Abstract
Sequencing the RNA in a biological sample can unlock a wealth of information, including the identity of bacteria and viruses, the nuances of alternative splicing or the transcriptional state of organisms. However, current methods have limitations due to short read lengths and reverse transcription or amplification biases. Here we demonstrate nanopore direct RNA-seq, a highly parallel, real-time, single-molecule method that circumvents reverse transcription or amplification steps. This method yields full-length, strand-specific RNA sequences and enables the direct detection of nucleotide analogs in RNA.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Accession codes
References
Wang, Z., Gerstein, M. & Snyder, M. RNA-Seq: a revolutionary tool for transcriptomics. Nat. Rev. Genet. 10, 57–63 (2009).
Wu, J.Q. et al. Systematic analysis of transcribed loci in ENCODE regions using RACE sequencing reveals extensive transcription in the human genome. Genome Biol. 9, R3 (2008).
Kozarewa, I. et al. Amplification-free Illumina sequencing-library preparation facilitates improved mapping and assembly of (G+C)-biased genomes. Nat. Methods 6, 291–295 (2009).
Lipson, D. et al. Quantification of the yeast transcriptome by single-molecule sequencing. Nat. Biotechnol. 27, 652–658 (2009).
Mamanova, L. et al. FRT-seq: amplification-free, strand-specific transcriptome sequencing. Nat. Methods 7, 130–132 (2010).
Ozsolak, F. et al. Direct RNA sequencing. Nature 461, 814–818 (2009).
Pan, Q., Shai, O., Lee, L.J., Frey, B.J. & Blencowe, B.J. Deep surveying of alternative splicing complexity in the human transcriptome by high-throughput sequencing. Nat. Genet. 40, 1413–1415 (2008).
Steijger, T. et al. Assessment of transcript reconstruction methods for RNA-seq. Nat. Methods 10, 1177–1184 (2013).
Thomas, S., Underwood, J.G., Tseng, E. & Holloway, A.K. Long-read sequencing of chicken transcripts and identification of new transcript isoforms. PLoS One 9, e94650 (2014).
Vilfan, I.D. et al. Analysis of RNA base modification and structural rearrangement by single-molecule real-time detection of reverse transcription. J. Nanobiotechnology 11, 8 (2013).
Clamer, M., Höfler, L., Mikhailova, E., Viero, G. & Bayley, H. Detection of 3′-end RNA uridylation with a protein nanopore. ACS Nano 8, 1364–1374 (2014).
Smith, A.M., Abu-Shumays, R., Akeson, M. & Bernick, D.L. Capture, unfolding, and detection of individual tRNA molecules using a nanopore device. Front. Bioeng. Biotechnol. 3, 91 (2015).
Wu, T.D. & Watanabe, C.K. GMAP: a genomic mapping and alignment program for mRNA and EST sequences. Bioinformatics 21, 1859–1875 (2005).
Pertea, M. et al. StringTie enables improved reconstruction of a transcriptome from RNA-seq reads. Nat. Biotechnol. 33, 290–295 (2015).
Byrne, A. et al. Nanopore long-read RNAseq reveals widespread transcriptional variation among the surface receptors of individual B cells. Nat. Commun. 8, 16027 (2017).
Deamer, D., Akeson, M. & Branton, D. Three decades of nanopore sequencing. Nat. Biotechnol. 34, 518–524 (2016).
Oxford Nanopore Technologies Ltd. Direct RNA sequencing https://community.nanoporetech.com/protocols/direct-rna-sequencing/v/drs_9026_v1_revj_15dec201 (2016).
The HDF Group. Hierarchical data format, version 5, 1997–2017. http://www.hdfgroup.org/HDF5/.
Quinlan, A.R. & Hall, I.M. BEDTools: a flexible suite of utilities for comparing genomic features. Bioinformatics 26, 841–842 (2010).
Krzywinski, M. et al. Circos: an information aesthetic for comparative genomics. Genome Res. 19, 1639–1645 (2009).
Larkin, M.A. et al. Clustal W and Clustal X version 2.0. Bioinformatics 23, 2947–2948 (2007).
Li, H. & Durbin, R. Burrows–Wheeler Alignment Tool http://bio-bwa.sourceforge.net/bwa.shtml (2012).
Fariselli, P., Martelli, P.L. & Casadio, R. A new decoding algorithm for hidden Markov models improves the prediction of the topology of all-beta membrane proteins. BMC Bioinformatics 6, S12 (2005).
