Abstract
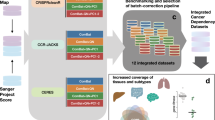
We developed a systematic approach to map human genetic networks by combinatorial CRISPR–Cas9 perturbations coupled to robust analysis of growth kinetics. We targeted all pairs of 73 cancer genes with dual guide RNAs in three cell lines, comprising 141,912 tests of interaction. Numerous therapeutically relevant interactions were identified, and these patterns replicated with combinatorial drugs at 75% precision. From these results, we anticipate that cellular context will be critical to synthetic-lethal therapies.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
References
Mani, R., St Onge, R.P., Hartman, J.L. IV, Giaever, G. & Roth, F.P. Proc. Natl. Acad. Sci. USA 105, 3461–3466 (2008).
Avery, L. & Wasserman, S. Trends Genet. 8, 312–316 (1992).
Collins, S.R. et al. Nature 446, 806–810 (2007).
Bandyopadhyay, S. et al. Science 330, 1385–1389 (2010).
Costanzo, M. et al. Science 353, af1420 (2016).
Lord, C.J., Tutt, A.N. & Ashworth, A. Annu. Rev. Med. 66, 455–470 (2015).
Krogan, N.J., Lippman, S., Agard, D.A., Ashworth, A. & Ideker, T. Mol. Cell 58, 690–698 (2015).
Srivas, R. et al. Mol. Cell 63, 514–525 (2016).
Mali, P., Esvelt, K.M. & Church, G.M. Nat. Methods 10, 957–963 (2013).
Gilbert, L.A. et al. Cell 159, 647–661 (2014).
Wong, A.S.L. et al. Proc. Natl. Acad. Sci. USA 113, 2544–2549 (2016).
Bassik, M.C. et al. Cell 152, 909–922 (2013).
Horn, T. et al. Nat. Methods 8, 341–346 (2011).
Laufer, C., Fischer, B., Billmann, M., Huber, W. & Boutros, M. Nat. Methods 10, 427–431 (2013).
Vogelstein, B. et al. Science 339, 1546–1558 (2013).
Neshat, M.S. et al. Proc. Natl. Acad. Sci. USA 98, 10314–10319 (2001).
Chari, R., Mali, P., Moosburner, M. & Church, G.M. Nat. Methods 12, 823–826 (2015).
Doench, J.G. et al. Nat. Biotechnol. 34, 184–191 (2016).
Xu, H. et al. Genome Res. 25, 1147–1157 (2015).
Shi, J. et al. Nat. Biotechnol. 33, 661–667 (2015).
Shen, J.P., Zhao, D., Sasik, R., Ideker, T. & Mali, P. Combinatorial CRISPR-Cas9 knockout screen. Protocol Exchange http://dx.doi.org/10.1038/protex.2017.063 (2017).
Kabadi, A.M., Ousterout, D.G., Hilton, I.B. & Gersbach, C.A. Nucleic Acids Res. 42, e147 (2014).
Briner, A.E. et al. Mol. Cell 56, 333–339 (2014).
Dang, Y. et al. Genome Biol. 16, 280 (2015).
Mali, P. et al. Nat. Biotechnol. 31, 833–838 (2013).
Sanjana, N.E., Shalem, O. & Zhang, F. Nat. Methods 11, 783–784 (2014).
Martin, M. EMBnet.journal 17, 10–12 (2011).
Baryshnikova, A., Costanzo, M., Myers, C.L., Andrews, B. & Boone, C. Annu. Rev. Genomics Hum. Genet. 14, 111–133 (2013).
Hart, T. et al. Cell 163, 1515–1526 (2015).
Storey, J.D. Ann. Stat. 31, 2013–2035 (2003).
Storey, J.D., Bass, A.J., Dabney, A. & Robinson, D. qvalue: Q-value estimation for false discovery rate control. R package version 2.4.2. (2015).
Benjamini, Y. & Hochberg, Y. J. R. Stat. Soc. B 57, 289–300 (1995).
Tusher, V.G., Tibshirani, R. & Chu, G. Proc. Natl. Acad. Sci. USA 98, 5116–5121 (2001).
Pollard, K.S. & van der Laan, M.J. J. Stat. Plan. Inference 125, 85–100 (2004).
Baryshnikova, A. et al. Nat. Methods 7, 1017–1024 (2010).
ENCODE Project Consortium. Nature 489, 57–74 (2012).
Lori, F. & Lisziewicz, J. Clin. Infect. Dis. 30 (Suppl. 2), S193–S197 (2000).
DeLean, A., Munson, P. & Rodbard, D. Am. J. Physiol. Gastrointest. Liver Physiol. 235, G97–G102 (1978).
