Abstract
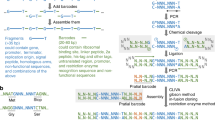
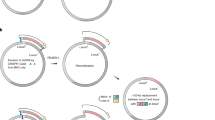
We describe a new cloning method, sequence and ligation–independent cloning (SLIC), which allows the assembly of multiple DNA fragments in a single reaction using in vitro homologous recombination and single-strand annealing. SLIC mimics in vivo homologous recombination by relying on exonuclease-generated ssDNA overhangs in insert and vector fragments, and the assembly of these fragments by recombination in vitro. SLIC inserts can also be prepared by incomplete PCR (iPCR) or mixed PCR. SLIC allows efficient and reproducible assembly of recombinant DNA with as many as 5 and 10 fragments simultaneously. SLIC circumvents the sequence requirements of traditional methods and functions much more efficiently at very low DNA concentrations when combined with RecA to catalyze homologous recombination. This flexibility allows much greater versatility in the generation of recombinant DNA for the purposes of synthetic biology.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Smith, H.O. & Wilcox, K.W. A restriction enzyme from Hemophilus influenzae. I. Purification and general properties. J. Mol. Biol. 51, 379–391 (1970).
Danna, K. & Nathans, D. Specific cleavage of simian virus 40 DNA by restriction endonuclease of Hemophilus influenzae. Proc. Natl. Acad. Sci. USA 68, 2913–2917 (1971).
Cohen, S.N., Chang, A.C., Boyer, H.W. & Helling, R.B. Construction of biologically functional bacterial plasmids in vitro. Proc. Natl. Acad. Sci. USA 70, 3240–3244 (1973).
Backman, K. & Ptashne, M. Maximizing gene expression on a plasmid using recombination in vitro. Cell 13, 65–71 (1978).
Hutchison, C.A. et al. Mutagenesis at a specific position in a DNA sequence. J. Biol. Chem. 253, 6551–6560 (1978).
Rumsby, G. An introduction to PCR techniques. PCR Methods Mol. Biol. 324, 75–89 (2006).
Saiki, R.K. et al. Enzymatic amplification of beta-globin genomic sequences and restriction site analysis for diagnosis of sickle cell anemia. Science 230, 1350–1354 (1985).
Ho, S.N., Hunt, H.D., Horton, R.M., Pullen, J.K. & Pease, L.R. Site-directed mutagenesis by overlap extension using the polymerase chain reaction. Gene 77, 51–59 (1989).
Liu, Q., Li, M.Z., Liebham, D., Cortez, D. & Elledge, S.J. The univector plasmid fusion system, a method for rapid construction of recombinant DNA without restriction enzymes. Curr. Biol. 8, 1300–1309 (1998).
Hartley, J.L., Temple, G.F. & Brasch, M.A. DNA cloning using in vitro site-specific recombination. Genome Res. 10, 1788–1795 (2000).
Walhout, A.J. et al. GATEWAY recombinational cloning: application to the cloning of large numbers of open reading frames or ORFeomes. Methods Enzymol. 328, 575–592 (2000).
Bethke, B. & Sauer, B. Segmental genomic replacement by Cre-mediated recombination: Genotoxic stress activation of the p53 promoter in single-copy transformants. Nucleic Acids Res. 25, 2828–2834 (1997).
Nebert, D.W., Dalton, T.P., Stuart, G.W. & Carvan, M.J. Gene-swap knock-in cassette in mice to study allelic differences in human genes. Ann. NY Acad. Sci. 919, 148–170 (2000).
Siegel, R.W. et al. Recombinatorial cloning using heterologous lox sites. Genome Res. 14, 1119–1129 (2004).
Li, M.Z. & Elledge, S.J. MAGIC: An in vivo genetic method for the rapid construction of recombinant DNA molecules. Nat. Genet. 37, 311–319 (2005).
Amundsen, S.K. & Smith, G.R. Interchangeable parts of the Escherichia coli recombination machinery. Cell 112, 741–744 (2003).
Kuzminov, A. Recombinational repair of DNA damage in Escherichia coli and bacteriophage lambda. Microbiol. Mol. Biol. Rev. 63, 751–813 (1999).
Konforti, B.B. & Davis, R.W. 3′ homologous free ends are required for stable joint molecule formation by the RecA and single-stranded binding proteins of Escherichia coli. Proc. Natl. Acad. Sci. USA 84, 690–694 (1987).
Tillett, D. & Neilan, B.A. Enzyme-free cloning: A rapid method to clone PCR products independent of vector restriction enzyme sites. Nucleic Acids Res. 27, e26 (1999).
Aslanidis, C. & de Jong, P.J. Ligation-independent cloning of PCR products (LIC-PCR). Nucleic Acids Res. 18, 6069–6074 (1990).
Haun, R.S., Servanti, I.M. & Moss, J. Rapid, reliable ligation-independent cloning of PCR products using modified plasmid vectors. Biotechniques 13, 515–518 (1992).
Aslanidis, C., de Jong, P.J. & Schmitz, G. Minimal length requirement of the single-stranded tails for ligation-independent cloning (LIC) of PCR products. PCR Methods Appl. 4, 172–177 (1994).
Cheo, D.L. et al. Concerted assembly and cloning of multiple DNA fragments using in vitro site-specific recombination: Functional analysis of multi-site expression clones. Gen. Res. 14, 2111–2120 (2004).
Acknowledgements
We thank B. Wanner, G. Hannon and T. Moore for providing plasmids, bacterial strains and advice concerning their use. We thank M. Schlabach for comments on the manuscript. This work was supported by a grant from US National Institutes of Health. S.J.E. is an investigator with the Howard Hughes Medical Institute.
Author information
Authors and Affiliations
Contributions
M.Z.L. performed all experiments. S.J.E. helped in experimental design. M.Z.L. and S.J.E. wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Fig. 1
Effect of insert to vector ratio on SLIC. (PDF 34 kb)
Supplementary Fig. 2
Effect of insert size on SLIC. (PDF 35 kb)
Supplementary Table 1
T4 DNA polymerase is the most efficient and reproducible exonuclease for SLIC cloning. (DOC 20 kb)
Supplementary Protocol 1
SLIC Sub-cloning using T4 DNA polymerase treated inserts without RecA. (PDF 56 kb)
Supplementary Protocol 2
SLIC Sub-cloning using T4 DNA polymerase treated inserts with RecA. (PDF 55 kb)
Supplementary Protocol 3
SLIC Sub-cloning using iPCR or mixed PCR products. (PDF 51 kb)
Rights and permissions
About this article
Cite this article
Li, M., Elledge, S. Harnessing homologous recombination in vitro to generate recombinant DNA via SLIC. Nat Methods 4, 251–256 (2007). https://doi.org/10.1038/nmeth1010
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nmeth1010
This article is cited by
-
High-throughput process development from gene cloning to protein production
Microbial Cell Factories (2023)
-
Amino acid positions near the active site determine the reduced activity of human ACOD1 compared to murine ACOD1
Scientific Reports (2023)
-
Optimal tagging strategies for illuminating expression profiles of genes with different abundance in zebrafish
Communications Biology (2023)
-
CRISPR-induced DNA reorganization for multiplexed nucleic acid detection
Nature Communications (2023)
-
Metabolic engineering of Escherichia coli for the production of (R)-α-lipoic acid
Biotechnology Letters (2023)