Abstract
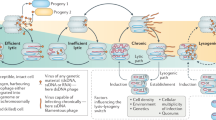
Temperate bacterial viruses (phages) may enter a symbiosis with their host cell, forming a unit called a lysogen. Infection and viral replication are disassociated in lysogens until an induction event such as DNA damage occurs, triggering viral-mediated lysis. The lysogen–lytic viral reproduction switch is central to viral ecology, with diverse ecosystem impacts. It has been argued that lysogeny is favoured in phages at low host densities. This paradigm is based on the fraction of chemically inducible cells (FCIC) lysogeny proxy determined using DNA-damaging mitomycin C inductions. Contrary to the established paradigm, a survey of 39 inductions publications found FCIC to be highly variable and pervasively insensitive to bacterial host density at global, within-environment and within-study levels. Attempts to determine the source(s) of variability highlighted the inherent complications in using the FCIC proxy in mixed communities, including dissociation between rates of lysogeny and FCIC values. Ultimately, FCIC studies do not provide robust measures of lysogeny or consistent evidence of either positive or negative host density dependence to the lytic–lysogenic switch. Other metrics are therefore needed to understand the drivers of the lytic–lysogenic decision in viral communities and to test models of the host density-dependent viral lytic–lysogenic switch.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Knowles, B. et al. Lytic to temperate switching of viral communities. Nature 531, 466–470 (2016).
Wang, Z. & Goldenfeld, N. Fixed points and limit cycles in the population dynamics of lysogenic viruses and their hosts. Phys. Rev. E 82, 011918 (2010).
Weitz, J. Quantitative Viral Ecology: Dynamics of Viruses and Their Microbial Hosts (Princeton University Press, 2015).
Jiang, S. C. & Paul, J. H. Significance of lysogeny in the marine environment: studies with isolates and a model of lysogenic phage production. Microb. Ecol. 35, 235–243 (1998).
Canchaya, C., Proux, C., Fournous, G., Bruttin, A. & Brussow, H. Prophage genomics. Microbiol. Mol. Biol. Rev. 67, 238–276 (2003).
Otsuji, N., Sekiguchi, M., Iijima, T. & Takagi, Y. Induction of phage formation in the lysogenic Escherichia coli K-12 by mitomycin C. Nature 184, 1079–1080 (1959).
Roberts, J. W. & Roberts, C. W. Proteolytic cleavage of bacteriophage lambda repressor in induction. Proc. Natl Acad. Sci. USA 72, 147–151 (1975).
Ackermann, H.-W. & DuBow, M. S. (eds) Viruses of Prokaryotes (CRC Press, 1987).
Muschel, L. H. & Schmoker, K. Activity of mitomycin C, other antibiotics, and serum against lysogenic bacteria. J. Bacteriol. 92, 967–971 (1966).
Jiang, S. C. & Paul, J. H. Seasonal and diel abundance of viruses and occurrence of lysogeny/bacteriocinogeny in the marine environment. Mar. Ecol. Prog. Ser. 104, 163–172 (1994).
Jiang, S. C. & Paul, J. H. Occurrence of lysogenic bacteria in marine microbial communities as determined by prophage induction. Mar. Ecol. Prog. Ser. 142, 27–38 (1996).
Weinbauer, M. G. & Suttle, C. A. Potential significance of lysogeny to bacteriophage production and bacterial mortality in coastal waters of the Gulf of Mexico. Appl. Environ. Microbiol. 62, 4374–4380 (1996).
Paul, J. H. & Weinbauer, M. Detection of lysogeny in marine environments. Man. Aquat. Viral Ecol. 4, 30–33 (2010).
Maurice, C. F., Bouvier, T., Comte, J., Guillemette, F. & Del Giorgio, P. A. Seasonal variations of phage life strategies and bacterial physiological states in three northern temperate lakes. Environ. Microbiol. 12, 628–641 (2010).
Evans, C. & Brussaard, C. P. D. Regional variation in lytic and lysogenic viral infection in the Southern Ocean and its contribution to biogeochemical cycling. Appl. Environ. Microbiol. 78, 6741–6748 (2012).
Payet, J. & Suttle, C. A. To kill or not to kill: the balance between lytic and lysogenic viral infection is driven by trophic status. Limnol. Oceanogr. 58, 465–474 (2013).