Acknowledgements
This work was generously supported by the following sources: UC San Diego School of Engineering and School of Medicine institutional funds (P.M., T.I., R.S. and A.B.), the Burroughs Wellcome Fund (1013926 to P.M.), the March of Dimes Foundation (5-FY15-450 to P.M.), the Sidney Kimmel Foundation (SKF-16-150 to P.M.), the California Institute for Regenerative Medicine (T.I. and J.P.S.), the National Cancer Institute (R21 CA199292 to N.E.L. and L30 CA171000 to J.P.S.), the National Institute for Environmental Health Sciences (R01 ES014811 to T.I.), the National Institute for General Medical Sciences (T32 GM008806 to J.L., R01 GM084279 to T.I. and N.K. and P50 GM085764 to T.I.), the UC San Diego Clinical and Translational Research Institute Grant (UL1TR001442 to R.S. and A.B.) and the Novo Nordisk Foundation Center for Biosustainability at DTU (NNF16CC0021858 to N.E.L.).
Author information
Authors and Affiliations
Contributions
J.P.S., D.Z., T.I. and P.M. conceived and supervised all experiments and wrote the paper. R.S. and J.L. performed computational analysis and wrote the paper. A.B., C.-C.K., A.T., N.L. and A.C. performed computational analysis. J.P.S., D.Z., A.B.G., K.L., K.K., D.P., A.N.B., K.S.S. and P.M. performed experiments. D.D., A.R., N.K. and L.Q. provided technical advice. J.K. provided technical advice and wrote the paper.
Corresponding authors
Ethics declarations
Competing interests
J.P.S., D.Z., R.S., T.I. and P.M. have filed a patent based on the described methods and findings; otherwise, the authors declare no competing financial interests.
Integrated supplementary information
Supplementary Figure 1 Schematic of cloning strategy for dual-gRNA-library synthesis.
Preparation of the dual-gRNA library involves a two-step cloning process whereby each synthesized oligonucleotide is assembled progressively with promoters and 3′ gRNA scaffolds.
Supplementary Figure 2 Evaluation of activity of engineered gRNA scaffolds.
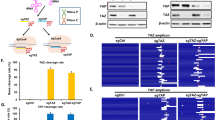
(a) A range of mutagenized gRNA scaffold sequences (designed to increase sequence diversity but maintain the primary hairpin loops in the gRNA scaffold via utilization of G-C vs. A-U interactions) were evaluated. (b) To assay their relative activity, NHEJ mediated mutagenesis of the AAVS1 locus was performed in HEK 293T cells and editing rates quantified 1-week post transduction of gRNA constructs. Two adjacent targets T1 and T2 (sequences labeled respectively in red and green) were targeted via corresponding gRNAs. Representative next generation sequencing results are shown, bar plots indicate ratio of reads showing indel relative to total reads. These experiments indicated that the wild-type scaffold (version 1) and an engineered scaffold (version 4) were the most active and showed the most consistent activity. (c) The effect of position of a gRNA in a dual-gRNA construct was evaluated via its ability to induce NHEJ mediated mutagenesis of EGFP loci in a HEK 293T cell line stably expressing the protein. In these experiments loss of GFP expression was assayed 96 hours post transduction of dual-gRNA constructs. The percentage of GFP+ cells are indicated in the FACS panel showing side scatter (SSC) vs. GFP signal. Bar graph shows average with error bar representing +/- SD from three technical replicates. These experiments confirmed that the two gRNA positions in the targeting construct were equally functional (left: hU6-gRNA; right: mU6-gRNA). (d) Similarly the simultaneous activity of two gRNAs in the same construct was evaluated by measuring loss of EGFR and mCherry expression in a HEK 293T cell line stably expressing both proteins. The percentage GFP+ and mCherry+ cells are indicated in each bar graph, error bar represents +/- SD from three technical replicates. Below is representative FACS panel showing mCherry vs. EGFP.
Supplementary Figure 3 Multiple time-point regression with Bayesian sampling analysis method.
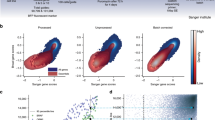
(a) Sample histograms of log2 relative frequencies of dual-gRNA constructs for HeLa cells. Red lines denote abundance cutoffs below which the data is deemed under-sampled and dropped. Note that after 28 days of selection there is a broader distribution with more gRNA falling under the threshold. (b) Representative data for fitting of a few selected gRNA constructs for HeLa cells. Black and blue points and lines correspond to data points and continuous fits for replicates 1 and 2, respectively. Empty symbols denote data points below the abundance threshold, which are not used for fitting. The slight curvature to the fitted lines is due to the nonlinear term in Eq. 3. (c) Comparison of unique gRNA probes for each gene for HeLa cells. The three gRNA probes targeting each gene are not equally active, however the fitness effect of each gRNA was highly correlated between replicates. Here the three gRNA for PTEN, VHL and the non-targeting are highlighted, showing that only two of the three gRNA have an effect greater than that of the non-targeting gRNA probes. These observations motivated our efforts to rank individual probes by activity and accordingly weight their effects in the overall analyses. (d) Kernel density plot for CRISPR-Cas9 screen probe ranks across screened genes for both HeLa and A549 cell lines. Red line indicates orthogonal regression result. Probe ranks assigned to each gene in both screens correlate significantly (Pearson r = 0.42, p = 5.3 ×10-11).
Supplementary Figure 4 Replicate analyses and cell-line comparisons.
