Abstract
Quantification of cell-cycle state at a single-cell level is essential to understand fundamental three-dimensional (3D) biological processes such as tissue development and cancer. Analysis of 3D in vivo images, however, is very challenging. Today's best practice, manual annotation of select image events, generates arbitrarily sampled data distributions, which are unsuitable for reliable mechanistic inferences. Here, we present an integrated workflow for quantitative in vivo cell-cycle profiling. It combines image analysis and machine learning methods for automated 3D segmentation and cell-cycle state identification of individual cell-nuclei with widely varying morphologies embedded in complex tumor environments. We applied our workflow to quantify cell-cycle effects of three antimitotic cancer drugs over 8 d in HT-1080 fibrosarcoma xenografts in living mice using a data set of 38,000 cells and compared the induced phenotypes. In contrast to results with 2D culture, observed mitotic arrest was relatively low, suggesting involvement of additional mechanisms in their antitumor effect in vivo.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Baker, B.M. & Chen, C.S. Deconstructing the third dimension: how 3D culture microenvironments alter cellular cues. J. Cell Sci. 125, 3015–3024 (2012).
Condeelis, J. & Weissleder, R. In vivo imaging in cancer. Cold Spring Harb. Perspect. Biol. 2, a003848 (2010).
Orth, J.D. et al. Analysis of mitosis and antimitotic drug responses in tumors by in vivo microscopy and single-cell pharmacodynamics. Cancer Res. 71, 4608–4616 (2011).
Pittet, M.J. & Weissleder, R. Intravital imaging. Cell 147, 983–991 (2011).
Thurber, G.M. et al. Single-cell and subcellular pharmacokinetic imaging allows insight into drug action in vivo. Nat. Commun. 4, 1504 (2013).
Sakaue-Sawano, A. et al. Visualizing spatiotemporal dynamics of multicellular cell-cycle progression. Cell 132, 487–498 (2008).
Carpenter, A.E. et al. CellProfiler: image analysis software for identifying and quantifying cell phenotypes. Genome Biol. 7, R100 (2006).
Held, M. et al. CellCognition: time-resolved phenotype annotation in high-throughput live cell imaging. Nat. Methods 7, 747–754 (2010).
Eliceiri, K.W. et al. Biological imaging software tools. Nat. Methods 9, 697–710 (2012).
Rasheed, S., Nelson-Rees, W.A., Toth, E.M., Arnstein, P. & Gardner, M.B. Characterization of a newly derived human sarcoma cell line (HT-1080). Cancer 33, 1027–1033 (1974).
Sakaue-Sawano, A., Kobayashi, T., Ohtawa, K. & Miyawaki, A. Drug-induced cell cycle modulation leading to cell-cycle arrest, nuclear mis-segregation, or endoreplication. BMC Cell Biol. 12, 2 (2011).
Umesh Adiga, P.S. & Chaudhuri, B.B. An efficient method based on watershed and rule-based merging for segmentation of 3-D histo-pathological images. Pattern Recognit. 34, 1449–1458 (2001).
Lin, G. et al. A hybrid 3D watershed algorithm incorporating gradient cues and object models for automatic segmentation of nuclei in confocal image stacks. Cytometry A 56A, 23–36 (2003).
Wählby, C., Sintorn, I.M., Erlandsson, F., Borgefors, G. & Bengtsson, E. Combining intensity, edge and shape information for 2D and 3D segmentation of cell nuclei in tissue sections. J. Microsc. 215, 67–76 (2004).
Lin, G. et al. Hierarchical, model-based merging of multiple fragments for improved three-dimensional segmentation of nuclei. Cytometry A 63A, 20–33 (2005).
Lin, G. et al. A multi-model approach to simultaneous segmentation and classification of heterogeneous populations of cell nuclei in 3D confocal microscope images. Cytometry A 71A, 724–736 (2007).
Al-Kofahi, Y., Lassoued, W., Lee, W. & Roysam, B. Improved automatic detection and segmentation of cell nuclei in histopathology images. IEEE Trans. Biomed. Eng. 57, 841–852 (2010).