Brum, J. R., Hurwitz, B. L., Schofield, O., Ducklow, H. W. & Sullivan, M. B. Seasonal time bombs: dominant temperate viruses affect Southern Ocean microbial dynamics. ISME J. 10, 437–449 (2015).
Weitz, J. S. et al. A multitrophic model to quantify the effects of marine viruses on microbial food webs and ecosystem processes. ISME J. 9, 1352–1364 (2015).
Bondy-Denomy, J. et al. Prophages mediate defense against phage infection through diverse mechanisms. ISME J. 10, 2854–2866 (2016).
Dedrick, R. M. et al. Prophage-mediated defence against viral attack and viral counter-defence. Nat. Microbiol. 2, 16251 (2017).
van Houte, S., Buckling, A. & Westra, E. R. Evolutionary ecology of prokaryotic immune mechanisms. Microbiol. Mol. Biol. Rev. 80, 745–763 (2016).
Breitbart, M. Marine viruses: truth or dare. Mar. Sci. Annu. Rev. 4, 425–448 (2012).
Paul, J. H. Prophages in marine bacteria: dangerous molecular time bombs or the key to survival in the seas? ISME J. 2, 579–589 (2008).
Cochran, P. K. & Paul, J. H. Seasonal abundance of lysogenic bacteria in a subtropical estuary. Appl. Environ. Microbiol. 64, 2308–2312 (1998).
Thomas, R., Berdjeb, L., Sime-Ngando, T. & Jacquet, S. Viral abundance, production, decay rates and life strategies (lysogeny versus lysis) in Lake Bourget (France). Environ. Microbiol. 13, 616–630 (2011).
Bettarel, Y., Bouvy, M., Dumont, C. & Sime-Ngando, T. Virus–bacterium interactions in water and sediment of West African inland aquatic systems. Appl. Environ. Microbiol. 72, 5274–5282 (2006).
Stopar, D., Černe, A., Žigman, M., Poljšak-Prijatelj, M. & Turk, V. Viral abundance and a high proportion of lysogens suggest that viruses are important members of the microbial community in the Gulf of Trieste. Microb. Ecol. 47, 1–8 (2004).
Robinson, W. S. Ecological correlations and the behavior of individuals. Am. Sociol. Rev. 15, 351–357 (1950).
Palesse, S., Colombet, J., Ram, A. S. P. & Sime-Ngando, T. Linking host prokaryotic physiology to viral lifestyle dynamics in a temperate freshwater lake (Lake Pavin, France). Microb. Ecol. 68, 740–750 (2014).
Maurice, C. F. et al. Disentangling the relative influence of bacterioplankton phylogeny and metabolism on lysogeny in reservoirs and lagoons. ISME J. 5, 831–842 (2011).
McNair, K., Bailey, B. A. & Edwards, R. A. PHACTS, a computational approach to classifying the lifestyle of phages. Bioinformatics 28, 614–618 (2012).
Cochran, P. K., Kellogg, C. A. & Paul, J. H. Prophage induction of indigenous marine lysogenic bacteria by environmental pollutants. Mar. Ecol. Prog. Ser. 164, 125–133 (1998).
Wommack, K. E. & Colwell, R. R. Virioplankton: viruses in aquatic ecosystems. Microbiol. Mol. Biol. Rev. 64, 69–114 (2000).
Franklin, R. B., Garland, J. L., Bolster, C. H. & Mills, A. L. Impact of dilution on microbial community structure and functional potential: comparison of numerical simulations and batch culture experiments. Appl. Environ. Microbiol. 67, 702–712 (2001).
Philippot, L. et al. Loss in microbial diversity affects nitrogen cycling in soil. ISME J. 7, 1609–1619 (2013).
Peter, H. et al. Function-specific response to depletion of microbial diversity. ISME J. 5, 351–361 (2011).
Roger, F., Bertilsson, S., Langenheder, S., Osman, O. A. & Gamfeldt, L. Effects of multiple dimensions of bacterial diversity on functioning, stability and multifunctionality. Ecology 97, 2716–2728 (2016).
Wilhelm, S. W., Brigden, S. M. & Suttle, C. A. A dilution technique for the direct measurement of viral production: a comparison in stratified and tidally mixed coastal waters. Microb. Ecol. 43, 168–173 (2002).
Roux, S., Enault, F., Hurwitz, B. L. & Sullivan, M. B. Virsorter: mining viral signal from microbial genomic data. PeerJ 3, e985 (2015).
