Abstract
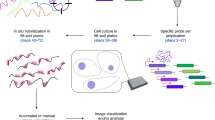
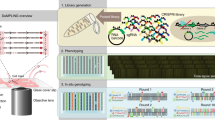
RNA interference (RNAi) is a powerful tool to study gene function in cultured cells. Transfected cell microarrays in principle allow high-throughput phenotypic analysis after gene knockdown by microscopy. But bottlenecks in imaging and data analysis have limited such high-content screens to endpoint assays in fixed cells and determination of global parameters such as viability. Here we have overcome these limitations and developed an automated platform for high-content RNAi screening by time-lapse fluorescence microscopy of live HeLa cells expressing histone-GFP to report on chromosome segregation and structure. We automated all steps, including printing transfection-ready small interfering RNA (siRNA) microarrays, fluorescence imaging and computational phenotyping of digital images, in a high-throughput workflow. We validated this method in a pilot screen assaying cell division and delivered a sensitive, time-resolved phenoprint for each of the 49 endogenous genes we suppressed. This modular platform is scalable and makes the power of time-lapse microscopy available for genome-wide RNAi screens.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
International Human Genome Sequencing Consortium. Finishing the euchromatic sequence of the human genome. Nature 431, 931–945 (2004).
Kittler, R. et al. An endoribonuclease-prepared siRNA screen in human cells identifies genes essential for cell division. Nature 432, 1036–1040 (2004).
Berns, K. et al. A large-scale RNAi screen in human cells identifies new components of the p53 pathway. Nature 428, 431–437 (2004).
Pelkmans, L. et al. Genome-wide analysis of human kinases in clathrin- and caveolae/raft-mediated endocytosis. Nature 436, 78–86 (2005).
Zhu, C. et al. Functional analysis of human microtubule-based motor proteins, the kinesins and dyneins, in mitosis/cytokinesis using RNA interference. Mol. Biol. Cell 16, 3187–3199 (2005).
Sonnichsen, B. et al. Full-genome RNAi profiling of early embryogenesis in Caenorhabditis elegans. Nature 434, 462–469 (2005).
Ziauddin, J. & Sabatini, D.M. Microarrays of cells expressing defined cDNAs. Nature 411, 107–110 (2001).
Erfle, H. et al. siRNA cell arrays for high-content screening microscopy. Biotechniques 37, 454–458, 460, 462 (2004).
Wheeler, D.B., Carpenter, A.E. & Sabatini, D.M. Cell microarrays and RNA interference chip away at gene function. Nat. Genet. 37 (Suppl.), S25–S30 (2005).
Gerlich, D. & Ellenberg, J. 4D imaging to assay complex dynamics in live specimens. Nat. Cell Biol. 4 (Suppl.), S14–S19 (2003).
Sumara, I. et al. Roles of polo-like kinase 1 in the assembly of functional mitotic spindles. Curr. Biol. 14, 1712–1722 (2004).
Kanda, T. & Wahl, G.M. The dynamics of acentric chromosomes in cancer cells revealed by GFP-based chromosome labeling strategies. J. Cell. Biochem. (Suppl.) 35, 107–114 (2000).
Liebel, U. et al. A microscope-based screening platform for large-scale functional protein analysis in intact cells. FEBS Lett. 554, 394–398 (2003).
Huang, K. & Murphy, R.F. From quantitative microscopy to automated image understanding. J. Biomed. Opt. 9, 893–912 (2004).
Conrad, C. et al. Automatic identification of subcellular phenotypes on human cell arrays. Genome Res. 14, 1130–1136 (2004).
Meraldi, P. & Sorger, P.K. A dual role for Bub1 in the spindle checkpoint and chromosome congression. EMBO J. 24, 1621–1633 (2005).
Liu, X., Lei, M. & Erikson, L. Normal cells, but not cancer cells, survive severe plk1 depletion. Mol. Cell. Biol. 26, 2093–2108 (2006).
Gruss, O.J. et al. Chromosome-induced microtubule assembly mediated by Tpx2 is required for spindle formation in HeLA cells. Nat. Cell Biol. 4, 871–879 (2002).
