Abstract
Millions of cis-regulatory elements are predicted to be present in the human genome, but direct evidence for their biological function is scarce. Here we report a high-throughput method, cis-regulatory element scan by tiling-deletion and sequencing (CREST-seq), for the unbiased discovery and functional assessment of cis-regulatory sequences in the genome. We used it to interrogate the 2-Mb POU5F1 locus in human embryonic stem cells, and identified 45 cis-regulatory elements. A majority of these elements have active chromatin marks, DNase hypersensitivity, and occupancy by multiple transcription factors, which confirms the utility of chromatin signatures in cis-element mapping. Notably, 17 of them are previously annotated promoters of functionally unrelated genes, and like typical enhancers, they form extensive spatial contacts with the POU5F1 promoter. These results point to the commonality of enhancer-like promoters in the human genome.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Accession codes
References
Gerstein, M.B. et al. Architecture of the human regulatory network derived from ENCODE data. Nature 489, 91–100 (2012).
Shen, Y. et al. A map of the cis-regulatory sequences in the mouse genome. Nature 488, 116–120 (2012).
Xie, W. et al. Epigenomic analysis of multilineage differentiation of human embryonic stem cells. Cell 153, 1134–1148 (2013).
Roadmap Epigenomics Consortium. et al. Integrative analysis of 111 reference human epigenomes. Nature 518, 317–330 (2015).
ENCODE Project Consortium. An integrated encyclopedia of DNA elements in the human genome. Nature 489, 57–74 (2012).
Ernst, J. et al. Mapping and analysis of chromatin state dynamics in nine human cell types. Nature 473, 43–49 (2011).
Thurman, R.E. et al. The accessible chromatin landscape of the human genome. Nature 489, 75–82 (2012).
Farh, K.K. et al. Genetic and epigenetic fine mapping of causal autoimmune disease variants. Nature 518, 337–343 (2015).
Gjoneska, E. et al. Conserved epigenomic signals in mice and humans reveal immune basis of Alzheimer's disease. Nature 518, 365–369 (2015).
Canver, M.C. et al. BCL11A enhancer dissection by Cas9-mediated in situ saturating mutagenesis. Nature 527, 192–197 (2015).
Korkmaz, G. et al. Functional genetic screens for enhancer elements in the human genome using CRISPR-Cas9. Nat. Biotechnol. 34, 192–198 (2016).
Rajagopal, N. et al. High-throughput mapping of regulatory DNA. Nat. Biotechnol. 34, 167–174 (2016).
Diao, Y. et al. A new class of temporarily phenotypic enhancers identified by CRISPR/Cas9-mediated genetic screening. Genome Res. 26, 397–405 (2016).
Sanjana, N.E. et al. High-resolution interrogation of functional elements in the noncoding genome. Science 353, 1545–1549 (2016).
Fulco, C.P. et al. Systematic mapping of functional enhancer-promoter connections with CRISPR interference. Science 354, 769–773 (2016).
Jinek, M. et al. A programmable dual-RNA-guided DNA endonuclease in adaptive bacterial immunity. Science 337, 816–821 (2012).
Mojica, F.J., Diez-Villasenor, C., Garcia-Martinez, J. & Almendros, C. Short motif sequences determine the targets of the prokaryotic CRISPR defence system. Microbiology 155, 733–740 (2009).
Sternberg, S.H., Redding, S., Jinek, M., Greene, E.C. & Doudna, J.A. DNA interrogation by the CRISPR RNA-guided endonuclease Cas9. Nature 507, 62–67 (2014).
Zwaka, T.P. & Thomson, J.A. Homologous recombination in human embryonic stem cells. Nat. Biotechnol. 21, 319–321 (2003).
Li, W. et al. MAGeCK enables robust identification of essential genes from genome-scale CRISPR/Cas9 knockout screens. Genome Biol. 15, 554 (2014).
Ware, C.B. et al. Derivation of naive human embryonic stem cells. Proc. Natl. Acad. Sci. USA 111, 4484–4489 (2014).
