Abstract
Complex biomedical analyses require the use of multiple software tools in concert and remain challenging for much of the biomedical research community. We introduce GenomeSpace (http://www.genomespace.org), a cloud-based, cooperative community resource that currently supports the streamlined interaction of 20 bioinformatics tools and data resources. To facilitate integrative analysis by non-programmers, it offers a growing set of 'recipes', short workflows to guide investigators through high-utility analysis tasks.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout

Similar content being viewed by others
References
Demchak, B. et al. F1000Res. 3, 151 (2014).
Shannon, P. et al. Genome Res. 13, 2498–2504 (2003).
Giardine, B. et al. Genome Res. 15, 1451–1455 (2005).
Reich, M. et al. Nat. Genet. 38, 500–501 (2006).
Segal, E., Friedman, N., Koller, D. & Regev, A. Nat. Genet. 36, 1090–1098 (2004).
Robinson, J.T. et al. Nat. Biotechnol. 29, 24–26 (2011).
Karolchik, D. et al. Nucleic Acids Res. 32, D493–D496 (2004).
Ben-Porath, I. et al. Nat. Genet. 40, 499–507 (2008).
Wong, D.J. et al. Cell Stem Cell 2, 333–344 (2008).
Sambrook, J., Fritsch, E.F. & Maniatis, T. Molecular Cloning: A Laboratory Manual vol. 3 (Cold Spring Harbor Laboratory Press, 1989).
Krivtsov, A.V. et al. Nature 442, 818–822 (2006).
Larsson, J. & Karlsson, S. Oncogene 24, 5676–5692 (2005).
Floratos, A., Smith, K., Ji, Z., Watkinson, J. & Califano, A. Bioinformatics 26, 1779–1780 (2010).
Saeed, A.I. et al. Biotechniques 34, 374–378 (2003).
Gorton, I., Wynne, A., Almquist, J. & Chatterton, J. in Software Architecture, 2008. WICSA 2008. Seventh Working IEEE/IFIP Conference (eds. Kruchten, P., Garlan, D. & Woods, E.) 95–104 (IEEE Computer Society, 2008).
Shannon, P.T., Reiss, D.J., Bonneau, R. & Baliga, N.S. BMC Bioinformatics 7, 176 (2006).
Acknowledgements
We thank other members of the GenomeSpace and GenePattern Teams for their contributions and input: P. Carr, B. Hill-Meyers, S.H. Lee and T. Tabor (Broad Institute of MIT and Harvard); J. Zhang (Stanford University); and H. Carter and M. Smoot (University of California, San Diego). Special thanks to D. Haussler and J. Kent (University of California, Santa Cruz) for their involvement in the nascent stages of the GenomeSpace project. We thank J. Bistline for help with the citations and figures, and L. Gaffney for help with the figures. This work has been supported by US National Institutes of Health–National Human Genome Research Institute P01 HG005062 and U41 HG007517, with additional initial support from Amazon Web Services (AWS).
Author information
Authors and Affiliations
Contributions
M.R., A.R. and J.P.M. conceived of the GenomeSpace concept. T.L., M.O. and M.R. designed and implemented the GenomeSpace software. K.Q., S.G., F.W. and N.P. implemented the driving biological projects within GenomeSpace with supervision and input from A.R., H.Y.C. and J.P.M. The recipes were implemented by S.G., F.W. and D.B.-R. The GenomeSpace seed tools were added to the system by J.T.R., B.D., T.H., G.B.-A., D.B., G.P.B., B.T.L., R.M.K., A.N., E.S. and T.I., who also consulted on the GenomeSpace architecture. H.T., M.R., A.R., H.Y.C. and J.P.M. supervised the GenomeSpace project. K.Q., S.G., F.W., H.T., M.R., H.C., A.R. and J.P.M. wrote the manuscript. All authors reviewed and approved the final manuscript as submitted.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
Supplementary Figure 1 GenomeSpace allows the biologist user to conduct complex analysis.
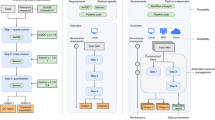
1a. Flow chart of our analytic approach to dissecting a stem cell-like gene regulatory network in human cancers. Rectangles indicate necessary input data. Rounded rectangles indicate the overall goal achieved by the analysis, grouped into specific analysis steps from the Supplementary Note 1 (brown text). Each group of analysis steps investigates a specific biological question (on the left). See Supplementary Figure 2 for definitions of some terms used in this example analysis. 1b. Detailed steps and tools involved in the integrative genomics analysis scenario for the driving biological project performed using GenomeSpace are numbered in order of execution, and correspond to the numbered steps in the Supplementary Note 1.
Supplementary Figure 2 Detailed definitions of some terms used in Supplementary Note 1 and Supplementary Figure 1.
Stemness signature: a set of genes which are upregulated in induced cancer stem cells (iCSCs), and enriched in embryonic stem cells (ESCs) and induced pluripotent stem cells (iPSCs). These genes represent a stem cell state. Non-stemness signature: a set of genes which are upregulated in iCSCs, but are not associated with the stemness signature. Stemness “ON”: a set of breast cancer tumor samples which have significant enrichment in the stemness signature. Stemness “OFF”: a set of breast cancer tumor samples which do not have enrichment of the stemness signature.
Supplementary Figure 3 Output from Genomica’s ModuleMap.
