Abstract
The ubiquity of scale-free topology in nature raises the question of whether this particular network design confers an evolutionary advantage1. A series of studies has identified key principles controlling the growth and the dynamics of scale-free networks2,3,4. Here, we use neuron-based networks of boolean components as a framework for modelling a large class of dynamical behaviours in both natural and artificial systems5,6,7. Applying a training algorithm, we characterize how networks with distinct topologies evolve towards a pre-established target function through a process of random mutations and selection8,9,10. We find that homogeneous random networks and scale-free networks exhibit drastically different evolutionary paths. Whereas homogeneous random networks accumulate neutral mutations and evolve by sparse punctuated steps11,12, scale-free networks evolve rapidly and continuously. Remarkably, this latter property is robust to variations of the degree exponent. In contrast, homogeneous random networks require a specific tuning of their connectivity to optimize their ability to evolve. These results highlight an organizing principle that governs the evolution of complex networks and that can improve the design of engineered systems.
Similar content being viewed by others
Main
In recent years, studies of the architecture of large complex networks have unveiled a topology, known as scale-free, in which the connectivity between elements is power-law distributed3. In biology, the intricate interactions of genes and proteins can be viewed as neuron-based networks that control biological signals13. It is also common to model large technological and social systems using similar neuronal networks. For example, electronic devices that carry out computational tasks are built using large networks of interconnected logic gates, whereas collective social behaviours emerge from a complex structure of social relations and the dynamics of personal influences14,15. In such real-world networks, the topology is important because it mediates the effect of modifications in local interaction that can sometimes affect the networks dynamical behaviour. Topology could thus be a determining factor to modify or evolve the function of networks16. In this picture, new dynamical behaviours emerge from ‘tinkering’ the local interactions from old systems. In living organisms, genetic and protein networks have evolved through a process of mutations and selection to carry out specific functions under specific environmental conditions. This process has inspired physical scientists to apply a similar evolutionary approach to explore solutions to difficult optimization problems and to search for novel designs of artificial systems17. For example, evolutionary algorithms have been used to train software and even reconfigurable electronic chips to carry out pre-determined tasks18. Also known as ‘evolvable hardware’, these electronic devices are composed of large numbers of neuron-like elements whose interactions are programmable19. To exploit the power of this evolutionary approach, we must identify the key organizing principles that govern the relationship between topology and the networks ability to evolve. As it is vital for any biological system to adapt and rapidly carry out new functions, long training time is a fundamental limitation of the evolutionary approach in engineered systems20. Therefore, understanding how specific architectures impose evolutionary constraints on complex networks is key to the design of artificial systems.
Here we use boolean threshold dynamics as a general framework to model the dynamics of large complex networks. This framework has previously been successful in modelling central biological dynamical processes such as the cell cycle in yeast6 and the expression of the polarity genes in Drosophila segments5 (see Supplementary Information, Fig. S2A). In engineering, boolean neural networks have been used in a wide variety of applications, such as electronic circuits (see Supplementary Information, Fig. S2B) and pattern recognition. We use boolean threshold networks as a theoretical test-bed to investigate how networks with distinct topologies evolve to carry out a pre-determined target function. To drive networks’ evolution, we use a standard evolutionary algorithm consisting of successive cycles of random mutations and selection. Similar algorithms have been applied to boolean networks to study the emergence of homeostasis and noise imprinting9, evolutionary plasticity of biological systems16, modularity21, and more recently the emergence of motifs8 in engineered systems. In this study, we wish to compare the evolutionary paths of networks with either scale-free or homogeneous random topologies. We use directed graphs in which nodes represent the network elements and the arrows between nodes represent interactions between elements. In random homogeneous networks, the number of connections per node, k, is Poisson distributed: Prand(k)=e−〈K〉〈K〉k/k!. The mean connectivity 〈K〉 is the relevant topological parameter to characterize the random network architecture. Conversely, the topology of scale-free networks is heterogeneous and the number of connections per node is power-law distributed  , where the associated topological parameter is the degree exponent γ. Scale-free networks here have both in- and out-degree distributions that are power-law distributed; we generate networks with a given degree distribution like in ref. 18 (Supplementary Information). To model the interactions between individual nodes we use a set of rules that determine the dynamical behaviour of the network. We approximate the activity of each element by a simple two-state model 22 (σi=1 or σi=0). The sum of all interactions, activating or inhibitory, determines the state of each node7,23 (Fig. 1a). The state of a whole network is defined at each time point by σ(t)={σ1,σ2,…,σN} and goes through a transient regime until it falls within a cycle.
