Abstract
Resolving the DNA-binding specificities of transcription factors (TFs) is of critical value for understanding gene regulation. Here, we present a novel, semiautomated protein–DNA interaction characterization technology, selective microfluidics-based ligand enrichment followed by sequencing (SMiLE-seq). SMiLE-seq is neither limited by DNA bait length nor biased toward strong affinity binders; it probes the DNA-binding properties of TFs over a wide affinity range in a fast and cost-effective fashion. We validated SMiLE-seq by analyzing 58 full-length human, mouse, and Drosophila TFs from distinct structural classes. All tested TFs yielded DNA-binding models with predictive power comparable to or greater than that of other in vitro assays. De novo motif discovery on all JUN–FOS heterodimers and several nuclear receptor-TF complexes provided novel insights into partner-specific heterodimer DNA-binding preferences. We also successfully analyzed the DNA-binding properties of uncharacterized human C2H2 zinc-finger proteins and validated several using ChIP-exo.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Matys, V. et al. TRANSFAC and its module TRANSCompel: transcriptional gene regulation in eukaryotes. Nucleic Acids Res. 34, D108–D110 (2006).
Mathelier, A. et al. JASPAR 2014: an extensively expanded and updated open-access database of transcription factor binding profiles. Nucleic Acids Res. 42, D142–D147 (2014).
Newburger, D.E. & Bulyk, M.L. UniPROBE: an online database of protein binding microarray data on protein-DNA interactions. Nucleic Acids Res. 37, D77–D82 (2009).
Kulakovskiy, I.V. et al. HOCOMOCO: expansion and enhancement of the collection of transcription factor binding sites models. Nucleic Acids Res. 44 D1, D116–D125 (2016).
Fulton, D.L. et al. TFCat: the curated catalog of mouse and human transcription factors. Genome Biol. 10, R29 (2009).
Vaquerizas, J.M., Kummerfeld, S.K., Teichmann, S.A. & Luscombe, N.M. A census of human transcription factors: function, expression and evolution. Nat. Rev. Genet. 10, 252–263 (2009).
Berger, M.F. & Bulyk, M.L. Universal protein-binding microarrays for the comprehensive characterization of the DNA-binding specificities of transcription factors. Nat. Protoc. 4, 393–411 (2009).
Meng, X., Brodsky, M.H. & Wolfe, S.A. A bacterial one-hybrid system for determining the DNA-binding specificity of transcription factors. Nat. Biotechnol. 23, 988–994 (2005).
Jolma, A. et al. Multiplexed massively parallel SELEX for characterization of human transcription factor binding specificities. Genome Res. 20, 861–873 (2010).
Deplancke, B., Alpern, D. & Gardeux, V. The genetics of transcription factor DNA binding variation. Cell 166, 538–554 (2016).
Ravasi, T. et al. An atlas of combinatorial transcriptional regulation in mouse and man. Cell 140, 744–752 (2010).
Jolma, A. et al. DNA-dependent formation of transcription factor pairs alters their binding specificity. Nature 527, 384–388 (2015).
O'Shea, E.K., Rutkowski, R. & Kim, P.S. Mechanism of specificity in the Fos-Jun oncoprotein heterodimer. Cell 68, 699–708 (1992).
Isakova, A., Berset, Y., Hatzimanikatis, V. & Deplancke, B. Quantification of cooperativity in heterodimer-DNA binding improves the accuracy of binding specificity models. J. Biol. Chem. 291, 10293–10306 (2016).
Rastinejad, F., Ollendorff, V. & Polikarpov, I. Nuclear receptor full-length architectures: confronting myth and illusion with high resolution. Trends Biochem. Sci. 40, 16–24 (2015).
Weirauch, M.T. et al. Evaluation of methods for modeling transcription factor sequence specificity. Nat. Biotechnol. 31, 126–134 (2013).
Maerkl, S.J. & Quake, S.R. A systems approach to measuring the binding energy landscapes of transcription factors. Science 315, 233–237 (2007).
Zimmermann, M., Hunziker, P. & Delamarche, E. Valves for autonomous capillary systems. Microfluid. Nanofluidics 5, 395–402 (2008).
Gupta, S., Stamatoyannopoulos, J.A., Bailey, T.L. & Noble, W.S. Quantifying similarity between motifs. Genome Biol. 8, R24 (2007).
Noyes, M.B. et al. A systematic characterization of factors that regulate Drosophila segmentation via a bacterial one-hybrid system. Nucleic Acids Res. 36, 2547–2560 (2008).
