Abstract
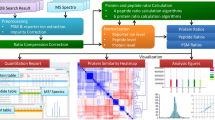
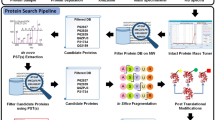
Tandem mass spectrometry (MS/MS) allows for the rapid identification of many types of post-translational modifications (PTMs), especially those that can be detected by a diagnostic mass shift in one or more peptide fragment ions (for example, phosphorylation). But some PTMs (for example, SUMOs and other ubiquitin-like modifiers) themselves produce multiple fragment ions; combined with fragments from the modified target peptide, a complex overlapping fragmentation pattern is thus generated, which is uninterpretable by standard peptide sequencing software. Here we introduce SUMmOn, an automated pattern recognition tool that detects diagnostic PTM fragment ion series within complex MS/MS spectra, to identify modified peptides and modification sites within these peptides. Using SUMmOn, we demonstrate for the first time that human SUMO-1 multimerizes in vitro primarily via three N-terminal lysines, Lys7, Lys16 and Lys17. Notably, our method is theoretically applicable to any type of modification or chemical moiety generating a unique fragment ion pattern.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Aebersold, R. & Goodlett, D.R. Mass spectrometry in proteomics. Chem. Rev. 101, 269–295 (2001).
Steen, H. & Mann, M. The ABC's (and XYZ's) of peptide sequencing. Nat. Rev. Mol. Cell Biol. 5, 699–711 (2004).
Schwartz, D.C. & Hochstrasser, M. A superfamily of protein tags: ubiquitin, SUMO and related modifiers. Trends Biochem. Sci. 28, 321–328 (2003).
Welchman, R.L., Gordon, C. & Mayer, R.J. Ubiquitin and ubiquitin-like proteins as multifunctional signals. Nat. Rev. Mol. Cell Biol. 6, 599–609 (2005).
Melchior, F. SUMO–nonclassical ubiquitin. Annu. Rev. Cell Dev. Biol. 16, 591–626 (2000).
Johnson, E.S. Protein modification by SUMO. Annu. Rev. Biochem. 73, 355–382 (2004).
Rosas-Acosta, G., Russell, W.K., Deyrieux, A., Russell, D.H. & Wilson, V.G. A universal strategy for proteomic studies of SUMO and other ubiquitin-like modifiers. Mol. Cell. Proteomics 4, 56–72 (2005).
Vertegaal, A.C. et al. A proteomic study of SUMO-2 target proteins. J. Biol. Chem. 279, 33791–33798 (2004).
Zhao, Y., Kwon, S.W., Anselmo, A., Kaur, K. & White, M.A. Broad spectrum identification of cellular small ubiquitin-related modifier (SUMO) substrate proteins. J. Biol. Chem. 279, 20999–21002 (2004).
Li, T. et al. Sumoylation of heterogeneous nuclear ribonucleoproteins, zinc finger proteins, and nuclear pore complex proteins: a proteomic analysis. Proc. Natl. Acad. Sci. USA 101, 8551–8556 (2004).
Gocke, C., Yu, H. & Kang, J. Systematic identification and analysis of mammalian SUMO substrates. J. Biol. Chem. 280, 5004–5012 (2005).
Denison, C. et al. A proteomic strategy for gaining insights into protein sumoylation in yeast. Mol. Cell. Proteomics 4, 246–254 (2005).
Wykoff, D. & O'Shea, E. Identification of sumoylated proteins by systematic immunoprecipitation of the budding yeast proteome. Mol. Cell. Proteomics 4, 73–83 (2005).
Panse, V.G., Hardeland, U., Werner, T., Kuster, B. & Hurt, E. A proteome-wide approach identifies sumoylated substrate proteins in yeast. J. Biol. Chem. 279, 41346–41351 (2004).
Hannich, J.T. et al. Defining the SUMO-modified proteome by multiple approaches in Saccharomyces cerevisiae. J. Biol. Chem. 280, 4102–4110 (2005).
Wohlschlegel, J.A., Johnson, E.S., Reed, S.I. & Yates, J.R., III. Global analysis of protein sumoylation in Saccharomyces cerevisiae. J. Biol. Chem. 279, 45662–45668 (2004).
Rodriguez, M.S., Dargemont, C. & Hay, R.T. SUMO-1 conjugation in vivo requires both a consensus modification motif and nuclear targeting. J. Biol. Chem. 276, 12654–12659 (2001).
Sampson, D.A., Wang, M. & Matunis, M.J. The small ubiquitin-like modifier-1 (SUMO-1) consensus sequence mediates Ubc9 binding and is essential for SUMO-1 modification. J. Biol. Chem. 276, 21664–21669 (2001).
Kamitani, T. et al. Identification of three major sentrinization sites in PML. J. Biol. Chem. 273, 26675–26682 (1998).