, where the associated topological parameter is the degree exponent γ. Scale-free networks here have both in- and out-degree distributions that are power-law distributed; we generate networks with a given degree distribution like in ref. 18 (Supplementary Information). To model the interactions between individual nodes we use a set of rules that determine the dynamical behaviour of the network. We approximate the activity of each element by a simple two-state model 22 (σi=1 or σi=0). The sum of all interactions, activating or inhibitory, determines the state of each node7,23 (Fig. 1a). The state of a whole network is defined at each time point by σ(t)={σ1,σ2,…,σN} and goes through a transient regime until it falls within a cycle.
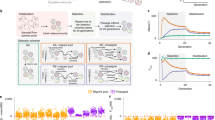
a, Boolean threshold node. Each connection (arrow) between nodes has a weight wi j indicating the relative strength of the interaction. For each node i with Ki inputs there are Ki non-zero weights. If a weight is positive, the interaction is activating; if it is negative, the interaction is inhibitory. The weights wi j are distributed uniformly in the interval (−1,1). A node is active or inactive (value 1 or 0) depending on the sign of the sum of its incoming interactions. We also implement specific rules for all Ki=1 nodes (Supplementary Information). b, Network dynamics and target function. One node from the network is the output (here σ6). The fitness is calculated by comparing the time series from the output node with the target function. c, Mutation and selection processes. Networks are mutated at fixed rate μ. Mutations include both changing an incoming connection of a node (arrows in bold) and changing the weight of a connection (numbers in bold). The networks with the highest fitness are then selected to form the population of the next generation. Each iteration of this process is one generation step.
We use a simple evolutionary algorithm to drive the evolution of a population of networks towards a pre-established target function9 (see the Methods section). A population of networks evolves through a sequence of random mutations and selection, such that the distribution of incoming connections remains fixed (Fig. 1c). The networks are then selected according to the distance of their output to the target, which defines the fitness function (Fig. 1b). Under these conditions, the fixed network topology has to be determined prior to this process. For example, a specific topology could emerge as the outcome of a higher-level selective or growth process. Finally, we want to produce robust functions, that is, cycles independent of the choice of initial conditions. Hence, we randomly choose initial values for each network at each generation, so that the final evolved networks are robust against variations of initial conditions. Robustness is a key property of biological and engineered networks because they need to maintain important functions under variations of environmental conditions24,25.
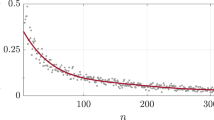
In Fig. 2 we show typical evolutionary runs for random (Fig. 2a) and scale-free networks (Fig. 2b). The parameters (〈K〉=1.9; γ=2.5) are chosen such that networks of both categories have the same average number of connections 〈K〉 and the same number of nodes N. It is also possible to compare networks with similar dynamical behaviour (see the Supplementary Information) but for the sake of simplicity we chose to identify networks by their average connectivity. The fitness of 50 statistically independent runs is shown in Fig. 3 for the same values of parameters. The evolutionary path of a population of homogeneous random networks and scale-free networks is initially rapid, but after just a few generations, the fitness behaviour differs greatly. In the population of homogeneous random networks the evolutionary path exhibits long plateaus where the fitness remains steady over hundreds of generations26. Plateaus are followed by punctuated jumps where the functions in the population come closer to the target function. This evolutionary behaviour resembles that of punctuated equilibrium described in ref. 12. We interpret the presence of the plateaus as the signature of flat directions in the fitness landscape between sparsely distributed local optima. During the plateaus the network population is dominated by a single function: a short cycle (usually with length L=1 or 2) that maximizes the fitness (Fig. 2a). Although there is no measurable improvement of the fitness, neutral mutations modify connections and weights in the connectivity matrix of the networks within the population but do not affect the function. When an additional advantageous mutation occurs, the number of copies of the associated mutant grows exponentially with the number of generations and becomes dominant in the population (Fig. 2a).
a,b, Average distance to the target is defined as 1-Fitness (Npop=50, N=500, Lc=10 and μ=0.02) for homogeneous networks (a) and scale-free networks (b). For random networks over 800 generations there is only one function in the whole population with 1-Fitness=0.3 for which the value of the output node has period L=2 (generation 500). After the advantageous jump, the population includes two functions (either 1-Fitness=0.3, L=2 or 1-Fitness=0.1, L=10) that the networks follow depending on the initial conditions (generation 1,200). In scale-free networks, after only a few generations (generation 10) the population consists of ∼25 different functions of various periods (L=1, 2, 5, 20, 40, 50). The perfect function, which matches the target function, appears as soon as the 12th generation, but does not dominate the population until ∼250 generations.
a,b, Distribution of the fitness of 50 populations as a function of the number of generations for homogeneous random networks (a) and for scale-free networks (b). Results for 〈K 〉=4.1 and γ=2.0 are presented in Supplementary Information, Fig. S8.
