Abstract
We present a strategy for tackling preferred specimen orientation in single-particle cryogenic electron microscopy by employing tilts during data collection. We also describe a tool to quantify the resulting directional resolution using 3D Fourier shell correlation volumes. We applied these methods to determine the structures at near-atomic resolution of the influenza hemagglutinin trimer, which adopts a highly preferred specimen orientation, and of ribosomal biogenesis intermediates, which adopt moderately preferred orientations.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
References
Glaeser, R.M. Nat. Methods 13, 28–32 (2016).
Glaeser, R.M. & Han, B.-G. Biophys. Rep. http://dx.doi.org/10.1007/s41048-016-0026-3 (2016).
Taylor, K.A. & Glaeser, R.M. J. Struct. Biol. 163, 214–223 (2008).
Barth, M., Bryan, R.K. & Hegerl, R. Ultramicroscopy 31, 365–378 (1989).
Penczek, P.A. & Frank, J. in Electron Tomography (ed. Frank, J.) 307–330 (Springer, 2007).
Yip, C.K., Murata, K., Walz, T., Sabatini, D.M. & Kang, S.A. Mol. Cell 38, 768–774 (2010).
Bartesaghi, A., Lecumberry, F., Sapiro, G. & Subramaniam, S. Structure 20, 2003–2013 (2012).
Radermacher, M., Wagenknecht, T., Verschoor, A. & Frank, J. J. Microsc. 146, 113–136 (1987).
Leschziner, A.E. & Nogales, E. J. Struct. Biol. 153, 284–299 (2006).
Su, M. et al. Sci. Adv. 3, e1700325 (2017).
Zhang, K. J. Struct. Biol. 193, 1–12 (2016).
Russo, C.J. & Passmore, L.A. Science 346, 1377–1380 (2014).
Zheng, S.Q. et al. Nat. Methods 14, 331–332 (2017).
Grant, T. & Grigorieff, N. eLife 4, e06980 (2015).
Rosenthal, P.B. & Henderson, R. J. Mol. Biol. 333, 721–745 (2003).
Scheres, S.H. J. Struct. Biol. 180, 519–530 (2012).
Penczek, P.A. Methods Enzymol. 482, 73–100 (2010).
Davis, J.H. et al. Cell 167, 1610–1622.e15 (2016).
Penczek, P.A. J. Struct. Biol. 138, 34–46 (2002).
Sorzano, C.O. et al. J. Struct. Biol. 133, 108–118 (2001).
Baker, M.R., Fan, G. & Serysheva, I.I. Eur. J. Transl. Myol. 25, 35–48 (2015).
Lyumkis, D., Tan, Y.Z. & Baldwin, P.R. Protoc. Exch. http://dx.doi.org/10.1038/protex.2017.055 (2017).
Diebolder, C.A., Faas, F.G., Koster, A.J. & Koning, R.I. J. Struct. Biol. 190, 215–223 (2015).
Dudkina, N.V., Kudryashev, M., Stahlberg, H. & Boekema, E.J. Proc. Natl. Acad. Sci. USA 108, 15196–15200 (2011).
Hohn, M. et al. J. Struct. Biol. 157, 47–55 (2007).
Tang, G. et al. J. Struct. Biol. 157, 38–46 (2007).
Kucukelbir, A., Sigworth, F.J. & Tagare, H.D. Nat. Methods 11, 63–65 (2014).
Wadell, H. J. Geol. 43, 250–280 (1935).
Lyumkis, D., Brilot, A.F., Theobald, D.L. & Grigorieff, N. J. Struct. Biol. 183, 377–388 (2013).
Baxter, W.T., Grassucci, R.A., Gao, H. & Frank, J. J. Struct. Biol. 166, 126–132 (2009).
Grigorieff, N. J. Struct. Biol. 157, 117–125 (2007).
Suloway, C. et al. J. Struct. Biol. 151, 41–60 (2005).
Mindell, J.A. & Grigorieff, N. J. Struct. Biol. 142, 334–347 (2003).
Voss, N.R., Yoshioka, C.K., Radermacher, M., Potter, C.S. & Carragher, B. J. Struct. Biol. 166, 205–213 (2009).
Sorzano, C.O. et al. J. Struct. Biol. 171, 197–206 (2010).
Roseman, A.M. J. Struct. Biol. 145, 91–99 (2004).