Buenrostro, J.D., Giresi, P.G., Zaba, L.C., Chang, H.Y. & Greenleaf, W.J. Transposition of native chromatin for fast and sensitive epigenomic profiling of open chromatin, DNA-binding proteins and nucleosome position. Nat. Methods 10, 1213–1218 (2013).
Ghirlando, R. & Felsenfeld, G. CTCF: making the right connections. Genes Dev. 30, 881–891 (2016).
Dixon, J.R., Gorkin, D.U. & Ren, B. Chromatin domains: the unit of chromosome organization. Mol. Cell 62, 668–680 (2016).
Yan, J. et al. Transcription factor binding in human cells occurs in dense clusters formed around cohesin anchor sites. Cell 154, 801–813 (2013).
MacArthur, S. et al. Developmental roles of 21 Drosophila transcription factors are determined by quantitative differences in binding to an overlapping set of thousands of genomic regions. Genome Biol. 10, R80 (2009).
Yip, K.Y. et al. Classification of human genomic regions based on experimentally determined binding sites of more than 100 transcription-related factors. Genome Biol. 13, R48 (2012).
Chandra, T. et al. Independence of repressive histone marks and chromatin compaction during senescent heterochromatic layer formation. Mol. Cell 47, 203–214 (2012).
Hnisz, D. et al. Super-enhancers in the control of cell identity and disease. Cell 155, 934–947 (2013).
Li, G. et al. Extensive promoter-centered chromatin interactions provide a topological basis for transcription regulation. Cell 148, 84–98 (2012).
Engreitz, J.M. et al. Local regulation of gene expression by lncRNA promoters, transcription and splicing. Nature 539, 452–455 (2016).
Chia, N.Y. et al. A genome-wide RNAi screen reveals determinants of human embryonic stem cell identity. Nature 468, 316–320 (2010).
Rogakou, E.P., Boon, C., Redon, C. & Bonner, W.M. Megabase chromatin domains involved in DNA double-strand breaks in vivo. J. Cell Biol. 146, 905–916 (1999).
Downs, J.A., Lowndes, N.F. & Jackson, S.P. A role for Saccharomyces cerevisiae histone H2A in DNA repair. Nature 408, 1001–1004 (2000).
Burma, S., Chen, B.P., Murphy, M., Kurimasa, A. & Chen, D.J. ATM phosphorylates histone H2AX in response to DNA double-strand breaks. J. Biol. Chem. 276, 42462–42467 (2001).
Dixon, J.R. et al. Chromatin architecture reorganization during stem cell differentiation. Nature 518, 331–336 (2015).
Core, L.J. et al. Analysis of nascent RNA identifies a unified architecture of initiation regions at mammalian promoters and enhancers. Nat. Genet. 46, 1311–1320 (2014).
Paralkar, V.R. et al. Unlinking an lncRNA from its associated cis element. Mol. Cell 62, 104–110 (2016).
Handoko, L. et al. CTCF-mediated functional chromatin interactome in pluripotent cells. Nat. Genet. 43, 630–638 (2011).
DeMare, L.E. et al. The genomic landscape of cohesin-associated chromatin interactions. Genome Res. 23, 1224–1234 (2013).
Kieffer-Kwon, K.R. et al. Interactome maps of mouse gene regulatory domains reveal basic principles of transcriptional regulation. Cell 155, 1507–1520 (2013).
Ji, X. et al. 3D chromosome regulatory landscape of human pluripotent cells. Cell Stem Cell 18, 262–275 (2016).
Tang, Z. et al. CTCF-mediated human 3D genome architecture reveals chromatin topology for transcription. Cell 163, 1611–1627 (2015).
Jin, F. et al. A high-resolution map of the three-dimensional chromatin interactome in human cells. Nature 503, 290–294 (2013).
Rao, S.S. et al. A 3D map of the human genome at kilobase resolution reveals principles of chromatin looping. Cell 159, 1665–1680 (2014).
Dixon, J.R. et al. Topological domains in mammalian genomes identified by analysis of chromatin interactions. Nature 485, 376–380 (2012).
Arnold, C.D. et al. Genome-wide quantitative enhancer activity maps identified by STARR-seq. Science 339, 1074–1077 (2013).