In contrast, a population of scale-free networks evolves continuously towards the target function (Fig. 2b). After only few generations, the population consists of many different functions which can also have different cycle lengths (Fig. 2b). We interpret the diversity of functions in scale-free networks as the signature of the fitness landscape that allows the population to escape local optima with very few mutations. Many mutated networks with an improved function occur at each generation, and produce a population composed of different functions until an optimal solution (1-Fitness=0) becomes dominant. Surprisingly, even poor solutions can survive over many generations along with the best fit. We then determine whether for each topology, independent runs with distinct initial conditions and mutations exhibit similar evolutionary paths. We find that evolutionary paths of random networks depend on rare advantageous mutative events and thus differ from one another (Fig. 3a). For scale-free networks, the distribution of independent evolutionary runs exhibits a similar trend to that of individual runs (Fig. 3b). Functions of high fitness emerge by evolving existing functions gradually towards the target function. Hence, a population of scale-free networks not only evolves faster than that of random networks but also has the capacity to produce a wide range of heritable functions27.
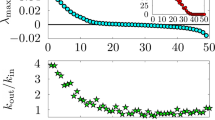
The dynamical behaviour of boolean networks is characterized by a phase transition from order to chaos. In the ordered phase, a small difference in the initial conditions fades away, whereas it spreads and dominates the dynamics in the chaotic phase28. These two phases correspond to low and high average connectivity respectively. We calculate the critical point using the annealed approximation introduced by Derrida and Pomeau28. The dynamical behaviour of boolean threshold networks depends not only on 〈K〉, like in Kauffman networks25, but on the higher moments of the distribution P(k). Scale-free and homogeneous random networks with the same average connectivity can exhibit distinct dynamical behaviour29. Random networks exhibit chaotic behaviour for 〈K〉 larger than Kc=3.83 and scale-free networks exhibit chaotic behaviour for exponents γ lower than γc=2.42 (see the Supplementary Information). We found that the ability for homogeneous random networks to evolve depends on the specific value of the parameter 〈K〉 (Fig. 4a). In particular, the fitness increases for values of 〈K〉 ranging from the critical to the chaotic phase. Conversely, the fitness of scale-free networks does not show large variations with different values of γ (Fig. 4b). It is conceivable that scale-free networks have a predetermined evolutionary advantage because the length of their cycle matches that of the target function. However, we found that the convergence of the fitness function was also better for scale-free networks even when the length of the target function, Lc, was much smaller (see the Supplementary Information). A qualitative argument allows us to understand the origin of these distinct evolutionary behaviours for the two topologies. First, we compute the average probability, 〈Pdyn〉, that a perturbation associated with a mutation of a given node would affect the dynamics of another directly connected node (see the Supplementary Information). Then, the probability that this latter mutation would affect the dynamics of the output node is 〈Pdyn〉ℓ, where ℓ is the average distance in the network. We found that the probability 〈Pdyn〉ℓ depends strongly on the connectivity distribution of the network and is larger for scale-free than for random networks. As a result, scale-free networks are more likely than random networks to produce a new function by several orders of magnitude. Scale-free topology alone allows the evolution of networks towards the target function faster than those with homogeneous random topology for any values of 〈K〉 or γ.
a,b, Populations of random (a) and scale-free (b) networks for various average connectivity 〈K 〉 and degree exponents γ. Average fitness of 50 independent evolutionary paths (Npop=50, N=500, Lc=10 and μ=0.02). For a comparison between random and scale-free networks in pairs of same average connectivity, see Supplementary Information, Fig. S6.
On the basis of the analysis of the phase diagram of boolean networks, Kauffman suggested that evolution of real systems may occur ‘at the edge of chaos’30. This hypothesis entails that systems not only need to be stable to carry out a function but they also need to be sensitive enough to perturbations to evolve towards new functions. Because the critical phase gathers the two latter properties, it was proposed that living systems may exhibit a similar critical behaviour to be evolvable. In Kauffman’s picture, the topological parameters have to be fine-tuned to specific values so that systems can exhibit a critical behaviour. In contrast, in our study, scale-free threshold networks evolve fast and continuously, and the associated evolutionary paths are robust to variations of the degree exponent γ. Consequently, the dynamical behaviour of scale-free networks does not need to be fine-tuned to the critical phase to promote evolution. On the other hand, homogeneous random networks evolve through a series of sparse punctuated changes, which inhibit their ability to evolve rapidly towards the desired function. In contrast with scale-free networks, the evolutionary paths of homogeneous networks depend on the specific values of the average connectivity 〈K〉: random networks with a low connectivity evolve slowly, whereas networks with larger connectivity evolve faster. This study indicates that topology governs the evolutionary paths of large complex networks, which suggests that the ubiquity of certain topologies in nature, such as scale-free, may also be the product of a selective process. The underlying physical principles of this approach are general and are applicable to a wide spectrum of neural-based systems that model real-world complex networks ranging from biological to engineered systems.
