Abstract
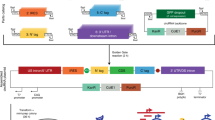
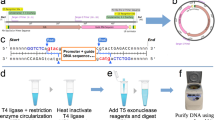
Here we report the development of a gene-synthesis technology, circular assembly amplification. In this approach, we first constructed exonuclease-resistant circular DNA via simultaneous ligation of oligonucleotides. Exonuclease- and subsequent mismatch cleaving endonuclease–mediated degradation of the resulting ligation mixture eliminated error-rich products, thereby substantially improving gene-synthesis quality. We used this method to construct genes encoding a small thermostable DNA polymerase, a highly repetitive DNA sequence and large (>4 kb) constructs.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Menzella, H.G. et al. Nat. Biotechnol. 23, 1171–1176 (2005).
Han, J.S. & Boeke, J.D. Nature 429, 314–318 (2004).
Stemmer, W.P., Crameri, A., Ha, K.D., Brennan, T.M. & Heyneker, H.L. Gene 164, 49–53 (1995).
Smith, H.O., Hutchison, C.A., III, Pfannkoch, C. & Venter, J.C. Proc. Natl. Acad. Sci. USA 100, 15440–15445 (2003).
Tian, J. et al. Nature 432, 1050–1054 (2004).
Kodumal, S.J. et al. Proc. Natl. Acad. Sci. USA 101, 15573–15578 (2004).
Xiong, A.S. et al. Nucleic Acids Res. 32, e98 (2004).
Fuhrmann, M., Oertel, W., Berthold, P. & Hegemann, P. Nucleic Acids Res. 33, e58 (2005).
Young, L. & Dong, Q. Nucleic Acids Res. 32, e59 (2004).
Carr, P.A. et al. Nucleic Acids Res. 32, e162 (2004).
Ling, H., Boudsocq, F., Woodgate, R. & Yang, W. Cell 107, 91–102 (2001).
Richardson, S.M., Wheelan, S.J., Yarrington, R.M. & Boeke, J.D. Genome Res. 16, 550–556 (2006).
SantaLucia, J., Jr. Proc. Natl. Acad. Sci. USA 95, 1460–1465 (1998).
Geu-Flores, F., Nour-Eldin, H.H., Nielsen, M.T. & Halkier, B.A. Nucleic Acids Res. 35, e55 (2007).
Acknowledgements
We thank M. Umbarger, M. Price and J. Aach for critical comments on the manuscript; F. Issacs for comments on experimental design; an anonymous reviewer's suggestion to test repetitive DNA sequence synthesis. We acknowledge funding from US Department of Energy for GTL Center support. D.B is a Damon Runyon Fellow supported by the Damon Runyon Cancer Research Foundation (DRG#1911-06).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
G.M.C. is a scientific advisory board member of Codon Devices.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–5, Supplementary Table 1, Supplementary Methods (PDF 1555 kb)
Rights and permissions
About this article
Cite this article
Bang, D., Church, G. Gene synthesis by circular assembly amplification. Nat Methods 5, 37–39 (2008). https://doi.org/10.1038/nmeth1136
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nmeth1136
This article is cited by
-
Ligation Based Assembly and Polymerase Chain Reaction-Based Assembly for Extraordinary Adenine/Thymine Rich DNA
Proceedings of the National Academy of Sciences, India Section B: Biological Sciences (2018)
-
Genomes by design
Nature Reviews Genetics (2015)
-
Sequential growth of long DNA strands with user-defined patterns for nanostructures and scaffolds
Nature Communications (2015)
-
Large-scale de novo DNA synthesis: technologies and applications
Nature Methods (2014)
-
Will recombinant DNA technology become obsolete?
Nature India (2012)