Thakore, P.I. et al. Highly specific epigenome editing by CRISPR-Cas9 repressors for silencing of distal regulatory elements. Nat. Methods 12, 1143–1149 (2015).
Diao, Y., Fang, R., Li, B. & Ren, B. A dual sgRNA mediated tiling-deletion based genetic screen to identify regulatory DNA sequence in mammalian cells. Protoc. exch. http://dx.doi.org/10.1038/protex.2017.037 (2017).
Langmead, B., Trapnell, C., Pop, M. & Salzberg, S.L. Ultrafast and memory-efficient alignment of short DNA sequences to the human genome. Genome Biol. 10, R25 (2009).
Wu, X., Kriz, A.J. & Sharp, P.A. Target specificity of the CRISPR-Cas9 system. Quant. Biol. 2, 59–70 (2014).
Meng, Z. et al. Berbamine inhibits the growth of liver cancer cells and cancer-initiating cells by targeting Ca2+/calmodulin-dependent protein kinase II. Mol. Cancer Ther. 12, 2067–2077 (2013).
Kolde, R., Laur, S., Adler, P. & Vilo, J. Robust rank aggregation for gene list integration and meta-analysis. Bioinformatics 28, 573–580 (2012).
Rajagopal, N. et al. RFECS: a random-forest based algorithm for enhancer identification from chromatin state. PLoS Comput. Biol. 9, e1002968 (2013).
Quinlan, A.R. & Hall, I.M. BEDTools: a flexible suite of utilities for comparing genomic features. Bioinformatics 26, 841–842 (2010).
Kelley, M.L., Strezoska, Ž., He, K., Vermeulen, A. & Smith, Av. Versatility of chemically synthesized guide RNAs for CRISPR-Cas9 genome editing. J. Biotechnol. 233, 74–83 (2016).
Heintzman, N.D. et al. Distinct and predictive chromatin signatures of transcriptional promoters and enhancers in the human genome. Nat. Genet. 39, 311–318 (2007).
Diao, Y. et al. Pax3/7BP is a Pax7- and Pax3-binding protein that regulates the proliferation of muscle precursor cells by an epigenetic mechanism. Cell Stem Cell 11, 231–241 (2012).
Zhang, Y. et al. Model-based analysis of ChIP-Seq (MACS). Genome Biol. 9, R137 (2008).
Ramírez, F. et al. deepTools2: a next generation web server for deep-sequencing data analysis. Nucleic Acids Res. 44, W160–W165 (2016).
Acknowledgements
We thank D. Gorkin and J. Yan for feedback on previous versions of the manuscript. We thank Z. Ye and S. Kuan for technical assistance. This work was supported by the US National Institutes of Health (NIH) (grants U54 HG006997, U01 DK105541, R01HG008135, 1UM1HG009402 and 2P50 GM085764 to B.R.), the Ludwig Institute for Cancer Research (to B.R.) and the Human Frontier Science Program (HFSP) (Long Term Postdoctoral Fellowship to Y.D.).
Author information
Authors and Affiliations
Contributions
Y.D. and B.R. conceived the idea for CREST-seq; R.F., Y.D. and B.L. conducted integrative data analysis with help from Y.Q., H.H. and I.J.; B.L. and Y.D. designed paired sgRNA libraries; Y.D., Z.M., J.Y., K.C.L., T.L., H.H., R.J.M. and Y.S. performed the experiment; Z.M., K.C.L. and K.-L.G. packaged the lentiviral library; and Y.D., R.F., B.L. and B.R. wrote the paper.
Corresponding author
Ethics declarations
Competing interests
B.R. is a cofounder of Arima Genomics, Inc.
Integrated supplementary information
Supplementary Figure 1 Design of sgRNA pairs.
(a) A genome browser screenshot illustrating the representative tiling design of CREST-seq sgRNA pairs in the POU5F1 locus. Each black bar represents a sequence targeted by a pair of sgRNAs. (b) Distribution of the sizes of deletions (top panel) and step sizes of two adjacent deletions (bottom panel).
