Abstract
We present an approach that enables us to simultaneously access structure and dynamics of a multidomain protein in solution. Dynamic domain arrangements are experimentally determined by combining self-consistent networks of distance distributions with known domain structures. Local structural dynamics are correlated with the global arrangements by analyzing networks of time-resolved single-molecule fluorescence parameters. The strength of this hybrid approach is shown by an application to the flexible multidomain protein Hsp90. The average solution structure of Hsp90's closed state resembles the known X-ray crystal structure with Angstrom precision. The open state is represented by an ensemble of conformations with interdomain fluctuations of up to 25 Å. The data reveal a state-specific suppression of the submillisecond fluctuations by dynamic protein–protein interaction. Finally, the method enables localization and functional characterization of dynamic elements and domain interfaces.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Henzler-Wildman, K. & Kern, D. Dynamic personalities of proteins. Nature 450, 964–972 (2007).
Krukenberg, K.A., Street, T.O., Lavery, L.A. & Agard, D.A. Conformational dynamics of the molecular chaperone Hsp90. Q. Rev. Biophys. 44, 229–255 (2011).
Pearl, L.H. & Prodromou, C. Structure and mechanism of the Hsp90 molecular chaperone machinery. Annu. Rev. Biochem. 75, 271–294 (2006).
Taipale, M. et al. A quantitative chaperone interaction network reveals the architecture of cellular protein homeostasis pathways. Cell 158, 434–448 (2014).
Neckers, L. & Workman, P. Hsp90 molecular chaperone inhibitors: are we there yet? Clin. Cancer Res. 18, 64–76 (2012).
Porter, J.R., Fritz, C.C. & Depew, K.M. Discovery and development of Hsp90 inhibitors: a promising pathway for cancer therapy. Curr. Opin. Chem. Biol. 14, 412–420 (2010).
Rohner, N. et al. Cryptic variation in morphological evolution: HSP90 as a capacitor for loss of eyes in cavefish. Science 342, 1372–1375 (2013).
Mickler, M., Hessling, M., Ratzke, C., Buchner, J. & Hugel, T. The large conformational changes of Hsp90 are only weakly coupled to ATP hydrolysis. Nat. Struct. Mol. Biol. 16, 281–286 (2009).
Ali, M.M. et al. Crystal structure of an Hsp90-nucleotide-p23/Sba1 closed chaperone complex. Nature 440, 1013–1017 (2006).
Sidera, K. & Patsavoudi, E. HSP90 inhibitors: current development and potential in cancer therapy. Recent Patents Anticancer. Drug Discov. 9, 1–20 (2014).
Krukenberg, K.A., Förster, F., Rice, L.M., Sali, A. & Agard, D.A. Multiple conformations of E. coli Hsp90 in solution: insights into the conformational dynamics of Hsp90. Structure 16, 755–765 (2008).
Jahn, M. et al. The charged linker of the molecular chaperone Hsp90 modulates domain contacts and biological function. Proc. Natl. Acad. Sci. USA 111, 17881–17886 (2014).
Comstock, M.J. et al. Protein structure. Direct observation of structure-function relationship in a nucleic acid-processing enzyme. Science 348, 352–354 (2015).
Schuler, B. Single-molecule FRET of protein structure and dynamics—a primer. J. Nanobiotechnology 11 (Suppl. 1), S2 (2013).
Ha, T. et al. Probing the interaction between two single molecules: fluorescence resonance energy transfer between a single donor and a single acceptor. Proc. Natl. Acad. Sci. USA 93, 6264–6268 (1996).
Hohlbein, J. et al. Conformational landscapes of DNA polymerase I and mutator derivatives establish fidelity checkpoints for nucleotide insertion. Nat. Commun. 4, 2131 (2013).
Mashaghi, A., Kramer, G., Lamb, D.C., Mayer, M.P. & Tans, S.J. Chaperone action at the single-molecule level. Chem. Rev. 114, 660–676 (2014).
Kalinin, S. et al. A toolkit and benchmark study for FRET-restrained high-precision structural modeling. Nat. Methods 9, 1218–1225 (2012).
Muschielok, A. et al. A nano-positioning system for macromolecular structural analysis. Nat. Methods 5, 965–971 (2008).
Brunger, A.T., Strop, P., Vrljic, M., Chu, S. & Weninger, K.R. Three-dimensional molecular modeling with single molecule FRET. J. Struct. Biol. 173, 497–505 (2011).
Prodromou, C. et al. Structural basis of the radicicol resistance displayed by a fungal hsp90. ACS Chem. Biol. 4, 289–297 (2009).
Lee, N.K. et al. Accurate FRET measurements within single diffusing biomolecules using alternating-laser excitation. Biophys. J. 88, 2939–2953 (2005).