Methods
Network output and target function
Each round of mutations and selection is known as a generation. A series of 10,000 generations for one population is one evolutionary run. At each generation a set of initial values is attributed randomly to each node. The value of each node is updated following the dynamical rules described in Fig. 1a. After a transient phase, the whole network follows a cycle of length L. Each network has a fixed output node throughout one evolutionary run. The values of the output node during the network’s cycle form a time series that is the ‘function’ of the network. This time series is compared with a target function of length Lc using a fitness function (Fig. 1b). The time series of 0s and 1s, which constitutes the target function, is randomly chosen for each evolutionary run and is kept fixed throughout the run. At each generation we randomly choose different initial conditions for each network so that we could select for networks that are robust to variations of initial conditions.
Evolutionary algorithm
First, we randomly generate a population of Npop=50 networks that have either a homogeneous random (with a fixed parameter 〈K〉) or heterogeneous scale-free topology (with a fixed parameter γ). Second, we increase the size of the population to include both the non-mutated parent networks and the mutated offspring. Each network from the initial population will have an offspring of three mutated networks. The size of the population will then be M=(3 mutants+1 parent)Npop. Third, we select a new generation of Npop networks that have the highest fitness among the non-mutated parent networks and the mutated offspring.
Mutations
At each generation we mutate each node of each network in the population with a fixed rate μ=0.02 so that the expected number of mutations in each network is μ N, where N is the size of the network. For each mutative process we randomly choose a node, and we either change one of its input connections or its weight in the associated updating rule (Fig. 1c). For such mutations the distribution of incoming connections remains unaltered, so that a population of networks that starts as scale-free (random) remains scale-free (random) throughout an evolutionary run. On the other hand, the distribution of outgoing connections can change. For random populations the final distribution of outgoing connections is identical to the initial one, whereas for scale-free populations the distribution evolves as it would if there were only random mutations without any selection (see the Supplementary Information).
Fitness function
The fitness is a real number ranging between 0 and 1. The value of the fitness function F will be about 0.5 when the networks are randomly chosen and 1 for a network output that exactly matches the target function. To calculate the fitness for each network, we update the network according to the dynamical rules until the networks falls within a cycle. The fitness of each network is calculated using the hamming distance of the output node to the target function:

The maximum is calculated over all cyclic permutations of the target function. The sum on the right-hand side is the time average of the distance between the output node and a given target function during the cycle of the network. The sum is taken over L·Lc, where Lc is the length of the target and L is the cycle length of the whole network (to save computational time, we calculate this sum using the least common multiple of Lc and L). Under this condition, the fitness is independent of the value at which the output node begins the cycle. In our calculations all sums cannot exceed tmax=35·Lc because of computational restrictions. This cutoff is not expected to affect the results because almost all networks have cycles and transient times much shorter than tmax (see Supplementary Information). The target function is a cycle of length Lc=10.
References
Albert, R., Jeong, H. & Barabási, A. L. Error and attack tolerance of complex networks. Nature 406, 378–382 (2000).
Barrat, A., Barthélemy, M. & Vespignani, A. Weighted evolving networks: Coupling topology and weight dynamics. Phys. Rev. Lett. 92, 228701 (2004).
Albert, R. & Barabási, A. L. Statistical mechanics of complex networks. Rev. Mod. Phys. 74, 47–97 (2002).
Bar-Yam, Y. & Epstein, I. R. Response of complex networks to stimuli. Proc. Natl Acad. Sci. USA 101, 4341–4345 (2004).
Albert, R. & Othmer, H. G. The topology of the regulatory interactions predicts the expression pattern of the segment polarity genes in Drosophila melanogaster. J. Theor. Biol. 223, 1–18 (2003).
Li, F. T., Long, T., Lu, Y., Ouyang, Q. & Tang, C. The yeast cell-cycle network is robustly designed. Proc. Natl Acad. Sci. USA 101, 4781–4786 (2004).
Hopfield, J. J. Neural networks and physical systems with emergent collective computational abilities. Proc. Natl Acad. Sci. USA 79, 2554–2558 (1982).