Zhu, C., Bossy-Wetzel, E. & Jiang, W. Recruitment of MKLP1 to the spindle midzone/midbody by INCENP is essential for midbody formation and completion of cytokinesis in human cells. Biochem. J. 389, 373–381 (2005).
Hirota, T. et al. Distinct functions of condensin I and II in mitotic chromosome assembly. J. Cell Sci. 117, 6435–6445 (2004).
Seul, M., Lawrence, O. & Sammon, M. Practical Algorithms for Image Analysis (Cambridge Univ. Press, Cambridge, UK, 2000).
Huang, K. & Murphy, R.F. Boosting accuracy of automated classification of fluorescence microscope images for location proteomics. BMC Bioinformatics 5, 78 (2004).
Acknowledgements
We thank S. Narumiya (Kyoto University, Kyoto) and T. Hirota (Institute of Molecular Pathology; IMP; Vienna) for HeLa 'Kyoto' cells; W. Huber (European Bioinformatics Institute; EBI; Hinxton) for advice on statistical analysis of kinetic data; O. Gruss (Zentrum für Molekulare Biologie Heidelberg; ZNBH; Heidelberg) for TPX2 antibody; J.-M. Peters (IMP; Vienna) for RPE cells; I. Hoffmann (Deutsches Krebsforschungszentrum; DKFZ; Heidelberg) for U2OS cells; H. Runz (Univ. Heidelberg) for primary human fibroblasts; Chroma Inc. for providing customized emission filter sets free of charge; EMBL's IT Services group (B. Kindler, M. Hemberger, R. Lück) for support; Olympus Biosystems, Hamilton and Bio-Rad for continuous support; Cenix BioScience GmbH for siRNA design and for providing the A549 cells; and Ambion Europe, Ltd. for providing siRNAs for validation. This project was funded by grants to J.E. within the MitoCheck consortium by the European Commission (FP6-503464) as well as in part by the Federal Ministry of Education and Research (BMBF) in the framework of the National Genome Research Network (NGFN) (NGFN-2 SMP-RNAi, FKZ01GR0403 to J.E. and NGFN-2 SMP-Cell FKZ01GR0423, NGFN-1 FKZ01GR0101, FKZ01KW0013 to R.P.).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Fig. 1
siRNA knock-down efficiency. (PDF 200 kb)
Supplementary Fig. 2
Examples of detected RNAi phenotypes. (PDF 379 kb)
Supplementary Table 1
Summary of siRNA sequences (PDF 85 kb)
Supplementary Table 2
Summary of RT-PCR Primer (PDF 62 kb)
Supplementary Video 1
siRNA SCRAMBLED (MOV 1154 kb)
Supplementary Video 2
siRNA INCENP - Segregation problem leading to multinucleated cells (MOV 360 kb)
Supplementary Video 3
siRNA SYNE2 - Metaphase alignment problem followed by segregation followed by apoptosis (MOV 377 kb)
Supplementary Video 4
siRNA PLK1 - Prometaphase arrest followed by apoptosis (MOV 391 kb)
Supplementary Video 5
siRNA CDC16 - Metaphase alignment problems followed by apoptosis (MOV 713 kb)
Supplementary Video 6
siRNA NUP107 - Metaphase alignment problems followed by apoptosis (MOV 604 kb)
Supplementary Video 7
siRNA NUMA1 - Apoptosis (MOV 310 kb)
Rights and permissions
About this article
Cite this article
Neumann, B., Held, M., Liebel, U. et al. High-throughput RNAi screening by time-lapse imaging of live human cells. Nat Methods 3, 385–390 (2006). https://doi.org/10.1038/nmeth876
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nmeth876
This article is cited by
-
Morphodynamical cell state description via live-cell imaging trajectory embedding
Communications Biology (2023)
-
Single object profiles regression analysis (SOPRA): a novel method for analyzing high-content cell-based screens
BMC Bioinformatics (2022)
-
ER-mitochondria contact sites in neurodegeneration: genetic screening approaches to investigate novel disease mechanisms
Cell Death & Differentiation (2021)
-
Genetic screening identifies a SUMO protease dynamically maintaining centromeric chromatin
Nature Communications (2020)
-
Systematic epistatic mapping of cellular processes
Cell Division (2017)