Arndt, D. et al. PHASTER: a better, faster version of the PHAST phage search tool. Nucleic Acids Res. 44, W16–W21 (2016).
Clasen, J. L., Brigden, S. M., Payet, J. P. & Suttle, C. A. Evidence that viral abundance across oceans and lakes is driven by different biological factors. Freshwat. Biol. 53, 1090–1100 (2008).
Wigington, C. H. et al. Re-examination of the relationship between marine virus and microbial cell abundances. Nat. Microbiol. 1, 15024 (2016).
Parikka, K. J., Le Romancer, M., Wauters, N. & Jacquet, S. Deciphering the virus-to-prokaryote ratio (VPR): insights into virus–host relationships in a variety of ecosystems. Biol. Rev. Camb. Philos. Soc. 92, 1081–1100 (2017).
Simpson, E. H. The interpretation of interaction in contingency tables. J. R. Stat. Soc. Ser. B 13, 238–241 (1951).
Breitbart, M., Wegley, L., Leeds, S., Schoenfeld, T. & Rohwer, F. Phage community dynamics in hot springs. Appl. Environ. Microbiol. 70, 1633–1640 (2004).
Colombet, J. et al. Depth-related gradients of viral activity in Lake Pavin. Appl. Environ. Microbiol. 72, 4440–4445 (2006).
Drewes, F., Peter, H. & Sommaruga, R. Are viruses important in the plankton of highly turbid glacier-fed lakes? Sci. Rep. 6, 24608 (2016).
Laybourn-Parry, J., Anesio, A. M., Madan, N. & Säwström, C. Virus dynamics in a large epishelf lake (Beaver Lake, Antarctica). Freshwat. Biol. 58, 1484–1493 (2013).
Lisle, J. T. & Priscu, J. C. The occurrence of lysogenic bacteria and microbial aggregates in the lakes of the McMurdo Dry Valleys, Antarctica. Microb. Ecol. 47, 427–439 (2004).
Lymer, D. & Lindström, E. S. Changing phosphorus concentration and subsequent prophage induction alter composition of a freshwater viral assemblage. Freshwat. Biol. 55, 1984–1996 (2010).
Pradeep Ram, A. S. & Sime-Ngando, T. Resources drive trade-off between viral lifestyles in the plankton: evidence from freshwater microbial microcosms. Environ. Microbiol. 12, 467–479 (2010).
Ram, A. S. P. et al. Variable viral and grazer control of prokaryotic growth efficiency in temperate freshwater lakes (French Massif Central). Microb. Ecol. 66, 906–916 (2013).
Ram, A. S. P., Palesse, S., Colombet, J., Thouvenot, A. & Sime-Ngando, T. The relative importance of viral lysis and nanoflagellate grazing for prokaryote mortality in temperate lakes. Freshwat. Biol. 59, 300–311 (2014).
Tapper, M. A. & Hicks, R. E. Temperate viruses and lysogeny in Lake Superior bacterioplankton. Limnol. Oceanogr. 43, 95–103 (1998).
Bettarel, Y. et al. Virioplankton distribution and activity in a tropical eutrophicated bay. Estuar. Coast. Shelf Sci. 80, 425–429 (2008).
Bettarel, Y. et al. Ecological traits of planktonic viruses and prokaryotes along a full-salinity gradient. FEMS Microbiol. Ecol. 76, 360–372 (2011).
Bongiorni, L., Magagnini, M., Armeni, M., Noble, R. & Danovaro, R. Viral production, decay rates, and life strategies along a trophic gradient in the North Adriatic Sea. Appl. Environ. Microbiol. 71, 6644–6650 (2005).
Bouvy, M. et al. Uncoupled viral and bacterial distributions in coral reef waters of Tuamotu Archipelago (French Polynesia). Mar. Pollut. Bull. 65, 506–515 (2012).
Laybourn-Parry, J., Marshall, W. A. & Madan, N. J. Viral dynamics and patterns of lysogeny in saline Antarctic lakes. Polar Biol. 30, 351–358 (2006).
Long, A., McDaniel, L. D., Mobberley, J. & Paul, J. H. Comparison of lysogeny (prophage induction) in heterotrophic bacterial and Synechococcus populations in the Gulf of Mexico and Mississippi River plume. ISME J. 2, 132–144 (2008).