Supplementary Figure 2 CREST-seq library construction and quality control.
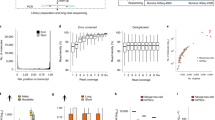
(a) (Top) Schematic illustration of the oligonculeotides containing the pairs of sgRNAs flanked by a common adaptor sequence required for two-step library cloning. (Bottom) Workflow of the two-step plasmid library cloning. The oligo library was synthesized by Custom Array (Seattle, WA), PCR amplified, and cloned into lentiCRISPRv2 backbone via Gibson Assembly. The first step cloning product were then digested by BsmBI and ligated with a DNA fragment containing tracRNA(EF) and mouse U6 promoter (mU6) sequence. tracRNA(EF): tracRNA with extended stem-loops and flipped A/T bases. (b) Lentiviral particles were packaged as described previously 13 and transduced into H1 hESC via spin infection. 36 hours after viral transduction, the cells were cultured in E8 media containing Puromycin for 72 hours, and in regular media for another 3 days. Genomic DNA was purified for genotyping PCR. The PCR products with smaller sizes indicate the genomic deletion at the target region. (c-e) After a two-step cloning procedure and plasmid DNA prep, the dual-sgRNA inserts were amplified from the final CREST-seq plasmid library and subjected to deep sequencing. The paired-end reads were mapped to CREST-seq oligo design file. (c) The plasmid library recovered 96.21% of oligos in the CREST-seq library design. (d, e) Distribution of CREST-seq oligo read counts (d) and cumulative frequency in the plasmid DNA library (e).
Supplementary Figure 3 Quality control of CREST-seq data from replicates.
Genomic DNA isolated from “Cis”, “High” and “Ctrl” cell populations was subjected to PCR amplification and then deep sequencing.
(a) Unsupervised clustering analysis shows correlation of biological replicates of five “Cis” (cis 1-5), three “high” (high 1-3) and two control (ctrl1, ctrl2) samples. (b) Scatter plots show that sgRNA read counts correlate well between replicates. (c) Genome browser screenshot showing the gene annotation in the 2Mbp POU5F1 locus (RefSeq genes), mean reads counts in control samples (“Ctrl”) and in Cis samples (“Cis”), -10log (Adjusted P-value) (green tracks) and log2(Fold change) (blue) of sgRNA pairs. We used edgeR to identify significantly enriched oligos (see Supplementary Methods for more details).
Supplementary Figure 4 CREST-seq identifies the promoter and known enhancers of POU5F1.
(a) Genome browser screenshot showing the CREST-seq peak predicted from “Cis” sample and “High” sample with the same peak calling method (detailed in Material and Methods). (b) Genome browser screenshot showing the CREST-seq peak (top, red bar), CREST-seq signal (dark green track), and the associate features surrounding POU5F1 gene body, promoter and well characterized enhancer (blue bar and the highlighted region by yellow). (c) Genome browser screenshot showing the functional sites identified by CREST-seq (red and green tracks on top) compared to previous single sgRNA based screen (orange bars in the middle, DHS_65, DHS_108, DHS_113 and DHS_115). The black box highlighted the 1Mbp POU5F1 locus surveyed in our previous screen.
Supplementary Figure 5 The POU5F1–eGFP reporter hESC line is highly similar to H1 hESCs.
(a) Genome browser snapshot comparing the ENCODE DHS and CTCF-ChIP-seq signal with POU5F1-eGFP reporter line ATAC-seq and CTCF ChIP-seq signal within the 2Mbp tested POU5F1 locus along with gene annotation. (b) PCA analysis showing the clustering of 122 public available DHS data sets (Supplementary Table 8), including data generated from K562 cell(10x), human lymphoblastoid cell lines(GM, 3x), human fibroblast (Ag, 5x), human dermal microvascular endothelial cells (Hmvec, 8x) and 96 other cell types. ENCODE H1 DHS data and POU5F1-eGFP reporter hESC ATAC-seq data are also included.
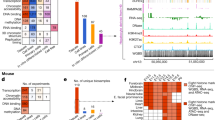
Supplementary Figure 6 Chromatin features enriched on CREs.