Kashtan, N. & Alon, U. Spontaneous evolution of modularity and network motifs. Proc. Natl Acad. Sci. USA 102, 13773–13778 (2005).
Stern, M. D. Emergence of homeostasis and ‘noise imprinting’ in an evolution model. Proc. Natl Acad. Sci. USA 96, 10746–10751 (1999).
Fontana, W. & Schuster, P. Continuity in evolution: On the nature of transitions. Science 280, 1451–1455 (1998).
Bornholdt, S. & Sneppen, K. Neutral mutations and punctuated equilibrium in evolving genetic networks. Phys. Rev. Lett. 81, 236–239 (1998).
Eldredge, N. & Gould, S. J. in Models in Paleobiology (ed. Schopf, T. J. M.) 82–115 (Cooper & Co, San Francisco, 1972).
Bray, D. Protein molecules as computational elements in living cells. Nature 376, 307–312 (1995) ibid. 378, 419 (1995).
Marsden, P. V. & Friedkin, N. E. Network studies of social-influence. Sociol. Methods Res. 22, 127–151 (1993).
Granovetter, M. Threshold models of collective behavior. Am. J. Sociol. 83, 1420–1443 (1978).
Wagner, A. Does evolutionary plasticity evolve? Evolution 50, 1008–1023 (1996).
Holland, J. H. Adaptation in Natural and Artificial Systems: An Introductory Analysis with Applications to Biology, Control, and Artificial Intelligence (MIT Press, Cambridge, Massachusetts, 1992).
Catanzaro, M. & Pastor-Satorras, R. Analytic solution of a static scale-free network model. Eur. Phys. J. B 44, 241–248 (2005).
Beiu, V., Quintana, J. M. & Avedillo, M. J. VLSI implementations of threshold logic—A comprehensive survey. IEEE Trans. Neural Netw. 14, 1217–1243 (2003).
Mitchell, M. An Introduction to Genetic Algorithms (MIT Press, Cambridge, Massachusetts, 1996).
Variano, E. A., McCoy, J. H. & Lipson, H. Networks, dynamics, and modularity. Phys. Rev. Lett. 92, 188701 (2004).
Kauffman, S. A. Metabolic stability and epigenesis in randomly constructed genetic nets. J. Theor. Biol. 22, 437–467 (1969).
Kurten, K. E. Correspondence between neural threshold networks and Kauffman boolean cellular automata. J. Phys. A 21, L615–L619 (1988).
Savageau, M. A. Parameter sensitivity as a criterion for evaluating and comparing performance of biochemical systems. Nature 229, 542–544 (1971).
Aldana, M. & Cluzel, P. A natural class of robust networks. Proc. Natl Acad. Sci. USA 100, 8710–8714 (2003).
Kauffman, S. & Levin, S. Towards a general-theory of adaptive walks on rugged landscapes. J. Theor. Biol. 128, 11–45 (1987).
Kirschner, M. & Gerhart, J. Evolvability. Proc. Natl Acad. Sci. USA 95, 8420–8427 (1998).
Derrida, B. & Pomeau, Y. Random networks of automata—A simple annealed approximation. Europhys. Lett. 1, 45–49 (1986).
Rohlf, T. & Bornholdt, S. Criticality in random threshold networks: annealed approximation and beyond. Physica A 310, 245–259 (2002).
Kauffman, S. A. The Origins of Order: Self Organization and Selection in Evolution (Oxford Univ. Press, New York, 1993).
Acknowledgements
We thank L. P. Kadanoff for stimulating discussions and we are also thankful to M. Aldana, C. Guet and T. Emonet for useful comments. This work was supported by the Materials Research Science and Engineering Center program of the National Science Foundation under NSF DMR-0213745. We gratefully acknowledge use of the ‘Jazz’ cluster operated by the Mathematics and Computer Science Division at Argonne National Laboratory.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Rights and permissions
About this article
Cite this article
Oikonomou, P., Cluzel, P. Effects of topology on network evolution. Nature Phys 2, 532–536 (2006). https://doi.org/10.1038/nphys359
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nphys359
This article is cited by
-
The nature and nurture of network evolution
Nature Communications (2023)
-
Exploratory adaptation in large random networks
Nature Communications (2017)
-
Latent phenotypes pervade gene regulatory circuits
BMC Systems Biology (2014)
-
Evolutionary design of non-frustrated networks of phase-repulsive oscillators
Scientific Reports (2012)
-
Survival of the sparsest: robust gene networks are parsimonious
Molecular Systems Biology (2008)