Shiau, A.K., Harris, S.F., Southworth, D.R. & Agard, D.A. Structural Analysis of E. coli hsp90 reveals dramatic nucleotide-dependent conformational rearrangements. Cell 127, 329–340 (2006).
Dollins, D.E., Warren, J.J., Immormino, R.M. & Gewirth, D.T. Structures of GRP94-nucleotide complexes reveal mechanistic differences between the Hsp90 chaperones. Mol. Cell 28, 41–56 (2007).
Graf, C., Lee, C.T., Eva Meier-Andrejszki, L., Nguyen, M.T. & Mayer, M.P. Differences in conformational dynamics within the Hsp90 chaperone family reveal mechanistic insights. Front Mol Biosci 1, 4 (2014).
Hellenkamp, B., Wortmann, P., Kandzia, F., Zacharias, M. & Hugel, T. Determination of structural ensembles and correlated dynamics of a multidomain protein. Protocol Exchange http://dx.doi.org/10.1038/protex.2016.078 (2016).
Müller, B.K., Zaychikov, E., Bräuchle, C. & Lamb, D.C. Pulsed interleaved excitation. Biophys. J. 89, 3508–3522 (2005).
Kudryavtsev, V. et al. Combining MFD and PIE for accurate single-pair Forster resonance energy transfer measurements. Chemphyschem 13, 1060–1078 (2012).
Arfken, G. Mathematical Methods for Physicists 3rd edn. 428–436 (Academic Press, 1985).
Case, D. et al. Amber14 (University of California, San Francisco, 2014).
Jorgensen, W.L., Chandrasekhar, J., Madura, J.D., Impey, R.W. & Klein, M.L. Comparison of simple potential functions for simulating liquid water. J. Chem. Phys. 79, 926–935 (1983).
Maier, J.A. et al. ff14SB: Improving the accuracy of protein side chain and backbone parameters from ff99SB. J. Chem. Theory Comput. 11, 3696–3713 (2015).
Darden, T., York, D. & Pedersen, L. Particle mesh Ewald: An N· log (N) method for Ewald sums in large systems. J. Chem. Phys. 98, 10089–10092 (1993).
Shrake, A. & Rupley, J.A. Environment and exposure to solvent of protein atoms. Lysozyme and insulin. J. Mol. Biol. 79, 351–371 (1973).
Humphrey, W., Dalke, A. & Schulten, K. VMD: visual molecular dynamics. J. Mol. Graph. 14, 33–38 (1996).
Acknowledgements
We thank J. Michaelis, C. Seidel, S. Schmid, M. Götz, M. Jahn, J. Buchner, S. Andrade and O. Einsle for helpful discussion. We thank J. Michaelis and C. Seidel for software support. We thank D. Elnatan, D. Agard and M. Götz for help with Δ131Δ expression and purification. This work is funded by the German Research Foundation SFB863 to T.H. (A4) and M.Z. (A10) and the European Research Council through grant agreement 681891 (T.H.).
Author information
Authors and Affiliations
Contributions
B.H., P.W. and T.H. designed the research; B.H. and P.W. performed the research; B.H. developed analytic tools and the software; F.K. and M.Z. performed MD simulations; B.H. analyzed the data in consultation with all authors; B.H. and T.H. wrote the manuscript in consultation with all authors.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
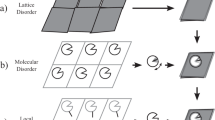
Supplementary Figure 1 Flow chart for dynamic domain arrangement.
To generate dynamic ensembles of protein structures the given steps should be followed. For details see main text and Online Methods.
Supplementary Figure 2 Comparison of Hsp90’s global states.
(a) Superposition of the average open structure of yeast Hsp90 (red) with the closed x-ray structure of yeast Hsp90 (blue) at the C-domains. (b) The -2σ-structure of yeast Hsp90 (reddish) and the x-ray structure of ADP-bound Grp94 (pdb: 2o1v, yellow) match with an RMSD of 4.5 Å. The conformations between M- and N-domain match with an RMSD of 3 Å. (c) Superposition of the 1σ-structure of yeast Hsp90 (reddish) and the open conformation of the bacterial homologue HtpG (pdb: 2ioq, orange).
Supplementary Figure 3 Domain interface dynamics from unrestrained MD simulations.
