Abstract
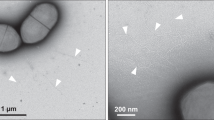
Type IV pili (T4P) are filamentous appendages found on many Bacteria and Archaea. They are helical fibres of pilin proteins assembled by a multi-component macromolecular machine we call the basal body. Based on pilin features, T4P are classified into type IVa pili (T4aP) and type IVb pili (T4bP)1,2. T4aP are more widespread and are involved in cell motility3, DNA transfer4, host predation5 and electron transfer6. T4bP are less prevalent and are mainly found in enteropathogenic bacteria, where they play key roles in host colonization7. Following similar work on T4aP machines8,9, here we use electron cryotomography10 to reveal the three-dimensional in situ structure of a T4bP machine in its piliated and non-piliated states. The specific machine we analyse is the Vibrio cholerae toxin-coregulated pilus machine (TCPM). Although only about half of the components of the TCPM show sequence homology to components of the previously analysed Myxococcus xanthus T4aP machine (T4aPM), we find that their structures are nevertheless remarkably similar. Based on homologies with components of the M. xanthus T4aPM and additional reconstructions of TCPM mutants in which the non-homologous proteins are individually deleted, we propose locations for all eight TCPM components within the complex. Non-homologous proteins in the T4aPM and TCPM are found to form similar structures, suggesting new hypotheses for their functions and evolutionary histories.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Strom, M. S. & Lory, S. Structure–function and biogenesis of the type IV pili. Annu. Rev. Microbiol. 47, 565–596 (1993).
Craig, L., Pique, M. E. & Tainer, J. A. Type IV pilus structure and bacterial pathogenicity. Nat. Rev. Microbiol. 2, 363–378 (2004).
Mattick, J. S. Type IV pili and twitching motility. Annu. Rev. Microbiol. 56, 289–314 (2002).
Chen, I. & Dubnau, D. DNA uptake during bacterial transformation. Nat. Rev. Microbiol. 2, 241–249 (2004).
Evans, K. J., Lambert, C. & Sockett, R. E. Predation by Bdellovibrio bacteriovorus HD100 requires type IV pili. J. Bacteriol. 189, 4850–4859 (2007).
Reguera, G. et al. Extracellular electron transfer via microbial nanowires. Nature 435, 1098–1101 (2005).
Roux, N., Spagnolo, J. & de Bentzmann, S. Neglected but amazingly diverse type IVb pili. Res. Microbiol. 163, 659–673 (2012).
Chang, Y. W. et al. Architecture of the type IVa pilus machine. Science 351, aad2001 (2016).
Gold, V. A., Salzer, R., Averhoff, B. & Kuhlbrandt, W. Structure of a type IV pilus machinery in the open and closed state. eLife 4, e07380 (2015).
Oikonomou, C. M. & Jensen, G. J. A new view into prokaryotic cell biology from electron cryotomography. Nat. Rev. Microbiol. 14, 205–220 (2016).
Craig, L. et al. Type IV pilus structure by cryo-electron microscopy and crystallography: implications for pilus assembly and functions. Mol. Cell 23, 651–662 (2006).
Tammam, S. et al. Characterization of the PilN, PilO and PilP type IVa pilus subcomplex. Mol. Microbiol. 82, 1496–1514 (2011).
Sampaleanu, L. M. et al. Periplasmic domains of pseudomonas aeruginosa PilN and PilO form a stable heterodimeric complex. J. Mol. Biol. 394, 143–159 (2009).
Karuppiah, V. & Derrick, J. P. Structure of the PilM-PilN inner membrane type IV pilus biogenesis complex from Thermus thermophilus. J. Biol. Chem. 286, 24434–24442 (2011).
Korotkov, K. V. et al. Structural and functional studies on the interaction of GspC and GspD in the type II secretion system. PLoS Pathogens 7, e1002228 (2011).
Karuppiah, V., Collins, R. F., Thistlethwaite, A., Gao, Y. & Derrick, J. P. Structure and assembly of an inner membrane platform for initiation of type IV pilus biogenesis. Proc. Natl Acad. Sci. USA 110, E4638–E4647 (2013).
Karuppiah, V., Hassan, D., Saleem, M. & Derrick, J. P. Structure and oligomerization of the PilC type IV pilus biogenesis protein from Thermus thermophilus. Proteins 78, 2049–2057 (2010).
Abendroth, J. et al. The three-dimensional structure of the cytoplasmic domains of EpsF from the type 2 secretion system of Vibrio cholerae. J. Struct. Biol. 166, 303–315 (2009).
Abendroth, J., Murphy, P., Sandkvist, M., Bagdasarian, M. & Hol, W. G. The X-ray structure of the type II secretion system complex formed by the N-terminal domain of EpsE and the cytoplasmic domain of EpsL of Vibrio cholerae. J. Mol. Biol. 348, 845–855 (2005).
Yamagata, A. & Tainer, J. A. Hexameric structures of the archaeal secretion ATPase GspE and implications for a universal secretion mechanism. EMBO J. 26, 878–890 (2007).