Eng, J.K., McCormack, A.L. & Yates, J.R., III. An approach to correlate tandem mass spectral data of peptides with amino acid sequences in a protein database. J. Am. Soc. Mass Spectrom. 5, 976–989 (1994).
Nesvizhskii, A.I., Keller, A., Kolker, E. & Aebersold, R. A statistical model for identifying proteins by tandem mass spectrometry. Anal. Chem. 75, 4646–4658 (2003).
Matunis, M.J., Coutavas, E. & Blobel, G. A novel ubiquitin-like modification modulates the partitioning of the Ran-GTPase-activating protein RanGAP1 between the cytosol and the nuclear pore complex. J. Cell Biol. 135, 1457–1470 (1996).
Matunis, M.J., Wu, J. & Blobel, G. SUMO-1 modification and its role in targeting the Ran GTPase-activating protein, RanGAP1, to the nuclear pore complex. J. Cell Biol. 140, 499–509 (1998).
Rogers, R.S., Horvath, C.M. & Matunis, M.J. SUMO modification of STAT1 and its role in PIAS-mediated inhibition of gene activation. J. Biol. Chem. 278, 30091–30097 (2003).
Tatham, M.H. et al. Polymeric chains of SUMO-2 and SUMO-3 are conjugated to protein substrates by SAE1/SAE2 and Ubc9. J. Biol. Chem. 276, 35368–35374 (2001).
Pichler, A., Gast, A., Seeler, J.S., Dejean, A. & Melchior, F. The nucleoporin RanBP2 has SUMO1 E3 ligase activity. Cell 108, 109–120 (2002).
Yang, M., Hsu, C-T., Ting, C-Y., Liu, L.F. & Hwang, J. Assembly of a polymeric chain of SUMO1 on human topoisomerase I in vitro. J. Biol. Chem. 281, 8264–8274 (2006).
Pedrioli, P.G. et al. A common open representation of mass spectrometry data and its application to proteomics research. Nat. Biotechnol. 22, 1459–1466 (2004).
Gygi, S.P., Rist, B., Griffin, T.J., Eng, J. & Aebersold, R. Proteome analysis of low-abundance proteins using multidimensional chromatography and isotope-coded affinity tags. J. Proteome Res. 1, 47–54 (2002).
Keller, A., Nesvizhskii, A.I., Kolker, E. & Aebersold, R. Empirical statistical model to estimate the accuracy of peptide identifications made by MS/MS and database search. Anal. Chem. 74, 5383–5392 (2002).
Acknowledgements
We thank A.-C. Gingras, J. Eng, O. Vitek, X. Li, A. Nesvizhskii and N. Zhang for technical advice and critical reading of the manuscript. This project was funded in part by federal funds from the US National Heart, Lung and Blood Institute, National Institutes of Health, under contract number N01-HV-28179. X.D.Z. and M.J.M. were supported by a grant from the National Institutes of Health (GM060980). B.R. holds the Canada Research Chair (Tier 2) in Proteomics and Molecular Medicine, with support from the Canada Foundation for Innovation, the Ontario Innovation Trust, and the McLaughlin Centre for Molecular Medicine.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Fig. 1
Screenshot of the introductory SUMmOn user interface. (PDF 456 kb)
Supplementary Fig. 2
SUMmOn identifies a known RanGAP1 conjugation site modified with SUMO-2 and a known RanGAP1 conjugation site modified with SUMO-3. (PDF 1788 kb)
Supplementary Fig. 3
SUMmOn identifies a known SUMO-2 conjugation site within SUMO-2. (PDF 320 kb)
Supplementary Fig. 4
SUMmOn identifies a known RanGAP1 conjugation site modified with Smt3. (PDF 1018 kb)
Supplementary Fig. 5
SUMmOn identifies SUMO conjugation sites in RanGAP1 and SUMO-1. (PDF 996 kb)
Supplementary Fig. 6
Identification of SUMO-1 multimerization sites. (PDF 540 kb)
Supplementary Table 1
Data used for modification site identification in this work. (PDF 93 kb)
Supplementary Table 2
SUMO sequence comparison. (PDF 218 kb)
Rights and permissions
About this article
Cite this article
Pedrioli, P., Raught, B., Zhang, XD. et al. Automated identification of SUMOylation sites using mass spectrometry and SUMmOn pattern recognition software. Nat Methods 3, 533–539 (2006). https://doi.org/10.1038/nmeth891
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nmeth891
This article is cited by
-
Prediction and analysis of multiple protein lysine modified sites based on conditional wasserstein generative adversarial networks
BMC Bioinformatics (2021)
-
A computationally engineered RAS rheostat reveals RAS–ERK signaling dynamics
Nature Chemical Biology (2017)
-
An improved method for identifying SUMOylation sites of viral proteins
Virologica Sinica (2017)
-
In vitro assay to determine SUMOylation sites on protein substrates
Nature Protocols (2016)
-
Proteome-wide identification of SUMO modification sites by mass spectrometry
Nature Protocols (2015)