(a) A close-up view of a 5kb CRE occupied by a cluster of TFs pointed by green arrowhead in Fig. 2b.
(b) A box plot shows that transcription factor binding sites more frequently cluster at CREs than at typical cis-regulatory elements represented by DHS. (Wilcoxon test P-value < 6e-11). (c) A bar chart shows the degree of enrichment of each chromatin feature in the CREs. To calculate the “Enrichment Test Score”, we first calculated the fraction of CREST-seq peaks that intersected with sites associated with each feature as a ratio between the observed over expected. An average ratio is calculated from 1,000 random permutations of the CREs. The enrichment test score is defined as the percentage that observed ratio is greater than expected. (*χ2 P-value < 0.01). (d) Bar plot shows the enrichment test score for 57 features (49 for TFBS and 8 for histone modifications) at CREs compared to random.
Supplementary Figure 7 Genotype information for the mutant clones with genomic deletion on selected CREs.
(a) Genomic DNA was isolated from each indicated mutant clones and the genotypes were confirmed by Sanger sequencing of genotyping PCR product after TOPO cloning. The targeted deletion regions are showing on top of each panel. The blue box, green box and red box contain the genotyping for bi-allelic, P1 allele or P2 allele deletion, respectively. P1 is the eGFP containing allele while P2 is the allele with wild-type sequence. The genome browser screenshot shows CREST-seq signal/peak, and other epigenetic features as indicated around each targeted locus including CRE(-694), CRE(-652), CRE(-571), CRE(-521), CRE(-449), CRE(+38) and CREST-seq negative region. (b) The eGFP levels on WT cells (WT Ctrl), bia-allelic deletion, P1 allele specific deletion and P2 allele specific deletion mutants was quantified with FlowJo. Both early passage cells (day 25) and long term cultured cells (day 50) were subjected to FACS analysis. Two-sample t-test was performed to compute the P-value, Error bars, s.d.
Supplementary Figure 8 Genotype information for core promoter mutant clones.
The genotype of each mutant clones were determined by genotyping PCR using genomic DNA as template, followed by Sanger sequencing for verification. The blue box, green box and red box highlight the genotyping for bi-allelic, P1 allele or P2 allele deletion, respectively. P1 is the eGFP containing allele while P2 is the allele with wild-type sequence. The genome browser screenshot shows CREST-seq signal/peak, and other epigenetic features as indicated around each targeted locus. From top to bottom: Genotype information of MSH5, NEU1, PRRC2A, and TCF19 core promoter deletion mutants, respectively.
Supplementary Figure 9 Characterization and quantification of eGFP levels in multiple core promoter deletion mutant clones.
(a,b) Total of 37 mutant clones were generated in the same way as described in Fig. 3A. In addition to the 12 mutant clones showing in Fig. 3A, the additional 25 multiple mutant clones were also subjected to FACS analysis at (a) day 25 and (b) day 40 after CRISPR/Cas9 transfection. (c) The FACS data of the mutant clones showing in (A), (B), and Fig. 3A were analyzed with FlowJo to quantify the eGFP level. P-value was calculated with two-sample t-test. Error bars, s.d.
Supplementary Figure 10 Quantification of POU5F1, MSH5, NEU1 and PRRC2A expression in various samples.
