Abstract
Mammalian genomes are folded in a hierarchy of compartments, topologically associating domains (TADs), subTADs and looping interactions. Here, we describe 3DNetMod, a graph theory-based method for sensitive and accurate detection of chromatin domains across length scales in Hi-C data. We identify nested, partially overlapping TADs and subTADs genome wide by optimizing network modularity and varying a single resolution parameter. 3DNetMod can be applied broadly to understand genome reconfiguration in development and disease.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
References
Bullmore, E. & Sporns, O. Nat. Rev. Neurosci. 10, 186–198 (2009).
Girvan, M. & Newman, M.E. Proc. Natl. Acad. Sci. USA 99, 7821–7826 (2002).
Lanctôt, C., Cheutin, T., Cremer, M., Cavalli, G. & Cremer, T. Nat. Rev. Genet. 8, 104–115 (2007).
Dekker, J., Marti-Renom, M.A. & Mirny, L.A. Nat. Rev. Genet. 14, 390–403 (2013).
Lieberman-Aiden, E. et al. Science 326, 289–293 (2009).
Phillips-Cremins, J.E. et al. Cell 153, 1281–1295 (2013).
Rao, S.S. et al. Cell 159, 1665–1680 (2014).
Dixon, J.R. et al. Nature 485, 376–380 (2012).
Nora, E.P. et al. Nature 485, 381–385 (2012).
Sexton, T. et al. Cell 148, 458–472 (2012).
Dali, R. & Blanchette, M. Nucleic Acids Res. 45, 2994–3005 (2017).
Forcato, M. et al. Nat. Methods 14, 679–685 (2017).
Newman, M.E. Proc. Natl. Acad. Sci. USA 103, 8577–8582 (2006).
Blondel, V.D., Guillaume, J.-L., Lambiotte, R. & Lefebvre, E. J. Stat. Mech. Theory. E. 2008, P10008 (2008).
Newman, M.E.J. Nat. Phys. 8, 25–31 (2012).
Won, H. et al. Nature 538, 523–527 (2016).
Rubinov, M. & Sporns, O. Neuroimage 56, 2068–2079 (2011).
Ball, B., Karrer, B. & Newman, M.E.J. Phys. Rev. E 84, 036103 (2011).
Weinreb, C. & Raphael, B.J. Bioinformatics 32, 1601–1609 (2016).
Jiang, Y. et al. Nat. Genet. 49, 1239–1250 (2017).
Servant, N. et al. Genome Biol. 16, 259 (2015).
Imakaev, M. et al. Nat. Methods 9, 999–1003 (2012).
Good, B.H., de Montjoye, Y.-A. & Clauset, A. Phys. Rev. E 81, 046106 (2010).
Traud, A.L., Kelsic, E.D., Mucha, P.J. & Porter, M.A. SIAM Rev. 53, 526–543 (2011).
Hubert, L. & Arabie, P. J. Classif. 2, 193–218 (1985).
Doron, K.W., Bassett, D.S. & Gazzaniga, M.S. Proc. Natl. Acad. Sci. USA 109, 18661–18668 (2012).
Lohse, C., Bassett, D.S., Lim, K.O. & Carlson, J.M. PLoS Comput. Biol. 10, e1003712 (2014).
Maslov, S. & Sneppen, K. Science 296, 910–913 (2002).
Acknowledgements
J.E.P.-C. is a New York Stem Cell Foundation (NYSCF) Robertson Investigator and an Alfred P. Sloan Foundation Fellow. This work was funded by The New York Stem Cell Foundation (J.E.P.-C.), the Alfred P. Sloan Foundation (J.E.P.-C.), the NIH Director's New Innovator Award (1DP2MH11024701; J.E.P.-C.), a 4D Nucleome Common Fund grant (1U01HL12999801; J.E.P.-C.) and a joint NSF-NIGMS grant to support research at the interface of the biological and mathematical sciences (1562665; J.E.P.-C.). This material is based upon work supported by the National Science Foundation Graduate Research Fellowship under DGE-1321851 (H.N.). D.S.B. would also like to acknowledge support from the John D. and Catherine T. MacArthur Foundation.