Jolma, A. et al. DNA-binding specificities of human transcription factors. Cell 152, 327–339 (2013).
Orenstein, Y. & Shamir, R. A comparative analysis of transcription factor binding models learned from PBM, HT-SELEX and ChIP data. Nucleic Acids Res. 42, e63 (2014).
ENCODE Project Consortium. An integrated encyclopedia of DNA elements in the human genome. Nature 489, 57–74 (2012).
Grant, C.E., Bailey, T.L. & Noble, W.S. FIMO: scanning for occurrences of a given motif. Bioinformatics 27, 1017–1018 (2011).
Geertz, M., Shore, D. & Maerkl, S.J. Massively parallel measurements of molecular interaction kinetics on a microfluidic platform. Proc. Natl. Acad. Sci. USA 109, 16540–16545 (2012).
Nielsen, R. et al. Genome-wide profiling of PPARgamma:RXR and RNA polymerase II occupancy reveals temporal activation of distinct metabolic pathways and changes in RXR dimer composition during adipogenesis. Genes Dev. 22, 2953–2967 (2008).
Taylor, P. & Hardin, P.E. Rhythmic E-box binding by CLK-CYC controls daily cycles in per and tim transcription and chromatin modifications. Mol. Cell. Biol. 28, 4642–4652 (2008).
Rey, G. et al. Genome-wide and phase-specific DNA-binding rhythms of BMAL1 control circadian output functions in mouse liver. PLoS Biol. 9, e1000595 (2011).
Glass, C.K. Differential recognition of target genes by nuclear receptor monomers, dimers, and heterodimers. Endocr. Rev. 15, 391–407 (1994).
Evans, R.M. & Mangelsdorf, D.J. Nuclear receptors, RXR, and the Big Bang. Cell 157, 255–266 (2014).
Shaulian, E. & Karin, M. AP-1 as a regulator of cell life and death. Nat. Cell Biol. 4, E131–E136 (2002).
Eferl, R. & Wagner, E.F. AP-1: a double-edged sword in tumorigenesis. Nat. Rev. Cancer 3, 859–868 (2003).
Ryseck, R.P. & Bravo, R. c-JUN, JUN B, and JUN D differ in their binding affinities to AP-1 and CRE consensus sequences: effect of FOS proteins. Oncogene 6, 533–542 (1991).
Gustems, M. et al. c-Jun/c-Fos heterodimers regulate cellular genes via a newly identified class of methylated DNA sequence motifs. Nucleic Acids Res. 42, 3059–3072 (2014).
Monje, P., Hernández-Losa, J., Lyons, R.J., Castellone, M.D. & Gutkind, J.S. Regulation of the transcriptional activity of c-Fos by ERK. A novel role for the prolyl isomerase PIN1. J. Biol. Chem. 280, 35081–35084 (2005).
Basuyaux, J.P., Ferreira, E., Stéhelin, D. & Butticè, G. The Ets transcription factors interact with each other and with the c-Fos/c-Jun complex via distinct protein domains in a DNA-dependent and -independent manner. J. Biol. Chem. 272, 26188–26195 (1997).
Persikov, A.V. et al. A systematic survey of the Cys2His2 zinc finger DNA-binding landscape. Nucleic Acids Res. 43, 1965–1984 (2015).
Najafabadi, H.S. et al. C2H2 zinc finger proteins greatly expand the human regulatory lexicon. Nat. Biotechnol. 33, 555–562 (2015).
Weirauch, M.T. et al. Determination and inference of eukaryotic transcription factor sequence specificity. Cell 158, 1431–1443 (2014).
Christensen, R.G. et al. A modified bacterial one-hybrid system yields improved quantitative models of transcription factor specificity. Nucleic Acids Res. 39, e83 (2011).
Gupta, A. et al. An improved predictive recognition model for Cys(2)-His(2) zinc finger proteins. Nucleic Acids Res. 42, 4800–4812 (2014).
Isakova, A., Groux, R., Ambrosini, G., Bucher, P. & Deplancke, B. SMiLE-seq: Selective Microfluidics-based Ligand Enrichment followed by sequencing. Protoc. Exch. 10.1038/protex.2016.089.
Zimmermann, M., Schmid, H., Hunziker, P. & Delamarche, E. Capillary pumps for autonomous capillary systems. Lab Chip 7, 119–125 (2007).
Thorsen, T., Maerkl, S.J. & Quake, S.R. Microfluidic large-scale integration. Science 298, 580–584 (2002).
Bailey, T.L. & Elkan, C. In Proc. Int. Conf. Intell. Syst. Mol. Biol. (Eds. Altman, R. et al.) 28–36 (AAAI Press, 1994).