Misic, A. M., Satyshur, K. A. & Forest, K. T. P. Aeruginosa PilT structures with and without nucleotide reveal a dynamic type IV pilus retraction motor. J. Mol. Biol. 400, 1011–1021 (2010).
Satyshur, K. A. et al. Crystal structures of the pilus retraction motor PilT suggest large domain movements and subunit cooperation drive motility. Structure 15, 363–376 (2007).
Lu, C. et al. Hexamers of the type II secretion ATPase GspE from Vibrio cholerae with increased ATPase activity. Structure 21, 1707–1717 (2013).
Kolappan, S. & Craig, L. Structure of the cytoplasmic domain of TcpE, the inner membrane core protein required for assembly of the Vibrio cholerae toxin-coregulated pilus. Acta. Crystallogr. D 69, 513–519 (2013).
Krebs, S. J. & Taylor, R. K. Protection and attachment of Vibrio cholerae mediated by the toxin-coregulated pilus in the infant mouse model. J. Bacteriol. 193, 5260–5270 (2011).
Taylor, R. K., Miller, V. L., Furlong, D. B. & Mekalanos, J. J. Identification of a pilus colonization factor that is coordinately regulated with cholera toxin. Ann. Sclavo. Collana Monogr. 3, 51–61 (1986).
Manning, P. A. The tcp gene cluster of Vibrio cholerae. Gene 192, 63–70 (1997).
Li, J., Egelman, E. H. & Craig, L. Structure of the Vibrio cholerae type IVb pilus and stability comparison with the Neisseria gonorrhoeae type IVa pilus. J. Mol. Biol. 418, 47–64 (2012).
Taylor, R. K., Miller, V. L., Furlong, D. B. & Mekalanos, J. J. Use of phoA gene fusions to identify a pilus colonization factor coordinately regulated with cholera toxin. Proc. Natl Acad. Sci. USA 84, 2833–2837 (1987).
Gao, Y., Hauke, C. A., Marles, J. M. & Taylor, R. K. Effects of tcpB mutations on biogenesis and function of the toxin-coregulated pilus, the type IVb pilus of Vibrio cholerae. J. Bacteriol. 198, 2818–2828 (2016).
Bose, N. & Taylor, R. K. Identification of a TcpC–TcpQ outer membrane complex involved in the biogenesis of the toxin-coregulated pilus of Vibrio cholerae. J. Bacteriol. 187, 2225–2232 (2005).
Tripathi, S. A. & Taylor, R. K. Membrane association and multimerization of TcpT, the cognate ATPase ortholog of the Vibrio cholerae toxin-coregulated-pilus biogenesis apparatus. J. Bacteriol. 189, 4401–4409 (2007).
Strom, M. S., Nunn, D. N. & Lory, S. A single bifunctional enzyme, PilD, catalyzes cleavage and N-methylation of proteins belonging to the type IV pilin family. Proc. Natl Acad. Sci. USA 90, 2404–2408 (1993).
Iredell, J. R. & Manning, P. A. Translocation failure in a type-4 pilin operon: rfb and tcpT mutants in Vibrio cholerae. Gene 192, 71–77 (1997).
Megli, C. J. & Taylor, R. K. Secretion of TcpF by the Vibrio cholerae toxin-coregulated pilus biogenesis apparatus requires an N-terminal determinant. J. Bacteriol. 195, 2718–2727 (2013).
Kirn, T. J., Bose, N. & Taylor, R. K. Secretion of a soluble colonization factor by the TCP type 4 pilus biogenesis pathway in Vibrio cholerae. Mol. Microbiol. 49, 81–92 (2003).
Hall, R. H., Vial, P. A., Kaper, J. B., Mekalanos, J. J. & Levine, M. M. Morphological studies on fimbriae expressed by Vibrio cholerae 01. Microb. Pathog. 4, 257–265 (1988).
Kolappan, S., Ng, D., Yang, G., Harn, T. & Craig, L. Crystal structure of the minor pilin CofB, the initiator of CFA/III pilus assembly in enterotoxigenic Escherichia coli. J. Biol. Chem. 290, 25805–25818 (2015).
Taniguchi, T. et al. Gene cluster for assembly of pilus colonization factor antigen III of enterotoxigenic Escherichia coli. Infect. Immun. 69, 5864–5873 (2001).
Souza, D. P. et al. A component of the Xanthomonadaceae type IV secretion system combines a VirB7 motif with a N0 domain found in outer membrane transport proteins. PLoS Pathogens 7, e1002031 (2011).
Berry, J. L. et al. Structure and assembly of a trans-periplasmic channel for type IV pili in Neisseria meningitidis. PLoS Pathogens 8, e1002923 (2012).
Reichow, S. L., Korotkov, K. V., Hol, W. G. & Gonen, T. Structure of the cholera toxin secretion channel in its closed state. Nat. Struct. Mol. Biol. 17, 1226–1232 (2010).
Tosi, T. et al. Structural similarity of secretins from type II and type III secretion systems. Structure 22, 1348–1355 (2014).
Lieberman, J. A. et al. Outer membrane targeting, ultrastructure, and single molecule localization of the enteropathogenic Escherichia coli type IV pilus secretin BfpB. J. Bacteriol. 194, 1646–1658 (2012).
