Abstract
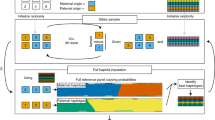
We describe an efficient, accurate and robust whole-genome genotyping (WGG) assay based on a two-color, single-base extension (SBE), single-nucleotide polymorphism (SNP)-scoring step. We report genotyping results for biallelic International HapMap quality control (QC) SNPs using a single probe per locus. We show scalability, throughput and accuracy of the system by resequencing homozygous loci from our 100k Human-1 Genotyping BeadChip.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
References
Syvanen, A.C. Nat. Genet. 37 (Suppl.), S5–10 (2005).
Craig, D.W. & Stephan, D.A. Expert Rev. Mol. Diagn. 5, 159–170 (2005).
Matsuzaki, H. et al. Nat. Methods 1, 109–111 (2004).
Carlson, C.S., Eberle, M.A., Kruglyak, L. & Nickerson, D.A. Nature 429, 446–452 (2004).
Gunderson, K.L., Steemers, F.J., Lee, G., Mendoza, L.G. & Chee, M.S. Nat. Genet. 37, 549–554 (2005).
Gunderson, K.L. et al. In Genetic variance detection: technologies for pharmacogenomics (ed. Hecker, K.H.) 221–235 (DNA Press, Eagleville, Pennsylvania, 2005).
Pastinen, T. et al. Genome Res. 10, 1031–1042 (2000).
Kuppuswamy, M.N. et al. Proc. Natl. Acad. Sci. USA 88, 1143–1147 (1991).
Pastinen, T., Kurg, A., Metspalu, A., Peltonen, L. & Syvanen, A.C. Genome Res. 7, 606–614 (1997).
Hirschhorn, J.N. et al. Proc. Natl. Acad. Sci. USA 97, 12164–12169 (2000).
Wang, H.Y. et al. Genome Res. 15, 276–283 (2005).
Gunderson, K.L. et al. Genome Res. 14, 870–877 (2004).
Pinkel, D., Straume, T. & Gray, J.W. Proc. Natl. Acad. Sci. USA 83, 2934–2938 (1986).
Fan, J.B. et al. Cold Spring Harb. Symp. Quant. Biol. LXVIII, 69–78 (2003).
Acknowledgements
We thank C. Allred for help with the grant applications and administration, M. Hansen for assistance in data analysis, E. Johnson, M. Graige and manufacturing department for bead pools and arrays, N. Espinosa and J. Stuelpnagel for manuscript suggestions. This work was supported in part by grants from the US National Institutes of Health (2R44CA103406-02) and National Cancer Institute (1R43CA108391-01).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
All authors have a financial interest in Illumina either via employment and/or stock ownership.
Supplementary information
Supplementary Fig. 1
Effect of SSB in the extension reaction. (PDF 71 kb)
Supplementary Table 1
Allele count and distribution over the SNP classes. (PDF 29 kb)
Rights and permissions
About this article
Cite this article
Steemers, F., Chang, W., Lee, G. et al. Whole-genome genotyping with the single-base extension assay. Nat Methods 3, 31–33 (2006). https://doi.org/10.1038/nmeth842
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nmeth842
This article is cited by
-
Integration of Infinium and Axiom SNP array data in the outcrossing species Malus × domestica and causes for seemingly incompatible calls
BMC Genomics (2021)
-
A distinct metabolic response characterizes sensitivity to EZH2 inhibition in multiple myeloma
Cell Death & Disease (2021)
-
Assessing the genetic diversity of cowpea [Vigna unguiculata (L.) Walp.] germplasm collections using phenotypic traits and SNP markers
BMC Genetics (2020)
-
Digging out molecular markers associated with low salinity tolerance of Nannochloropsis oceanica through bulked mutant analysis
Journal of Oceanology and Limnology (2020)
-
Inheritance of pre-emergent metribuzin tolerance and putative gene discovery through high-throughput SNP array in wheat (Triticum aestivum L.)
BMC Plant Biology (2019)