Author information
Authors and Affiliations
Contributions
J.E.P.-C., D.S.B. and H.K.N. conceived of the study. H.K.N., D.J.E., S.G., H.H., K.R.T. and J.K. implemented the computational pipeline. J.E.P.-C., H.K.N., D.J.E. and D.S.B. wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
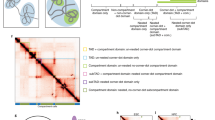
Supplementary Figure 1 3D genome folding data can be represented mathematically as a network.
(A) Schematic representation of a Hi-C heatmap with Megabase-scale topologically associated domains (TADs) and nested sub-TADs within the first TAD. Node numbers correspond to genomic bins. Yellow arrows indicate the corners of domains. (B) Model of the potential community structure underlying proximity ligation data depicted in panel (A). Black dots with numbers represent nodes corresponding to the genomic coordinates represented in each bin. (C) Spring force diagram representation of a genome-folding network. Edge colors depict edge weight ranging from low (grey) to high (red).
Supplementary Figure 2 Effect of partition number in 3DNetMod-MMCP on consensus partition.
(A-C) An approximately 6 Mb Hi-C region of human cortical plate cells1. Communities identified using 3DNetMod-MMCP with (A) 20 underlying partitions, (B) 100 underlying partitions and (C) 1000 underlying partitions at a γ value of 1.03 are outlined in green. (D-F) The underlying individual partitions in the partition block are used to determine the consensus partition with a γvalue of 1.03.
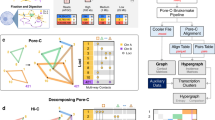
Supplementary Figure 3 Overview of 3DNetMod method.
(A) Each chromosome is parsed into overlapping regions with user-defined size and overlap. (B-D) Gamma Plateau Sweep (3DNetMod-GPS). (B) Five representative regions equally distributed along the chromosome are randomly rewired to create null networks. (C) Modularity is computed across a range of γ values for the five representative regions and their randomly rewired counterparts. The convergence point between the modularity curves of real and random networks is determined for each region (grey vertical line). The maximal informative γ value is computed for each chromosome as the maximum γ at the convergence points for the five regions. (D) To identify γ values that give rise to high-confidence communities, we defined ‘plateaus’ of finely incremented (0.01 step size) γ values that give rise to the same number of communities. The midpoints for all plateaus that are greater than a user-defined minimal plateau size (indicated with a star) are selected as γ values for high-confidence community detection. (E-F) Modularity maximization and consensus partition (3DNetMod-MMCP). (E) For each selected γ value, the network is partitioned 20 times into communities. (F) A consensus partition is computed from the 20 partitions. (G) 3DNetMod-MMCP is repeated across all γ values selected with 3DNetMod-GPS, resulting in the initial detected set of communities across all length scales. Small communities (less than or equal to 4 nodes) and communities at region edges are removed. (H-L) Hierarchical Spatial Variance Minimization (3DNetMod-HSVM). (H) First, communities are stratified by size: ≤400kb (magenta), 401kb – 800kb (green), 801kb – 1600kb (cyan), 1601kb – 3Mb (indigo) and > 3Mb (orange). (I) Spatial variance is then computed at each community boundary. (J) Boundary spatial variance values for each community of a given size stratification across the entire chromosome are pooled. A user-defined variance threshold is computed for each size-stratified set of communities. (K) If a community has a boundary with higher spatial variance than the size-specific threshold, it is removed from further consideration. (L) The resulting variance-thresholded communities can be further refined by merging significantly overlapping domains or domains called in regions devoid of nested substructure (see Supplemental Methods). The final community list represents high-confidence 3DNetMod domains.
Supplementary Figure 4 Effect of region size on domains detected.
(A) Representative 3 Mb Hi-C heatmap from human cortical plate1. Domains identified with 3DNetMod-GPS, 3DNetMod-MMCP and 3DNetMod-HSVM using 3Mb regions are outlined. (B) Top panel: Heatmap of the same 3Mb in (A) with domains identified using 6Mb outlined. Bottom panel: The full 6Mb region used for region detection. (C) Domains identified using 12Mb regions.
Supplementary Figure 5 Gamma plateau sweep (3DNetMod-GPS) sensitively partitions Hi-C data into a hierarchy of partially overlapping TADs and subTADs genome-wide.