Lyumkis, D., Vinterbo, S., Potter, C.S. & Carragher, B. J. Struct. Biol. 184, 417–426 (2013).
Lander, G.C. et al. J. Struct. Biol. 166, 95–102 (2009).
Rubinstein, J.L. & Brubaker, M.A. J. Struct. Biol. 192, 188–195 (2015).
Pettersen, E.F. et al. J. Comput. Chem. 25, 1605–1612 (2004).
Punjani, A., Rubinstein, J.L., Fleet, D.J. & Brubaker, M.A. Nat. Methods 14, 290–296 (2017).
Campbell, M.G., Veesler, D., Cheng, A., Potter, C.S. & Carragher, B. eLife 4, e06380 (2015).
Elmlund, D. & Elmlund, H. J. Struct. Biol. 180, 420–427 (2012).
Acknowledgements
We thank N. Grigorieff, T. Grant, A. Rohou, A. Cheng and A. Noble for their invaluable advice; T. Goddard for incorporating the result of 3D FSC into UCSF Chimera. Molecular graphics and analyses were performed with the UCSF Chimera package (supported by NIGMS P41-GM103311). We thank B. Anderson and J.C. Ducom at TSRI for help with EM data collection and network infrastructure; and we thank F. Dwyer for computational support at the Salk Institute. The work was supported by Agency for Science, Technology and Research Singapore (to Y.Z.T.); the Leona M. and Harry B. Helmsley Charitable Trust Grant 2017-PG-MED001 (to D.L.); the US National Institutes of Health (NIH) (DP5 OD021396-01 to D.L.); the Jane Coffin Child's Foundation postdoctoral fellowship (to J.H.D.); the National Institute of Aging K99 transitional award (AG050749 to J.H.D.); National Institute of General Medical Sciences (GM103310 to C.S.P. and B.C.; GM053757 to J.R.W.); and Simons Foundation (349247 to C.S.P. and B.C.).
Author information
Authors and Affiliations
Contributions
D.L. conceived the idea for this study. B.C. and C.P. provided guidance throughout the study. Y.Z.T. and D.L. performed the cryo-EM experiments, data collection, processing and analysis. D.L. and Y.Z.T. generated the synthetic data sets. P.R.B. and Y.Z.T. coded the 3D FSC program suite. J.H.D. and J.R.W. provided the L17-depleted 50S ribosomal samples. Y.Z.T. and D.L. wrote the manuscript with input from all authors.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
Supplementary Figure 1 Schematic demonstrating the preferred orientation sampling problem in single-particle cryo-EM.
Specimens sticking to the air-water interface or a substrate support of a cryo-EM grid in a single preferred orientation always adopt random in-plane orientations. Tilting of the specimen stage therefore leads to (a) a precession of views around the axis of preferred orientation in a conical manner (precession indicated in green with grey arrows, axis of preferred orientation is indicated by a red arrow). (b) Projections of the object in (a) are displayed for 30° and 60° tilts (theta angle), both at a ϕ angle increment of 15°. Whereas the 30° tilted projections resemble more the “top” views of the object, the 60° tilted projections resemble more the “side” views of the object. (c) In reciprocal space, each Fourier slice inserted into the reconstruction is perpendicular to the direction of its real-space projection image. Therefore, a large missing cone is formed from a reconstruction using 30° tilted images, and a much smaller missing cone is present from a reconstruction using 60° tilted images. No missing cone would be present for (hypothetically) 90° tilted images, although the X/Y plane would be characterized by 1D lines rather than a complete 2D slice. The practical implications of this diagram and anisotropic sampling are demonstrated in Supplementary Figure 2.
Supplementary Figure 2 3D reconstructions of a synthetic, absolutely preferred dataset of an HA trimer presumed to be tilted at various angles.
(a) Euler angle distribution profiles of the synthetic projections. (b) The resulting 3D reconstructions. The red arrow shows the direction of preferred orientation. The grey arrows show the angle at which the synthetic projection images were generated at the various tilt angles. A synthetic dataset of equally sampled HA trimer is also shown for comparison in the last row. (c) Slice through the 3D FSC volumes describing the 3D reconstructions along the X/Z plane. The 3D FSC was thresholded at 0.143, while sphericity was calculated by thresholding the 3D FSC at 0.5 and applying a Gaussian filter of 2 pixels. Due to C3 symmetry along the Z-axis of the reconstruction, only the X/Z slice of the 3D FSC is shown. Missing cones are apparent within the 30°, and especially the 60° 3D FSCs. (d) Plots of the global FSC after reconstruction (solid blue line), the spread of the directional resolutions defined by plus and minus one standard deviation from the mean of the directional resolutions (green area encompassed by green dotted lines) and the histogram of one hundred directional resolutions evenly sampled over the 3D FSC (yellow bars). Labels on the left Y-axis refer to global FSC curves, while labels on the right Y-axis refer to directional FSC histograms. The grey dotted line indicates when FSC = 0.143.
