Abstract
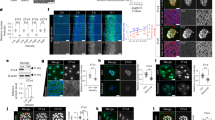
Cell size is specific to each species and impacts cell function. Various phenomenological models for cell size regulation have been proposed, but recent work in bacteria has suggested an ‘adder’ model, in which a cell increments its size by a constant amount between each division. However, the coupling between cell size, shape and constriction remains poorly understood. Here, we investigate size control and the cell cycle dependence of bacterial growth using multigenerational cell growth and shape data for single Caulobacter crescentus cells. Our analysis reveals a biphasic mode of growth: a relative timer phase before constriction where cell growth is correlated to its initial size, followed by a pure adder phase during constriction. Cell wall labelling measurements reinforce this biphasic model, in which a crossover from uniform lateral growth to localized septal growth is observed. We present a mathematical model that quantitatively explains this biphasic ‘mixer’ model for cell size control.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Iyer-Biswas, S. et al. Scaling laws governing stochastic growth and division of single bacterial cells. Proc. Natl Acad. Sci. USA 111, 15912–15917 (2014).
Wright, C. S. et al. Intergenerational continuity of cell shape dynamics in Caulobacter crescentus. Sci. Rep. 5, 9155 (2015).
Wang, P. et al. Robust growth of Escherichia coli. Curr. Biol. 20, 1099–1103 (2010).
Campos, M. et al. A constant size extension drives bacterial cell size homeostasis. Cell 159, 1433–1446 (2014).
Taheri-Araghi, S. et al. Cell-size control and homeostasis in bacteria. Curr. Biol. 25, 385–391 (2015).
Sauls, J. T., Li, D. & Jun, S. Adder and a coarse-grained approach to cell size homeostasis in bacteria. Curr. Opin. Cell Biol. 38, 38–44 (2016).
Amir, A. Cell size regulation in bacteria. Phys. Rev. Lett. 112, 208102 (2014).
Deforet, M., van Ditmarsch, D. & Xavier, J. B. Cell-size homeostasis and the incremental rule in a bacterial pathogen. Biophys. J. 109, 521–528 (2015).
Voorn, W., Koppes, L. & Grover, N. Mathematics of cell division in Escherichia coli. Curr. Top. Mol. Genet. 1, 187–194 (1993).
Jun, S. & Taheri-Araghi, S. Cell-size maintenance: universal strategy revealed. Trends Microbiol. 23, 4–6 (2015).
Schaechter, M., Williamson, J. P., Hood, J. Jr & Koch, A. L. Growth, cell and nuclear divisions in some bacteria. J. Gen. Microbiol. 29, 421–434 (1962).
Koppes, L., Meyer, M., Oonk, H., De Jong, M. & Nanninga, N. Correlation between size and age at different events in the cell division cycle of Escherichia coli. J. Bacteriol. 143, 1241–1252 (1980).
Osella, M., Nugent, E. & Lagomarsino, M. C. Concerted control of Escherichia coli cell division. Proc. Natl Acad. Sci. USA 111, 3431–3435 (2014).
Tanouchi, Y. et al. A noisy linear map underlies oscillations in cell size and gene expression in bacteria. Nature 523, 357–360 (2015).
Donachie, W., Begg, K. & Vicente, M. Cell length, cell growth and cell division. Nature 264, 328–333 (1976).
Kubitschek, H. Bilinear cell growth of Escherichia coli. J. Bacteriol. 148, 730–733 (1981).
Harris, L. K. & Theriot, J. A. Relative rates of surface and volume synthesis set bacterial cell size. Cell 165, 1479–1492 (2016).
Wang, J. D. & Levin, P. A. Metabolism, cell growth and the bacterial cell cycle. Nat. Rev. Microbiol. 7, 822–827 (2009).
Mitchison, J. M. The Biology of the Cell Cycle (CUP Archive, 1971).
Marshall, W. F. et al. What determines cell size? BMC Biol. 10, 101 (2012).
Banerjee, S., Scherer, N. F. & Dinner, A. R. Shape dynamics of growing cell walls. Soft Matter 12, 3442–3450 (2016).
Lin, Y., Li, Y., Crosson, S., Dinner, A. R. & Scherer, N. F. Phase resetting reveals network dynamics underlying a bacterial cell cycle. PLoS Comput. Biol. 8, e1002778 (2012).
Ursell, T. S. et al. Rod-like bacterial shape is maintained by feedback between cell curvature and cytoskeletal localization. Proc. Natl Acad. Sci. USA 111, E1025–E1034 (2014).
Aaron, M. et al. The tubulin homologue FtsZ contributes to cell elongation by guiding cell wall precursor synthesis in Caulobacter crescentus. Mol. Microbiol. 64, 938–952 (2007).
Kuru, E. et al. In situ probing of newly synthesized peptidoglycan in live bacteria with fluorescent d-amino acids. Angew. Chem. Int. Ed. 51, 12519–12523 (2012).
Reshes, G., Vanounou, S., Fishov, I. & Feingold, M. Cell shape dynamics in Escherichia coli. Biophys. J. 94, 251–264 (2008).
Sibarita, J.-B. in Microscopy Techniques (ed. Rietdorf, J. ) 201–243 (Springer, 2005).
Acknowledgements
The authors thank C. Wright and S. Iyer-Biswas for measurements and shape analysis of C. crescentus single-cell data1,2. The authors thank S. Crosson and A. Fiebig for contributing reagents, materials and discussions. The authors acknowledge funding from the National Science Foundation Physics of Living Systems (NSF PHY-1305542), the National Science Foundation Materials Research Science and Engineering Center (MRSEC) at the University of Chicago (NSF DMR-1420709), the W. M. Keck Foundation and the Graduate Program in Biophysical Sciences at the University of Chicago (T32 EB009412/EB/NIBIB NIH HHS/United States). S.B. acknowledges support from the University College London for completion of part of this work.
Author information
Authors and Affiliations
Contributions
S.B., K.L., A.R.D. and N.F.S. designed the research. S.B., K.L., A.S., M.K.D. and T.K. performed the research. S.B., A.R.D. and N.F.S. wrote the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Information
Supplementary Methods, Supplementary Notes 1–6, Supplementary Discussion, Supplementary References, Supplementary Figures 1–13. (PDF 21093 kb)
Rights and permissions
About this article
Cite this article
Banerjee, S., Lo, K., Daddysman, M. et al. Biphasic growth dynamics control cell division in Caulobacter crescentus. Nat Microbiol 2, 17116 (2017). https://doi.org/10.1038/nmicrobiol.2017.116
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/nmicrobiol.2017.116
This article is cited by
-
Mechanical feedback promotes bacterial adaptation to antibiotics
Nature Physics (2021)
-
Probing bacterial cell wall growth by tracing wall-anchored protein complexes
Nature Communications (2021)
-
Untargeted metabolomics links glutathione to bacterial cell cycle progression
Nature Metabolism (2020)
-
Modeling homeostasis mechanisms that set the target cell size
Scientific Reports (2020)
-
Cell size sensing—a one-dimensional solution for a three-dimensional problem?
BMC Biology (2019)