Daniel, A. et al. Interaction and localization studies of enteropathogenic Escherichia coli type IV bundle-forming pilus outer membrane components. Microbiology 152, 2405–2420 (2006).
Gomez-Duarte, O. G. et al. Genetic diversity of the gene cluster encoding longus, a type IV pilus of enterotoxigenic Escherichia coli. J. Bacteriol. 189, 9145–9149 (2007).
McCallum, M. et al. PilN binding modulates the structure and binding partners of the Pseudomonas aeruginosa type IVa pilus protein PilM. J. Biol. Chem. 291, 11003-15 (2016).
Bischof, L. F., Friedrich, C., Harms, A., Sogaard-Andersen, L. & van der Does, C. The type IV pilus assembly ATPase PilB of Myxococcus xanthus interacts with the inner membrane platform protein PilC and the nucleotide-binding protein PilM. J. Biol. Chem. 291, 6946–6957 (2016).
Friedrich, C., Bulyha, I. & Sogaard-Andersen, L. Outside-in assembly pathway of the type IV pilus system in Myxococcus xanthus. J. Bacteriol. 196, 378–390 (2014).
Tivol, W. F., Briegel, A. & Jensen, G. J. An improved cryogen for plunge freezing. Microsc. Microanal. 14, 375–379 (2008).
Zheng, S. Q. et al. UCSF tomography: an integrated software suite for real-time electron microscopic tomographic data collection, alignment, and reconstruction. J. Struct. Biol. 157, 138–147 (2007).
Kremer, J. R., Mastronarde, D. N. & McIntosh, J. R. Computer visualization of three-dimensional image data using IMOD. J. Struct. Biol. 116, 71–76 (1996).
Agulleiro, J. I. & Fernandez, J. J. Fast tomographic reconstruction on multicore computers. Bioinformatics 27, 582–583 (2011).
Nicastro, D. et al. The molecular architecture of axonemes revealed by cryoelectron tomography. Science 313, 944–948 (2006).
Ulrich, L. E. & Zhulin, I. B. The MiST2 database: a comprehensive genomics resource on microbial signal transduction. Nucleic Acids Res. 38, D401–D407 (2010).
Camacho, C. et al. BLAST+: architecture and applications. BMC Bioinformatics 10, 421 (2009).
Adebali, O., Ortega, D. R. & Zhulin, I. B. CDvist: a webserver for identification and visualization of conserved domains in protein sequences. Bioinformatics 31, 1475–1477 (2015).
Soding, J. Protein homology detection by HMM-HMM comparison. Bioinformatics 21, 951–960 (2005).
Juncker, A. S. et al. Prediction of lipoprotein signal peptides in Gram-negative bacteria. Protein Sci. 12, 1652–1662 (2003).
Kawahara, K. et al. Homo-trimeric structure of the type IVb minor pilin CofB suggests mechanism of CFA/III pilus assembly in human enterotoxigenic Escherichia coli. J. Mol. Biol. 428, 1209–1226 (2016).
Acknowledgements
The authors thank C. Oikonomou and C. Shaffer for discussions and editorial assistance. This work was supported by NIH grant R01 AI127401 to G.J.J., the Howard Hughes Medical Institute and the John Templeton Foundation as part of the Boundaries of Life project. The opinions expressed in this publication are those of the authors and do not necessarily reflect the views of the John Templeton Foundation.
Author information
Authors and Affiliations
Contributions
Y.-W.C. and A.K. collected, processed and analysed the ECT data. D.R.O. performed the bioinformatics analyses. L.A.R. assisted with ECT data processing. G.K., J.A.S. and R.K.T. provided the V. cholerae strains. Y.-W.C., A.K., D.R.O. and G.J.J. wrote the paper.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Information
Supplementary Discussion, Supplementary Figures 1–7, Supplementary Tables 1–3, Supplementary References. (PDF 14686 kb)
Supplementary Video 1
Comparison of the TCPM conformation in piliated and non-piliated states. Conformational differences between piliated and non-piliated TCPMs are revealed by morphing from one density to another. (MOV 575 kb)
Rights and permissions
About this article
Cite this article
Chang, YW., Kjær, A., Ortega, D. et al. Architecture of the Vibrio cholerae toxin-coregulated pilus machine revealed by electron cryotomography. Nat Microbiol 2, 16269 (2017). https://doi.org/10.1038/nmicrobiol.2016.269
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/nmicrobiol.2016.269
This article is cited by
-
Bdellovibrio predation cycle characterized at nanometre-scale resolution with cryo-electron tomography
Nature Microbiology (2023)
-
CryoEM structure of the type IVa pilus secretin required for natural competence in Vibrio cholerae
Nature Communications (2020)
-
Type IV pili: dynamics, biophysics and functional consequences
Nature Reviews Microbiology (2019)
-
Multiple conformations facilitate PilT function in the type IV pilus
Nature Communications (2019)
-
In vivo structure of the Legionella type II secretion system by electron cryotomography
Nature Microbiology (2019)