The ModuleMap tool is used to identify a shared set of genes induced in embryonic (ESC) or induced pluripotent stem cells (iPSC) and also in cancer stem cells. Main panel: Expression pattern of 550 “stemness genes” in the indicated samples. Each row is a gene and each column is an array. Top panel: Enrichment of the indicated gene sets in each sample. Each row is a gene set and each column is an array. Red indicates coordinate gene set induction (P < 0.05, FDR < 0.05) in a corresponding array; green indicates coordinate gene set repression (P < 0.05, FDR <0.05). For example, stem cell genes are induced in ESC and iPSC arrays, but are not enriched in differentiated arrays. Right panel: Red pixel indicates overlap between a stemness gene and membership in the iCSC gene signature. There is 100% overlap, as the 550 stemness genes were also found in the iCSC gene signature.
Supplementary Figure 4 IGV visualization of CNV profiles.
Tumor samples with a “stemness ON” gene signature are found to have recurrent amplification of chromosome 8q. IGV is used to visualize the genome-wide chromosome copy number variation of all breast cancer tumor samples, ranked by the presence or absence of the stemness gene signature. Red indicates DNA copy number gain; blue indicates DNA copy number loss.
Supplementary Figure 5 Cytoscape visualization of gene regulatory network.
Cytoscape is used to visualize the gene regulatory network in human cancer stem cells. Each node is a gene, and each edge is a predicted regulatory connection between genes.
Supplementary Figure 6 GenomeSpace web application.
Users interact with GenomeSpace through a web-based interface, the GS-UI. (1) A menu bar gives access to all of the GS-UI functionality. A scrollable tool bar below the menu bar contains icons to launch each GenomeSpace tool, with options to send one or more input files to the tool. Users can rearrange the icons and hide icons for tools they use less frequently. Here the tool bar contains (2) analysis and visualization tools, (3) data resources, and (4) integrated portals. (5) Clicking on the arrow will scroll the tool bar to show more tools. (6, 7) A view of the data stored in a user’s GenomeSpace account uses a familiar file system representation for navigation. (8) The file navigator also displays a number of other directories, including a GenomeSpace Public directory that contains data hosted by the GenomeSpace team, a directory with links to files that other users have shared with this account, and directories the user has mounted from other data sources, such as Dropbox, Google Drive, and Amazon S3.
Supplementary Figure 7 Representation of the workflow for the recipe "Find subnetworks of differentially expressed genes and identify associated biological functions.”
This cartoon workflow represents the recipe steps and includes three sections: (1) input data; (2) analyses completed in GenomeSpace tools (GenePattern and Cytoscape); and (3) the output of the recipe. Specifically, it passes an input gene expression dataset from two conditions or phenotypes (e.g., naïve and mature cells, tumor and normal) to GenePattern to find the list of the top 50 differentially expressed genes. Next, the GeneMANIA Cytoscape plugin is used to identify a network connecting these differentially expressed genes, derived from a resource collection of interaction networks, e.g., co-expression, co-localization, or protein-protein interactions. This network is then probed for highly interconnected subnetworks from which active underlying biological processes can be inferred, e.g., by consisting of genes with similar functional annotations.
Supplementary Figure 8 Output from Example Recipe, “Find subnetworks of differentially expressed genes and identify associated biological functions.”
8a. Cytoscape network visualization of the top 50 genes differentially expressed between normal granulocyte/macrophage progenitor cells and leukemia stem cells. Nodes represent genes. Edges indicate pathways and interactions identified using the GeneMANIA functional association database. Subnetworks of highly connected nodes are identified using the MCODE algorithm and are displayed in the side panel. 8b. Cytoscape visualization of one such subnetwork. The bottom panel displays the names of genes found in the subnetwork along with metadata, e.g., descriptions and Gene Ontology (GO) Terms, associated with each gene. This subnetwork, for example, contains the SMAD1 gene, which is involved in the BMP signaling pathway.
Supplementary Figure 9 Representation of the workflow for the recipe "Identify biological functions for genes in CNV regions.”
This cartoon workflow represents the recipe steps and includes three sections: (1) input data, including data imported from a GenomeSpace tool (UCSC Table Browser); (2) analyses completed in GenomeSpace tools (Galaxy and MSigDB); and (3) the output of the recipe.
Supplementary Figure 10 Output from Example Recipe, "Identify biological functions for genes in CNV regions.”
MSigDB results for the top gene sets found to be amplified or deleted in CNV regions. Top panel indicates the top gene sets enriched in the list of genes found in CNV regions; bottom panel is a gene set matrix illustrating which gene sets the genes in CNV regions belong to. 10a. Results for genes amplified in CNV regions. 10b. Results for genes depleted in CNV regions.
Supplementary Figure 11 GenomeSpace architecture.
Cartoon illustrating of the GenomeSpace underlying architecture. GenomeSpace presents a “connection layer” that includes a collection of web services with well-defined entry points to the GenomeSpace server that provides the core system functionality.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–11, Supplementary Table 1 and Supplementary Notes 1–3. (PDF 1808 kb)
Rights and permissions
About this article
Cite this article
Qu, K., Garamszegi, S., Wu, F. et al. Integrative genomic analysis by interoperation of bioinformatics tools in GenomeSpace. Nat Methods 13, 245–247 (2016). https://doi.org/10.1038/nmeth.3732
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nmeth.3732
This article is cited by
-
Galaxy External Display Applications: closing a dataflow interoperability loop
Nature Methods (2020)
-
SyNDI: synchronous network data integration framework
BMC Bioinformatics (2018)