(a) Root mean square deviations (RMSD) from start structure vs. simulation time for the full-length structure (black) and the structure without N-domains (gray) for the closed (left) and open state (right). (b) Normalized buried surface area per atom (surface illustration) and number of contributions per residue to the buried surface area (cartoon illustration) for the CM-interface. The values are color-coded for the M-domain (top) and C-domain (bottom) for the closed (blue) and open state (red) (see also Online Methods). (c) Distances between charged residues that are located at the CM-interface are plotted vs. simulation time to test for salt bridges. Several charged residues capture relative distances of 2.9 Å, which is characteristic for ionic or hydrogen bonds1,2. These bonds are stable for long time periods of several 10 ns and seem to be protected from water. Some salt bridges interchange in an anti-correlated pattern. A second plateau at 4.5-5 Å could be explained by hydrogen bonds interacting with an enclosed water molecule. The interacting charged residues are well conserved. All amino acids that are well conserved among Hsp90 eukaryotes (conservation score > 80%) are color-coded in red (cartoon plot on the right). A correlation between conservation degree and interface contacts can be seen by comparing this plot with Figure 5 in the main text.
1 Xu, D., Tsai, C.-J. & Nussinov, R. Hydrogen bonds and salt bridges across protein-protein interfaces. Protein engineering 10, 999-1012 (1997). 2 Musafia, B., Buchner, V. & Arad, D. Complex salt bridges in proteins: statistical analysis of structure and function. J Mol Biol 254, 761-770 (1995).
Supplementary Figure 4 Time-correlated distance distributions between different Hsp90 domains.
Time-correlated distance distributions – i.e. distance fluctuations versus observation time – are quantified for the closed state (blue) and the open state (red). Shown are the results for two different FRET pairs between the M-domains (a-b), for one FRET pair between M- and N-domain within a monomer (c), and for one FRET pair between C- and M-domain (d).
Supplementary Figure 5 The effect of cysteine point mutations, labeling and heating on the ATPase activity of Hsp90.
Protein activity was checked by a regenerating ATPase assay1. ATPase rates were measured at 30°C in 40 mM Hepes, 150 mM KCl, 10 mM MgCl2, pH 7.5. ATPase background could be detected by specific inhibition of Hsp90 with radicicol (Sigma) and was subtracted.
The ATPase activity of various constructs is compared to the wild type (WT) activity. The point mutation 51C does show a similar activity, but loses more than 70% of activity upon dye attachment (Atto550-Mal). In addition, this construct (51C, Atto550-Mal) did not close completely after addition of AMP-PNP (as monitored by smFRET). Therefore, this data was excluded from the FRET-network.
For the 517C point mutation the situation is very different. Here the ATPase activity does not significantly change upon the addition of dyes (Atto550-Mal or Atto647N-Mal). Even the exchange a 1:1 mix of Hsp90 517C-Atto550 and -Atto647N, which was heated 20 minutes at 47°C, did not significantly affect the ATPase activity. In addition, these constructs did completely close after the addition of AMP-PNP. This data was included for the FRET-network. Some constructs showed an increase in ATPase, but complete closing, they were also included for the FRET-network.
ATPase activity was measured with a regenerating assay with 1 μM protein under saturating ATP concentration (2 mM) at 37°C. The error bars represent the standard deviations from three independent measurements.
1 Hessling, M., Richter, K. & Buchner, J. Dissection of the ATP-induced conformational cycle of the molecular chaperone Hsp90. Nat Struct Mol Biol 16, 287-293 (2009).
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–5 and Supplementary Notes 1–6 (PDF 1941 kb)
Supplementary Table 1
Mean distance 〈R〉 and standard deviation σR of Gaussian distributed distances extracted from the measured efficiency histograms with the distance distribution analysis are listed for each FRET pair for the closed and open state of Hsp90, together with the deviation from the model (RM-〈R〉), the combined anisotropy rC and the selection criteria for every rejected distance. (XLSX 42 kb)
Supplementary Software
Multidomain Arrangement Software and MD Restraining Support. (ZIP 2568 kb)
Rights and permissions
About this article
Cite this article
Hellenkamp, B., Wortmann, P., Kandzia, F. et al. Multidomain structure and correlated dynamics determined by self-consistent FRET networks. Nat Methods 14, 174–180 (2017). https://doi.org/10.1038/nmeth.4081
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nmeth.4081
This article is cited by
-
Fluorescence resonance energy transfer at the single-molecule level
Nature Reviews Methods Primers (2024)
-
An automated single-molecule FRET platform for high-content, multiwell plate screening of biomolecular conformations and dynamics
Nature Communications (2023)
-
Reliability and accuracy of single-molecule FRET studies for characterization of structural dynamics and distances in proteins
Nature Methods (2023)
-
Structural dynamics of DNA strand break sensing by PARP-1 at a single-molecule level
Nature Communications (2022)
-
Can DyeCycling break the photobleaching limit in single-molecule FRET?
Nano Research (2022)