Supplementary Figure 3 HA trimer per-tilt analysis.
HA trimer cryo-EM data was collected (a) untilted, or at tilts of (b) 10°, (c) 20°, (d) 30°, (e) 40°, and (f) 50°. Representative 2D class averages for each tilt angle are shown as insert. (g) The average frame-to-frame shift for each tilt angle is shown for both the first 15 frames and also for the entire range of 100 frames (insert). The exact cause of the slightly higher beam-induced movement within the 0° dataset is unclear and was not observed for the ribosome dataset (Supplementary Figure 6g).
Supplementary Figure 4 Tilting improves resolution and directional isotropy in a direct comparison of HA trimer reconstructions without and with tilting.
(a) Euler angle distribution of the particles. (b) 3D reconstruction superimposed onto a projection of the HA trimer crystal structure (pink). Loss of axial density at typical display thresholds is clearly evident within reconstructions from 0°, 10°, and 20° tilted images. (c) Close-up of a particular region within the reconstruction. Circled region indicates “false positive” density resulting from elongation of the reconstruction along the Z-axis, which progressively disappears within reconstructions from higher tilts characterized by improved directional resolution isotropy. (d) Slice through the 3D FSC describing (top, solid blue outline) half-map, resolution evaluating internal consistency and (bottom, dotted purple outline) map-to-model resolution evaluating external consistency, along with their corresponding sphericity values (e). 3D FSC sphericity values. The slight dip in 3D FSC sphericity at 10° is caused by greater improvements in directional resolution perpendicular to the preferred orientation axis that are not met with a concomitant increase in Z resolution. (f) Graph showing the global half-map resolution (solid blue), map-to-model resolution (dotted purple), and spread of directional resolution (refer to Supplementary Figure 2 for detailed graph description).
Supplementary Figure 5 HA particle titration experiment.
HA trimer reconstructions using subsets of particles from (a) untilted and (b) 40° tilted images. Random subsets of particles from the 130,000 total particles, in multiples of 13,000, were selected for each titration point and refined independently. The half map resolution (using 0.143 threshold) and map-to-model resolution (using 0.5 threshold) against the structure of HA (PDB 3WHE) is shown for each titration point. The Euler angle distribution of the reconstructions at each titration point for the (c) dataset from 0° images and (d) 40° tilted images show that the apparent side views in the untilted reconstruction (green arrows) only appear when more particles are added to the reconstruction, indicating overfitting.
Supplementary Figure 6 Per-tilt analysis of L17-depleted 50S ribosomal assembly intermediates (LSUbL17dep).
Cryo-EM data was collected (a) untilted and at tilts of (b) 10°, (c) 20°, (d) 30°, (e) 40° and (f) 50°. The average frame-to-frame shift for each tilt angle is shown for both the first 15 frames (g) and also for the entire range of 50 frames (insert). (h) The 4 super-classes of L17-depleted 50S ribosomal assembly intermediates are shown.
Supplementary Figure 7 LSUbL17dep – Class B.
For each tilt angle, the figure shows (a) Euler angle distributions of the particles, (b,c,d) 3D FSC at orthogonal views, and (e) global half-map and map-to-model FSC plots with the spread of directional resolutions defined by the green area and 3D FSC values overlaid as histograms. Direction of preferred orientation is indicated by the red arrow.
Supplementary Figure 8 LSUbL17dep – Class C.
For each tilt angle, the figure shows (a) Euler angle distributions of the particles, (b) an alpha helix density from uL29 protein, (c,d,e) 3D FSC at orthogonal views, and (f) global half-map and map-to-model FSC plots with the spread of directional resolutions defined by the green area and 3D FSC values overlaid as histograms. Direction of preferred orientation is indicated by the red arrow.
Supplementary Figure 9 LSUbL17dep – Class D.