Schütz, F. & Delorenzi, M. MAMOT: hidden Markov modeling tool. Bioinformatics 24, 1399–1400 (2008).
Hume, M.A., Barrera, L.A., Gisselbrecht, S.S. & Bulyk, M.L. UniPROBE, update 2015: new tools and content for the online database of protein-binding microarray data on protein-DNA interactions. Nucleic Acids Res. 43, D117–D122 (2015).
Barde, I., Salmon, P. & Trono, D. Production and titration of lentiviral vectors. Current Protoc. Neurosci. 53, 4.21.1 . (2010).
Serandour, A.A., Brown, G.D., Cohen, J.D. & Carroll, J.S. Development of an Illumina-based ChIP-exonuclease method provides insight into FoxA1-DNA binding properties. Genome Biol. 14, R147 (2013).
Acknowledgements
We would like to thank S. Maerkl (EPFL) for his guidance in applying microfluidic technologies; R. Dreos (EPFL) for helpful discussions on data analysis; and our lab members P. Schwalie and V. Gardeux (EPFL) for providing feedback on the manuscript. We also thank K. Harshman and B. Mangeat for their assistance in sample sequencing, as well as the VITAL-IT for providing the infrastructure for our computational analyses. This work has been supported by funds from the Swiss National Science Foundation (grant nos. 31003A_162735 and CRSII3_147684), by SystemsX.ch Special Opportunity Project 2015/323, and by institutional support from the EPFL.
Author information
Authors and Affiliations
Contributions
A.I. and B.D. conceived and planned the study and prepared the manuscript. A.I. performed the SMiLE-seq experiments. A.I. and R.G. analyzed SMiLE-seq data. P.R., D.A., and R.D. performed validation experiments including ChIP-seq. M.I. and D.T. performed ChIP-exo. R.G., G.A., and P.B. developed and implemented new bioinformatics methods and performed web server setup. All the authors discussed the results and commented on the paper.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
Supplementary Figure 1 SMiLE-seq set-up.
Top right. SMiLE-seq set-up. Each SMiLE-seq device consists of a PDMS chip (approximately 2 x 5 cm) bonded to a plasma-activated glass slide. The SMiLE-seq device is placed on the microscope table and is connected to the microcontroller-based control unit. The microscope camera, connected to an external display, enables chip observation during a SMiLE-seq experiment. Center. Schematic design of a SMiLE-seq microchip. Blue and green colors denote flow and control layers respectively. Each unit of the device is connected to the collector unit on one side and the capillary pump on the other1. All units of the device are connected together by the continuous flow channel with four inlets (F1-F4) and three outlets (F5-F7). Switching between these two access modes can be done through the use of control micro valves (C1-C11).
1. Zimmermann, M., Hunziker, P. & Delamarche, E. Valves for autonomous capillary systems. Microfluid. Nanofluidics 5, 395–402 (2008).
Supplementary Figure 2 SMiLE-seq capacity and reproducibility.
a and b. Motifs for mouse (a) and Drosophila (b) TFs. c-f. Scatter plots showing enrichment of top 2000 k-mers, from two independent SMiLE-seq experiments for PAX7 (c), SRY (d), MAX (e) and FLI1 (f) TFs. rp denotes for Pearson correlation coefficient.
Supplementary Figure 3 AUC profiles of TF binding predicted by SMiLE-seq models.
Each plot represents the AUC value computed for SMiLE-seq, HT-SELEX, JASPAR, UniPROBE (if available) and HOCOMOCO DNA binding models on intervals of 500 peaks obtained from ranked (from high-to-low) ENCODE ChIP-seq peak data.
Supplementary Figure 4 The predictive power of SMiLE-seq data.