(A) A 6 Mb-sized Hi-C region from human cortical plate tissue1. (B) Representative randomly rewired 6Mb Hi-C network in which the locations of existing edges have been reassigned according to expected edge weights. (C) Modularity, Q, versus structural resolution parameter, γ, for a representative 6Mb region and its corresponding randomly re-wired network. The maximal useful γ value is the point of convergence between real and random networks and is indicated by a vertical grey line. (D) Number of communities as a function of γ for the Hi-C region shown in (A). Three or more consecutive γ values with the same number of communities are grouped into plateaus. The midpoint of each plateau (indicated by a star) is selected as a valid γ for community detection. (E) Heatmap with domains identified across the selected resolution parameter values. (F) Partition blocks (partitions 1 through 20) and consensus partition (‘C’) for community assignments across the selected γ values. Domains that contact the edge of the network and domains < 200kb have been removed from consideration and are greyed out on partition blocks.
Supplementary Figure 6 Effect of plateau method on 3DNetMod domain detection.
(A-C) Hi-C heatmaps of 2 representative 6 Mb regions (2 rows) from human embryonic cortical plate tissue1 with communities detected by 3DNetMod-GPS and 3DNetMod-MMCP (A) without the plateau method or with plateau sizes of (B) 3 or (C) 8.
Supplementary Figure 7 High variance at potential A/B compartment boundaries.
(A) A 6 Mb Hi-C region from human embryonic cortical plate tissue1. The blocks representing 20 partitions and the corresponding consensus partition are shown beneath the heatmap. Community assignments have no variability across the 20 partitions. An illustrative domain with zero variability in assignment across the 20 partitions is indicated with green arrows. (B) Another 6 Mb Hi-C region from human embryonic cortical plate tissue1. The boundaries of a putative A/B compartment-like structure exhibit high spatial variance across 20 partitions (magenta arrows).
Supplementary Figure 8 Hierarchical spatial variance minimization (3DNetMod-HSVM) dissects bona fide chromatin domains from higher-order architectural features.
(A-B) Two different 6Mb genomic regions from human cortical plate tissue Hi-C1 after gamma plateau sweep (3DNetMod-GPS) to sensitively partition a hierarchy of partially overlapping TADs and subTADs genome-wide. A putative A/B compartment with alternating, non-uniform signal is indicated with an arrow in each heatmap. (C-D) Twenty partitions and consensus partition (labeled as ‘C’) for γ values at which the compartments illustrated in A-B, respectively, were detected. Domain calls at edges of the network are removed and colored in grey in the partition block and consensus partition. The spatial variance across the 20 partitions for each boundary is computed. (E) Histograms of spatial variance from all domain boundaries from chromosome 7 stratified by the size of domain. A variance threshold can be selected for each domain size stratum (red vertical line) to filter higher-order compartments and low-confidence domains from the full detected set. (F-G) Domain calls are removed from consideration (thresholded consensus partition, ‘T’) if either of its boundaries exceeds the length scale-specific variance threshold. (H-I) Final nested hierarchy of domains after 3DNetMod-HSVM.
Supplementary Figure 9 CTCF enrichment at high and low variance boundaries.
(A-D) Average number of wild type mouse neuronal CTCF peaks2 per 40kb centered at boundaries of domains identified with 3DNetMod. Boundaries of domains in the (A) 100 - 400 kb, (B) 401 - 800 kb and (C) 801 -1600 kb size stratum with low (left) or high variance and were removed with variance thresholding (right). (D) Boundaries of domains in the 1601 kb - 3 Mb size stratum. No variance thresholding was performed at this size stratum.
Supplementary Figure 10 Selection of variance thresholds on human embryonic cortical plate tissue Hi-C1.
(A-C) 6 Mb Hi-C heatmaps from human embryonic cortical plate tissue binned at 40 kb1. Domains identified with 3DNetMod-GPS and 3DNetMod-MMCP at the 200 – 400 kb size stratum with 3 different variance thresholds during hierarchical spatial variance minimization (3DNetMod-HSVM). Domains at (A) 0 variance, (B) 70% AUC and (C) 100% AUC thresholds outlined in magenta. The selected threshold is indicated with a green box. (D-L) 3 different variance thresholds at the (D-F) 401 – 800 kb size stratum, (G-I) 801 -1600 kb and (J-L) 1601 kb – 3 Mb size strata.