The H1 POU5F1-eGFP cells were transfected with either control scrambled siRNA or siRNAs targeting each gene as indicated. Each gene is targeted by two sets of siRNAs (siRNA #1 and #2) with different sequence. 48 hours after transfection, the total RNA was collected from the cells for RT-qPCR analysis. We also packaged lentiviral expressing two sets of shRNAs targeting each gene as indicated (shRNA#1 and shRNA#2). 16 days after lentiviral infection and antibiotic selection (1mg/ml puromycin), the cells were collected for RNA purification followed by qPCR analysis. We also selected some mutant clones with core promoter deletion specified as in Fig. S9C for qPCR analysis. (a-c) RT-qPCR analysis of NEU1, MSH5 and PRRC2A in the samples treated with siRNA, shRNA expressing lentiviral, or deletion on core promoter sequence as indicated. * P-value < 0.01, N.S. not significant, t-test, error bars, s.d. (d) RT-qPCR quantification of POU5F1 mRNA levels in the samples with long-term knockdown of MSH5, NEU1 and PRRC2A. * P-value < 0.01, N.S. not significant, t-test, error bars, s.d. (e) The raw gel image used to make Fig. 3b. (f) Bar chart showing the results from reporter assays testing four different POU5F1-regulatory core promoters. H1 hESC cells were transfected with various luciferase reporter plasmid as indicated. 48 hours post-transfection, cells were lysed and subjected to analysis of luciferase activities. All tested elements are cloned into the downstream of luciferase gene coding sequence in the control reporter (Ctrl) plasmid, which contains the 360bp POU5F1 minimal core promoter sequence to drive reporter gene expression. The reporter activity of each element was compared to the control reporter plasmid containing POU5F1 promoter only. (*t-test: P-value<0.05, error bars, s.d.)
Supplementary Figure 11 The reduced eGFP expression in biallelic and P1-allelic specific mutants is not due to DSB-induced transcription repression.
(a)The mutant clones with bi-allelic deletion (blue curves) or P1 allele deletion (green curves) on targeted CRE sites were dissociated into single cells and stained with PE- or PerCP-Cy5.5- conjugated antibodies specifically recognizing HLA-C or γH2AX, respectively. The black curves represent the signal obtained from WT POU5F1-eGFP reporter cells. Grey curves: WT cells without antibody staining; magenta curve: WT cells treated with 250μM of Etoposide for 6 hours to induce DNA double strand break (positive control for γH2AX staining signal). (b) WT POU5F1-eGFP reporter cells (top) and CRE(+12) biallelic (-/-) mutant (bottom, day 25 after CRISPR/Cas9 transfection) were stained with HLA-C antibody, followed by FACS analysis.
Supplementary Figure 12 Promoter CREs are associated with active gene expression.
(a) A Pie-chart shows that 14 promoter-intersected CREST-seq peaks contain active promoter signatures (Pol2/H3K4me3/H3K27ac). (b) A Bar chart shows that POU5F1-regulating promoters are enriched for active promoter signatures (Pol2/H3K4me3/H3K27ac) compared to random promoters in the region (permutation P-value < 0.01). To estimate the degree of the enrichment, we randomly shuffled 45 CREST-seq peaks within the 2Mbp region and calculated the ratio of peaks that contain active promoter marks (Pol2/H3K4me3/H3K27ac) as expected active promoter ratio. This is repeated for 1,000 times, allowing definition of permutation P-value as the percentage of observations that active-promoter ratio is above an observed ratio (78%) (see Supplementary Methods for more details). (c) A Violin plot shows that transcriptional activities of the POU5F1-regulating promoters are higher than other gene promoters in the 2Mbp region (Wilcoxon P-value < 0.01). We used gene expression profiles from ENCODE previously quantified and normalized using ENCODE uniform pipeline.
Supplementary Figure 13 A list of features that distinguish POU5F1-regulatory promoters from other non-POU5F1-regulatory promoters.
The bar plot reveals the relative importance of each feature to the prediction made by random forest model.
Supplementary Figure 14 Analysis of cis- and trans-regulatory elements with the dual sgRNA tiling deletion screen.