a. The predictive power of SMiLE-seq motifs compared to the motifs that are retrievable from HT-SELEX data or computed from HT-SELEX data cycle 1 using the HMM-based analysis pipeline. For each motif, we computed area under the ROC curve (AUC) values on the 500 top peaks of the ENCODE ChIP-seq datasets for a given TF. The heat map represents the AUC values computed for SMiLE-seq, HT-SELEX and HT-SELEX cycle1 motifs on the respective ChIP-seq datasets that were selected based on the highest mean AUC values among all five models. b. Each box plot represents the AUC value computed for SMiLE-seq, HT-SELEX, JASPAR and HOCOMOCO DNA binding models on a 500bp peak interval obtained from ranked (from high-to-low) ENCODE ChIP-seq data. c-f. Egr1 binding affinity. (c) Correlation between the k-mer enrichment of all possible SNP variants of the GCGTGGGCG 9-mer data derived from either the SMiLE-seq experiment or different selection cycles of HT-SELEX (SRA ID: ERR185027 for cycle 2, ERR185028 for cycle 3 and ERR185029 for cycle 4) and corresponding binding affinities computed from Kd values2 of the Egr1 mouse TF. (d) Same, but the binding affinities of 9-mers computed from Kon/Koff values. (e-f). Correlation between normalized PBM (UniPROBE Accession Number: UP00007) 9-mer counts of all possible GCGTGGGCG SNP variants as well as the respective 9-mer SMiLE-seq counts and corresponding binding affinity values of Egr1 TF computed either from Kds (e) or Kon/Koff values (f). rp and rs denote Pearson and Spearman correlation coefficients respectively.
2. Geertz, M., Shore, D. & Maerkl, S. J. Massively parallel measurements of molecular interaction kinetics on a microfluidic platform. Proc. Natl. Acad. Sci. U. S. A. 109, 16540–16545 (2012).
Supplementary Figure 5 Identification of binding motifs for TF heterodimers using SMiLE-seq.
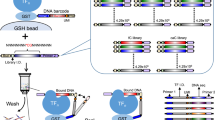
a. Schematic representation of the experimental setup. Step 1. Biotinylated anti-eGFP antibody is immobilized under the button of the SMiLE-seq device. Step 2. Dimerizing transcription factor (TF1) fused to an eGFP tag, dimer partner (TF2) tagged with mCherry and Cy5-labeled DNA baits are introduced into the chip. Step 3. Antibody-immobilized complexes consisting of TF1, TF2, and DNA are trapped under the flexible PDMS membrane; dimer formation is confirmed by fluorescent read-out. Step 4. Unbound molecules as well as molecular complexes are washed away. b. TOMTOM3 comparison of JASPAR and SMiLE-seq binding motifs for mouse PPARγ:RXRα and human ARNTL:CLOCK heterodimers.
3. Gupta, S., Stamatoyannopoulos, J. A., Bailey, T. L. & Noble, W. Quantifying similarity between motifs. Genome Biol. 8, R24 (2007).
Supplementary Figure 6 JUN:FOS motifs.
Primary (top) and secondary (bottom) motifs identified for JUN:FOS heterodimers.
Supplementary Figure 7 Genomic regions bound by KRAB ZFPs.
Peak annotation of the genomic regions bound by ZFP14 (a), ZNF135 (b), ZNF682 (c) obtained from HOMER4 and GREAT5 analyses.
4. Heinz, S. et al. Simple combinations of lineage-determining transcription factors prime cis-regulatory elements required for macrophage and B cell identities. Mol. Cell 38, 576–589 (2010).
5. McLean, C. Y. et al. GREAT improves functional interpretation of cis-regulatory regions. Nat. Biotechnol. 28, 495–501 (2010).
Supplementary Figure 8 An example of an initial HMM with a seed sequence 'ATGCCC'.
The emission states in the boxes correspond to 'A', 'C', 'G' and 'T' respectively. The red values are the values that are not subjected to EM training.
Supplementary information
Supplementary Text and Figures
Supplementary Figure 1–8 and Supplementary Tables 3–6 (PDF 1823 kb)
Supplementary Table 1
TFs used in the study. (XLSX 107 kb)
Supplementary Table 2
AUC values computed for SMiLE-seq, HTSELEX, JASPAR, HOCOMOCO and UniPROBE models on ChIP-seq peak intervals. (XLSX 132 kb)
Supplementary Data
SMiLE-seq-derived PWMs (ZIP 36 kb)
Rights and permissions
About this article
Cite this article
Isakova, A., Groux, R., Imbeault, M. et al. SMiLE-seq identifies binding motifs of single and dimeric transcription factors. Nat Methods 14, 316–322 (2017). https://doi.org/10.1038/nmeth.4143
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nmeth.4143
This article is cited by
-
ExplaiNN: interpretable and transparent neural networks for genomics
Genome Biology (2023)
-
PAPerFly: Partial Assembly-based Peak Finder for ab initio binding site reconstruction
BMC Bioinformatics (2023)
-
Hidden modes of DNA binding by human nuclear receptors
Nature Communications (2023)
-
Double DAP-seq uncovered synergistic DNA binding of interacting bZIP transcription factors
Nature Communications (2023)
-
Modeling binding specificities of transcription factor pairs with random forests
BMC Bioinformatics (2022)