Supplementary Figure 11 Selection of variance thresholds on mouse cortical tissue Hi-C3.
(A-C) 6 Mb Hi-C heatmaps from mouse cortical tissue binned at 40 kb3. Domains identified with 3DNetMod-GPS and 3DNetMod-MMCP at the 200 – 400 kb size stratum with 3 different variance thresholds during hierarchical spatial variance minimization (3DNetMod-HSVM). Domains at (A) 0 variance, (B) 70% AUC and (C) 100% AUC thresholds outlined in magenta. The selected threshold is indicated with a green box. (D-L) 3 different variance thresholds at the (D-F) 401 – 800 kb, (G-I) 801-1600 kb and (J-L) 1601 kb – 3 Mb size strata.
Supplementary Figure 12 Selection of variance thresholds on mouse neuron Hi-C2.
(A-C) 3 Mb Hi-C heatmaps from mouse neurons binned at 20 kb2. Domains identified with 3DNetMod-GPS and 3DNetMod-MMCP at the 200 – 400 kb size stratum with 3 different variance thresholds during hierarchical spatial variance minimization (3DNetMod-HSVM). Domains at (A) 0 variance, (B) 70% AUC and (C) 100% AUC thresholds outlined in magenta. The selected threshold is indicated with a green box. (D-L) 3 different variance thresholds at the (D-F) 401 – 800 kb, (G-I) 801-1600 kb and (J-L) 1601 kb - 3Mb size strata.
Supplementary Figure 13 Comparison of domain-calling performance across methods on simple simulated Hi-C data (corresponding to Figure 2).
(A) Simulated Hi-C data with simple, non-overlapping domains of variable size. Expected domains (left), DI-HMM + DI sweep (center left), TADtree (center right) and 3DNetMod (right) are shown. (B) Receiver Operating Characteristic (ROC) curves showing the true positive rate and false positive rate of 3DNetMod (magenta), DI-HMM + DI Sweep (teal) and TADtree (blue) domain detection performance in the simple simulated Hi-C network. (C) Simulated Hi-C data with nested, partially overlapping domain structure. Expected domains (left), DI-HMM + DI sweep domains (center left), TADtree domains (center right) and 3DNetMod domains (right) are shown. (D) Full view of the zoomed in simulations shown in (C) (green box).
Supplementary Figure 14 Comparison of domains identified in 40 kb binned human embryonic cortical plate tissue Hi-C1 by leading 3D chromatin domain detection methods.
Three representative 6 Mb Hi-C heatmaps from human embryonic cortical plate tissue1. Domains outlined as identified by (A) Arrowhead4, (B) DI-HMM Domain Caller3 with a sweep of directionality index (DI) values, (C) TADtree5 and (D) 3DNetMod.
Supplementary Figure 15 Comparison of domains identified in 20 kb binned mouse neuron Hi-C2 by leading 3D chromatin domain detection methods.
(A-D) Four representative 3 Mb Hi-C heatmaps from mouse neurons2. Domains outlined as identified by (A) Arrowhead4, (B) TADtree5 with S=50, (C) TADtree with S=100 default parameter and (D) 3DNetMod. Due to intractable run times of TADtree on 20kb-bnined data with S=100, TADtree was run on half of chromosome 18.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–15 and Supplementary Tables 1–3 (PDF 4799 kb)
Source data
Rights and permissions
About this article
Cite this article
Norton, H., Emerson, D., Huang, H. et al. Detecting hierarchical genome folding with network modularity. Nat Methods 15, 119–122 (2018). https://doi.org/10.1038/nmeth.4560
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nmeth.4560
This article is cited by
-
Computational methods for analysing multiscale 3D genome organization
Nature Reviews Genetics (2024)
-
Communities in C. elegans connectome through the prism of non-backtracking walks
Scientific Reports (2023)
-
Mapping robust multiscale communities in chromosome contact networks
Scientific Reports (2023)
-
High-throughput Oligopaint screen identifies druggable 3D genome regulators
Nature (2023)
-
A comparison of topologically associating domain callers over mammals at high resolution
BMC Bioinformatics (2022)