(a) 18 and 20 single clones were randomly picked from the non-sorted control population and the eGFP-/POU5F1+ “Cis” population, respectively. Genomic DNA was isolated followed by PCR amplification of paired sgRNA sequence and Sanger sequencing. After confirming the sgRNA sequence, genotyping PCR was performed to check the sgRNA targeting genomic DNA sequence. (b, c) POU5F1-eGFP reporter cells were infected with control (Ctrl) lentiviral or shRNA targeting PRDM14 and selected with 1mg/ml puromycin for 3 days. At day 5 and day 10 after infection, (b) total RNA was collected and subjected to qPCR analysis to quantify the knockdown effect. * P-value < 0.01, t-test. Error bars, s.d. (c) The cells were dissociated and analyzed by FACS. (d-f) FACS analysis of H1 POU5F1-eGFP cells transduced with CREST-seq lentiviral library (right) 14 days post transduction. The eGFP-/POU5F1- cells (d) and eGFP- cells (f) were collected for further studies. (e) The counts of sgRNA reads from eGFP-/POU5F1+ cells (left, Cis) and eGFP-/POU5F1- (right) are compared to those from a non-sorted control population (Ctrl). The fold changes represent the ratios between the “Cis” or “eGFP-/POU5F1-” sample compared to “Ctrl” sample, with the enrichment significance calculated by negative binomial test using edgeR package. Green dots denote eGFP targeting gRNA pairs; Red dots correspond to positive oligos enriched in the testing population with P-value < 0.05 and log2 (fold change) > 1; blue dots indicate negative control oligos which are enriched with P-value < 0.05 and log2 (fold change) > 1 in the testing samples compared to Ctrl. Grey dots for the rest of sgRNAs. (g) The eGFP- cells were collected, processed and analyzed in the same way as Cis samples. With same peak calling pipeline and cutoff, we identified 45 CREs (blue) and 52 GFP- peaks (orange), with 35 sites overlapped. (h) FACS data showing that 45 CREs contains cis-regulatory elements with strong (red) and weak (blue) effect on POU5F1/eGFP expression while the 52 GFP- sites cover strong cis- and strong trans- elements.
Supplementary Figure 15 The eGFP levels correlate with P1-allele-specific POU5F1 expression.
(a) Schemtic of phasing eGFP (P1) and non-eGFP (P2) alleles of H1 POU5F1-eGFP line. We performed PCR from genomic DNA in the 3' UTR between primer pairs (indicated by black arrows) that would be broken by the inserted transgene, so the only allele that can be amplified is the native one. We then infer what the SNPs on the nontargeted allele are to deduce whether P1 or P2 is the targeted vs. non-targeted allele. (b) Total RNA was purified from WT and promoter-CRE mutant clones followed by qPCR analysis to quantify POU5F1 mRNA levels. * t-test, P-value<0.01, Error bars, s.d.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–15 and Supplementary Notes 1–7 (PDF 2871 kb)
Supplementary Protocol
Supplementary Protocol (PDF 627 kb)
Supplementary Table 1
Oligo sequence of CREST-seq design (XLSX 863 kb)
Supplementary Table 2
CREST-seq oligo read count (XLSX 1968 kb)
Supplementary Table 3
Statistical enrichment of each sgRNA pair in the cis samples compared with the control samples (XLSX 1375 kb)
Supplementary Table 4
Statistical significance of rank bias for each 50-bp genomic bin (XLSX 2766 kb)
Supplementary Table 5
Genomic coordinates of 45 predicted CREST-seq peaks (XLSX 2612 kb)
Supplementary Table 6
Quantification and statistics of FACS eGFP levels in the mutant clones (XLSX 38 kb)
Supplementary Table 7
A full list of genomic features used in machine learning and PCA analysis (XLSX 50 kb)
Supplementary Table 8
List of DNA and CRISPR RNA oligo sequences used in this study (XLSX 12 kb)
Rights and permissions
About this article
Cite this article
Diao, Y., Fang, R., Li, B. et al. A tiling-deletion-based genetic screen for cis-regulatory element identification in mammalian cells. Nat Methods 14, 629–635 (2017). https://doi.org/10.1038/nmeth.4264
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nmeth.4264
This article is cited by
-
Multicenter integrated analysis of noncoding CRISPRi screens
Nature Methods (2024)
-
Cis-regulatory atlas of primary human CD4+ T cells
BMC Genomics (2023)
-
Dynamic network-guided CRISPRi screen identifies CTCF-loop-constrained nonlinear enhancer gene regulatory activity during cell state transitions
Nature Genetics (2023)
-
Young LINE-1 transposon 5′ UTRs marked by elongation factor ELL3 function as enhancers to regulate naïve pluripotency in embryonic stem cells
Nature Cell Biology (2023)
-
Human-specific genetics: new tools to explore the molecular and cellular basis of human evolution
Nature Reviews Genetics (2023)