).
Banchereau, J. et al. Immunobiology of dendritic cells. Annu. Rev. Immunol. 18, 767–811 (2000).
Heuzé, M. L. et al. Migration of dendritic cells: physical principles, molecular mechanisms, and functional implications. Immunol. Rev. 256, 240–254 (2013).
Steinman, R. M. & Swanson, J. The endocytic activity of dendritic cells. J. Exp. Med. 182, 283–288 (1995).
Swanson, J. A. & Watts, C. Macropinocytosis. Trends Cell Biol. 5, 424–428 (1995).
Sallusto, F., Cella, M., Danieli, C. & Lanzavecchia, A. Dendritic cells use macropinocytosis and the mannose receptor to concentrate macromolecules in the major histocompatibility complex class II compartment: downregulation by cytokines and bacterial products. J. Exp. Med. 182, 389–400 (1995).
Bergert, M. et al. Force transmission during adhesion-independent migration. Nature Cell Biol. 17, 524–529 (2015).
Guckenheimer, J. & Holmes, P. Nonlinear Oscillations, Dynamical Systems, and Bifurcations of Vector Fields Vol. 42 (Springer Science and Business Media, 1983).
Solanes, P. et al. Space exploration by dendritic cells requires maintenance of myosin II activity by ip3 receptor 1. EMBO J. 34, 798–810 (2015).
Zhang, J., Guo, W.-H. & Wang, Y.-L. Microtubules stabilize cell polarity by localizing rear signals. Proc. Natl Acad. Sci. USA 111, 16383–16388 (2014).
Szabo, B. et al. Auto-reverse nuclear migration in bipolar mammalian cells on micropatterned surfaces. Cell Motil. Cytoskeleton 59, 38–49 (2004).
Fraley, S. I., Feng, Y., Giri, A., Longmore, G. D. & Wirtz, D. Dimensional and temporal controls of three-dimensional cell migration by zyxin and binding partners. Nature Commun. 3, 719 (2012).
Camley, B. A., Zhao, Y., Li, B., Levine, H. & Rappel, W.-J. Periodic migration in a physical model of cells on micropatterns. Phys. Rev. Lett. 111, 158102 (2013).
Veltman, D. M. Drink or drive: competition between macropinocytosis and cell migration. Biochem. Soc. Trans. 43, 129–132 (2015).
Prentice-Mott, H. V. et al. Directional memory arises from long-lived cytoskeletal asymmetries in polarized chemotactic cells. Proc. Natl Acad. Sci. USA 113, 1267–1272 (2016).
Acknowledgements
The authors thank M. Chabaud and M. Heuzé who performed the experimental study23 and have kindly contributed the data; R. Attia, M. Bretou, P. Vargas and P. Maiuri for discussions; and A. Monge and E. Bellin for language editing. N.S.G. would like to thank the ISF grant 580/12 for support. This work is made possible through the historic generosity of the Perlman family.
Author information
Authors and Affiliations
Contributions
I.L. created the analytic model with input by all authors, designed and implemented the analysis of both the model and the experiments, and prepared the manuscript; M.P. and A.-M.L.-D. conceived and supervised the experiments and contributed the data; N.S.G. jointly conceived the theoretical study with R.V., supervised the work and edited the manuscript. All authors reviewed the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary information
Supplementary information (PDF 14394 kb)
Supplementary Movie 1
Supplementary Movie (MP4 270 kb)
Supplementary Movie 2
Supplementary Movie (MP4 51 kb)
Supplementary Movie 3
Supplementary Movie (MP4 101 kb)
Supplementary Movie 4
Originally mislabelled as Movie 3 (MP4 65 kb)
Supplementary Movie 5
Originally mislabelled as Movie 4 (MP4 112 kb)
Supplementary Movie 6
Originally mislabelled as Movie 5 (MP4 141 kb)
Supplementary Movie 7
Originally mislabelled as Movie 6 (MP4 162 kb)
Rights and permissions
About this article
Cite this article
Lavi, I., Piel, M., Lennon-Duménil, AM. et al. Deterministic patterns in cell motility. Nature Phys 12, 1146–1152 (2016). https://doi.org/10.1038/nphys3836
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nphys3836
This article is cited by
-
Polarity and mixed-mode oscillations may underlie different patterns of cellular migration
Scientific Reports (2023)
-
Cell migration guided by long-lived spatial memory
Nature Communications (2021)
-
Contact enhancement of locomotion in spreading cell colonies
Nature Physics (2017)
-
How to eat on the go
Nature Physics (2016)