For each tilt angle, the figure shows (a) Euler angle distributions of the particles, (b) an alpha helix density from uL29 protein, (c,d,e) 3D FSC at orthogonal views, and (f) global half-map and map-to-model FSC plots with the spread of directional resolutions defined by the green area and 3D FSC values overlaid as histograms. Direction of preferred orientation is indicated by the red arrow.
Supplementary Figure 10 LSUbL17dep – Class E.
For each tilt angle, the figure shows (a) Euler angle distributions of the particles, (b) an alpha helix density from uL29 protein, (c,d,e) 3D FSC at orthogonal views, and (f) global half-map and map-to-model FSC plots with the spread of directional resolutions defined by the green area and 3D FSC values overlaid as histograms. Direction of preferred orientation is indicated by the red arrow.
Supplementary Figure 11 Tilting improves resolution and angular isotropy for all four super-classes of L17-depleted 50S ribosomal intermediates.
For the four super-classes, B-E, the changes in (a) resolution, (b) half-map 3D FSC sphericity, and (c) map-to-model 3D FSC sphericity are plotted with respect to the dataset from untilted images and as a function of tilt angle (see Supplementary Figures 7, 8, 9, 10 for all raw data). Map densities of (d) beta sheets and (e) alpha helices from reconstructions at various tilts for class E are also shown to illustrate the effects of change in resolution and resolution anisotropy (the two are interrelated). Beta sheets are from uL22 and the alpha helices are from uL29. Side chain densities are marked by red asterisks, while beta sheet separation is indicated by a green arrow. The direction of preferred orientation is indicated by the red double-headed arrow. As expected, map isotropy steadily improves with increasing angular tilt, which also affects global resolution and can accordingly facilitate interpretation of structural features. For example, smearing of beta-strands (d) and alpha-helices (e) parallel to the direction of preferred orientation is ameliorated with tilts, which can be especially important at borderline resolutions for interpreting atomic models.
Supplementary Figure 12 Tilt angles up to 50° provide near-atomic resolution single-particle reconstructions.
(a) 3D reconstruction from a preferred oriented LSUbL17dep ribosomal intermediate dataset, where particles were combined from all super-classes but only using data from 50° tilted images. The sample heterogeneity from combining different super-classes is reflected in the local resolution colored onto the reconstruction. Whereas the peripheral density changes, the homogeneous core components are much better resolved. (b) Global half-map 3D FSC and (c) map-to-model 3D FSC. The high tilt angle used for collection results in nearly spherical 3D FSCs, with sphericity of 0.98 and 0.92 respectively. (d) The ten best resolved ribosomal proteins have resolutions of between 4.2 Å and 4.8 Å. (e-g) Map density of (e) bL21, (f) bL20 and (g) uL29 show side chain density and separation of beta strands.
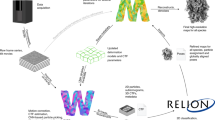
Supplementary Figure 13 Workflow for collection of single particle data at tilts.
Data can be collected at either one or multiple tilt angles. Other than the need to tilt the stage and perform per-particle CTF estimation, the other processing steps are similar to a conventional single particle cryo-EM workflow. In order to analyze the degree of directional anisotropy, 3D FSC is performed at the end (isosurface shown) and its results are visualized in Chimera - when rotating the map, the map’s color and associated directional FSC line component changes on the fly, enabling immediate assessment of resolution anisotropy.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–13 and Supplementary Notes 1–4 (PDF 5577 kb)
Rights and permissions
About this article
Cite this article
Tan, Y., Baldwin, P., Davis, J. et al. Addressing preferred specimen orientation in single-particle cryo-EM through tilting. Nat Methods 14, 793–796 (2017). https://doi.org/10.1038/nmeth.4347
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nmeth.4347
This article is cited by
-
Structural basis of Acinetobacter type IV pili targeting by an RNA virus
Nature Communications (2024)
-
Structural and mechanistic characterization of bifunctional heparan sulfate N-deacetylase-N-sulfotransferase 1
Nature Communications (2024)
-
High-resolution cryo-EM of the human CDK-activating kinase for structure-based drug design
Nature Communications (2024)
-
Structure of the γ-tubulin ring complex-capped microtubule
Nature Structural & Molecular Biology (2024)
-
Time-resolved cryo-EM of G-protein activation by a GPCR
Nature (2024)