Maurice, C. F., Bouvier, C., Wit, R. & Bouvier, T. Linking the lytic and lysogenic bacteriophage cycles to environmental conditions, host physiology and their variability in coastal lagoons. Environ. Microbiol. 15, 2463–2475 (2013).
Muck, S. et al. Fracture zones in the Mid Atlantic Ridge lead to alterations in prokaryotic and viral parameters in deep-water masses. Front. Microbiol. 5, 264 (2014).
Nguyen-Kim, H. et al. Coral mucus is a hot spot for viral infections. Appl. Environ. Microbiol. 81, 5773–5783 (2015).
Ortmann, A. C., Lawrence, J. E. & Suttle, C. A. Lysogeny and lytic viral production during a bloom of the cyanobacterium Synechococcus spp. Microb. Ecol. 43, 225–231 (2002).
Weinbauer, M. G. & Suttle, C. A. Lysogeny and prophage induction in coastal and offshore bacterial communities. Aquat. Microb. Ecol. 18, 217–225 (1999).
Weinbauer, M. G., Brettar, I. & Höfle, M. G. Lysogeny and virus-induced mortality of bacterioplankton in surface, deep, and anoxic marine waters. Limnol. Oceanogr. 48, 1457–1465 (2003).
Williamson, S. J., Houchin, L. A., McDaniel, L. & Paul, J. H. Seasonal variation in lysogeny as depicted by prophage induction in Tampa Bay, Florida. Appl. Environ. Microbiol. 68, 4307–4314 (2002).
Mei, M. L. & Danovaro, R. Virus production and life strategies in aquatic sediments. Limnol. Oceanogr. 49, 459–470 (2004).
Montanié, H. et al. Microbial interactions in marine water amended by eroded benthic biofilm: a case study from an intertidal mudflat. J. Sea Res. 92, 74–85 (2014).
Ghosh, D. et al. Prevalence of lysogeny among soil bacteria and presence of 16S rRNA and trzN genes in viral-community DNA. Appl. Environ. Microbiol. 74, 495–502 (2008).
Haas, A. F. et al. Unraveling the unseen players in the ocean—a field guide to water chemistry and marine microbiology. J. Vis. Exp. 21, 1–16 (2014).
Grasis, J. A. et al. Species-specific viromes in the ancestral holobiont Hydra. PLoS ONE 9, e109952 (2014).
Acknowledgements
The authors thank R. Young for microbiological insight and guidance. Canadian Institute for Advanced Research Integrated Microbial Biodiversity Program Fellowship Award 141679, National Science Foundation grants OISE-1243541 and DEB-1046413, a Gordon and Betty Moore Foundation Investigator Award GBMF-3781 (to F.R.) and National Science Foundation grants OCE-1538567 (to L.W.K.), IOS-1456301 and DEB-1555854 (to M.B.) funded this work. The authors thank G. Gueiros and K. Furby for critiquing the manuscript.
Author information
Authors and Affiliations
Contributions
B.K. and F.R. designed, conducted and wrote up the study. B.B., L.B., M.B., A.C.-G., J.d.C., R.E., B.F., J.G., A.H., P.K., L.W.K., A.L., J.N., G.P., L.P., N.R., S.S., A.S., C.S. and M.Y. contributed data, analysis and manuscript preparation.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Information
Supplementary Figure 1, Supplementary Tables 1 and 2. (PDF 217 kb)
Rights and permissions
About this article
Cite this article
Knowles, B., Bailey, B., Boling, L. et al. Variability and host density independence in inductions-based estimates of environmental lysogeny. Nat Microbiol 2, 17064 (2017). https://doi.org/10.1038/nmicrobiol.2017.64
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/nmicrobiol.2017.64
This article is cited by
-
Biogeographic patterns and drivers of soil viromes
Nature Ecology & Evolution (2024)
-
Interactions between bacterial and phage communities in natural environments
Nature Reviews Microbiology (2022)
-
Identification and characterization of a novel Enterococcus bacteriophage with potential to ameliorate murine colitis
Scientific Reports (2021)
-
Confocal microscopy reveals alterations of thylakoids in Limnospira fusiformis during prophage induction
Protoplasma (2021)
-
High cell densities favor lysogeny: induction of an H20 prophage is repressed by quorum sensing and enhances biofilm formation in Vibrio anguillarum
The ISME Journal (2020)