Scatterplot of fitness (fg) for single gene knockouts for biological replicates in (a) HeLa, (b) A549, and (c) 293T cells. Scatterplot of raw genetic interaction scores (π gg') for biological replicates in (d) HeLa, (e) A549, and (f) 293T cells. The significant interactions underlying the correlation calculation are highlighted in bold. (g) Genes that have large fitness effects when they are disrupted have greater number of genetic interactions (node degree, includes positive interactions as well as synthetic lethal interactions). This was true for Hela (blue), A549 (pink) and 293T (black) cells. Interaction hubs (genes with highest node degree) are highlighted for each cell line. (h) Scatterplot of fitness (fg) for single gene knockouts in HeLa versus 293T. (i) Scatterplot of fitness (fg) for single gene knockouts in A549 versus 293T. (j) Scatterplot of interaction scores in HeLa versus 293T. (k) Scatterplot of interaction scores in A549 versus 293T.
Supplementary Figure 5 Single-gene fitness effects relative to gene expression.
Scatterplot of expression vs. single gene knockout fitness effect (fg) for (a) HeLa, (b) A549, and (c) 293T cells. Pearson correlation r values and p values are indicated.
Supplementary Figure 6 Single-cell-line synthetic-lethal networks.
Z-score vs. False Discovery Rate (FDR) plots for (a) HeLa, (b) A549, and (c) 293T cells. Synthetic lethal networks for (d) HeLa, (e) A549, and (f) 293T cells. Circles indicate TSG, squares druggable targets, squares outlined in bold indicate FDA approval of drug targeting that gene. Color of node indicates single gene knockout fitness effect, red: positive fitness effect, blue: negative fitness effect. Previously reported interactions are indicated as red edges, and interactions validated in drug-drug assays in this study are shown in green. Black edges were identified in multiple cell lines in this study.
Supplementary Figure 7 Selection of ‘drug 2’ concentrations.
Single drug dose-response curves are shown for all drugs used as ‘drug2’ in a drug1-drug2 assay described in the manuscript: (a) trametinib (b) 5-FU (c) fludarabine (d) KU-60019 (e) dinaciclib (f) hydroxyurea (g) NU7441. X-axis is log10 of the concentration of the indicated drug in micromolar units. In drug1-drug2 assays, concentration of drug2 was set at the IC20 value (dotted line). Viability normalized to solvent-only controls independently for HeLa and A549 cells. Error bars represent +/- SD.
Supplementary Figure 8 Validation of synthetic-lethal interactions in drug–drug assays.
Dose-response curves for inhibitors of the following gene pairs (a) CHEK1-MAP2K1 (b) CHEK1-TYMS (c) ADA-CHEK1 (d) ATM-CHEK1 (e) CDK9-CHEK1 (f) PRKDC-RRM2 (g) CDK9-PRKDC (h) CDK4-PRKDC; HeLa on left and A549 on right of each pair; error bars represent +/- SD. X-axis is log10 of the concentration of the indicated drug in micromolar units. Gray curves show single agent dose-response of Drug#1 and are normalized to solvent control. Pink curves show dose response in presence of fixed concentration of Drug#2 and are normalized to effect of Drug#2 at indicated dose. To assess for synthetic lethal interaction, the IC50 values for gray and pink curves are compared for each plot using the sum-of-squares F test; visually the IC50 concentration is the point along the x-axis at which the curve crosses 50% viability (dashed line). When the pink curve crosses the 50% viability to the left of the gray this indicates a synergistic, or synthetic lethal interaction; p values are indicated with significant interactions (p < 0.05) highlighted in red. (i) Summary of synthetic-lethal interactions re-tested in arrayed drug-drug viability assays. (j) Matrix summarizing results from drug-drug validation assays for HeLa and A549 cell lines. A total of 8 gene pairs were each tested in both HeLa and A549 cells for a total of 16 tests.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–8 (PDF 1653 kb)
Supplementary Table 1
Annotated list of the 73 genes assayed in this study (XLSX 15 kb)
Supplementary Table 2
Raw and processed data from CRISPR competitive growth assay (XLSX 3612 kb)
Supplementary Table 3
List of hit genetic interactions from CRISPR assay (XLSX 38 kb)
Supplementary Table 4
Summary of drug-drug validation experiments (XLSX 14 kb)
Rights and permissions
About this article
Cite this article
Shen, J., Zhao, D., Sasik, R. et al. Combinatorial CRISPR–Cas9 screens for de novo mapping of genetic interactions. Nat Methods 14, 573–576 (2017). https://doi.org/10.1038/nmeth.4225
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nmeth.4225
This article is cited by
-
X-CHIME enables combinatorial, inducible, lineage-specific and sequential knockout of genes in the immune system
Nature Immunology (2024)
-
Efficient gene knockout and genetic interaction screening using the in4mer CRISPR/Cas12a multiplex knockout platform
Nature Communications (2024)
-
Recent advances and applications of CRISPR-Cas9 in cancer immunotherapy
Molecular Cancer (2023)
-
CRISPR screening in hematology research: from bulk to single-cell level
Journal of Hematology & Oncology (2023)
-
CRISPR-Combo–mediated orthogonal genome editing and transcriptional activation for plant breeding
Nature Protocols (2023)