Author information
Authors and Affiliations
Contributions
D.R.G., A.J.H., J. Clarke and D.J.T. conceived the experiments. D.R.G. led the project. D.R.G., E.A.S., D.J., A.J.H., J.H.L., P.J., A.W., M.J., J.K., S.M. and L.M. designed and performed the experiments. J.H.L. tested, engineered and developed the motor protein. J.H.L., S.M., L.M., D.R.G., E.A.S., A.J.H., M.B., D.J., A.W. and E.J.W. designed or assessed motor protein mutations and the sequencing adaptor. D.J.T., D.R.G. and E.A.S. developed the library preparation. E.A.S. and J.K. created custom RNA templates. B.S. wrote custom analysis tools and performed analysis of all sequence data sets. N.P., T.A. and M.B. expressed and purified proteins. M.J., J. Ciccone and S.S. designed and prepared plasmids. M.J., E.J.W., L.J., S.Y., D.R.G., E.A.S., D.J., A.J.H., M.B., J.H.L. and D.B. assessed sequencing performance of buffers, voltages and pores. C.W. wrote squiggle-consensus algorithms. J.B., C.W., D.B., J.H.L., M.B. and S.Y. trained RNA basecallers or analyzed modified base data. D.J.T., B.S., D.R.G., S.J. and C.W. wrote the manuscript. A.J.H., S.Y. and P.J. contributed to the figures or to editing of the manuscript.
Corresponding author
Ethics declarations
Competing interests
All authors are employees of Oxford Nanopore Technologies and are shareholders and/or share option holders.
Integrated supplementary information
Supplementary Figure 2 Analysis of direct RNA method
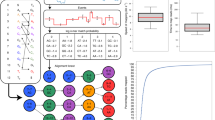
a) Distribution of mean quality values for all reads in the direct RNA yeast dataset. b) Distribution of read accuracies from the retrained direct RNA basecaller.
Supplementary Figure 3 Technical replicates of the direct RNA method.
The correlation between read counts after mapping to the yeast transcriptome for 5 technical replicates of the Direct RNA method. The five technical replicates were separate library preparations of yeast run on separate MinION Chips. Above the diagonal are pairwise scatter plots and below the diagonal are pairwise density plots (Rho from Spearman’s rank correlation is shown over each plot). Each scatter or density plot includes all transcripts in the annotation: n = 6713 transcripts.
Supplementary Figure 4 Effect of increasing number of PCR cycles
The effect of number of PCR cycles on bias, read length and deviation from expected read counts for ERCC spike-ins. Three independent replicates were performed at each cycle number totaling 24 separate nanopore cDNA sequencing runs. Error bars denote s.e.m..
Supplementary Figure 5 Direct RNA versus Illumina: comparison of bias.
Correlation between read counts and transcript length for a) direct RNA (Pearson’s r = 0.13, p = 5.4e-29) or b) Illumina (Pearson’s r = 0.3, p = 7e-141) yeast datasets. Correlation between read counts and GC content for c) direct RNA (Pearson’s r = 0.013, p = 0.29) or d) Illumina (Pearson’s r = 0.19, p = 1.6e-58) yeast datasets. In each of (a-d), all transcripts were included: n = 6713 transcripts. e) Correlation between mean quality of aligned read portions and the GC content of aligned reference portions for direct RNA yeast dataset (Pearson’s r = 0.082, p = 0, n = 2,777,523 alignments). The correlation coefficients and the corresponding two-sided p-values were calculated using the stats.pearsonr function from the scipy Python package.
Supplementary Figure 6 Gene-level and transcript-level correlations to SIRV control.
Reads aligned using the spliced-alignment strategy and correlations calculated a) at the transcript level (Spearman’s Rho = 0.62, p = 9.5e-9, n = 69 transcripts) or b) at the gene level (Spearman’s Rho = 0.61, p = 0.15, n = 7 genes) for the SIRV E2 dataset. The correlation coefficients and the corresponding two-sided p-values were calculated using the stats.spearmanr function from the scipy Python package.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–7 (PDF 908 kb)
Rights and permissions
About this article
Cite this article
Garalde, D., Snell, E., Jachimowicz, D. et al. Highly parallel direct RNA sequencing on an array of nanopores. Nat Methods 15, 201–206 (2018). https://doi.org/10.1038/nmeth.4577
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nmeth.4577
This article is cited by
-
Single-molecule RNA sizing enables quantitative analysis of alternative transcription termination
Nature Communications (2024)
-
NAP-seq reveals multiple classes of structured noncoding RNAs with regulatory functions
Nature Communications (2024)
-
Single-molecule epitranscriptomic analysis of full-length HIV-1 RNAs reveals functional roles of site-specific m6As
Nature Microbiology (2024)
-
Direct RNA sequencing coupled with adaptive sampling enriches RNAs of interest in the transcriptome
Nature Communications (2024)
-
SQANTI3: curation of long-read transcriptomes for accurate identification of known and novel isoforms
Nature Methods (2024)