Vincent, L. Morphological grayscale reconstruction in image analysis: applications and efficient algorithms. IEEE Trans. Image Process. 2, 176–201 (1993).
Meyer, F. Topographic distance and watershed lines. Signal Process. 38, 113–125 (1994).
Amat, F. et al. Fast, accurate reconstruction of cell lineages from large-scale fluorescence microscopy data. Nat. Methods 11, 951–958 (2014).
Mitchison, T.J. The proliferation rate paradox in antimitotic chemotherapy. Mol. Biol. Cell 23, 1–6 (2012).
Komlodi-Pasztor, E., Sackett, D., Wilkerson, J. & Fojo, T. Mitosis is not a key target of microtubule agents in patient tumors. Nat. Rev. Clin. Oncol. 8, 244–250 (2011).
Spencer, S.L. & Sorger, P.K. Measuring and modeling apoptosis in single cells. Cell 144, 926–939 (2011).
Jordan, M.A. & Wilson, L. Microtubules as a target for anticancer drugs. Nat. Rev. Cancer 4, 253–265 (2004).
Janssen, A., Beerling, E., Medema, R. & van Rheenen, J. Intravital FRET imaging of tumor cell viability and mitosis during chemotherapy. PLoS ONE 8 (2013).
Milas, L. et al. Kinetics of mitotic arrest and apoptosis in murine mammary and ovarian tumors treated with taxol. Cancer Chemother. Pharmacol. 35, 297–303 (1995).
Zasadil, L.M. et al. Cytotoxicity of paclitaxel in breast cancer is due to chromosome missegregation on multipolar spindles. Sci. Transl. Med. 6, 229ra43 (2014).
Gascoigne, K.E. & Taylor, S.S. Cancer cells display profound intra- and interline variation following prolonged exposure to antimitotic drugs. Cancer Cell 14, 111–122 (2008).
Shi, J., Orth, J.D. & Mitchison, T. Cell type variation in responses to antimitotic drugs that target microtubules and kinesin-5. Cancer Res. 68, 3269–3276 (2008).
Purcell, J.W. et al. Activity of the kinesin spindle protein inhibitor ispinesib (SB-715992) in models of breast cancer. Clin. Cancer Res. 16, 566–576 (2010).
Towle, M.J. et al. Broad spectrum preclinical antitumor activity of eribulin (Halaven(R)): optimal effectiveness under intermittent dosing conditions. Anticancer Res. 32, 1611–1619 (2012).
Manfredi, M.G. et al. Characterization of alisertib (MLN8237), an investigational small-molecule inhibitor of aurora A kinase using novel in vivo pharmacodynamic assays. Clin. Cancer Res. 17, 7614–7624 (2011).
Towle, M.J. et al. Eribulin induces irreversible mitotic blockade: implications of cell-based pharmacodynamics for in vivo efficacy under intermittent dosing conditions. Cancer Res. 71, 496–505 (2011).
Mori, T. et al. Retention of paclitaxel in cancer cells for 1 week in vivo and in vitro. Cancer Chemother. Pharmacol. 58, 665–672 (2006).
Vanhoefer, U., Cao, S., Harstrick, A., Seeber, S. & Rustum, Y.M. Comparative antitumor efficacy of docetaxel and paclitaxel in nude mice bearing human tumor xenografts that overexpress the multidrug resistance protein (MRP). Ann. Oncol. 8, 1221–1228 (1997).
Friedrich, J., Seidel, C., Ebner, R. & Kunz-Schughart, L.A. Spheroid-based drug screen: considerations and practical approach. Nat. Protoc. 4, 309–324 (2009).
Tosoni, D., Di Fiore, P.P. & Pece, S. Functional purification of human and mouse mammary stem cells. Methods Mol. Biol. 916, 59–79 (2012).
Fan, J. Notes on Poisson distribution-based minimum error thresholding. Pattern Recognit. Lett. 19, 425–431 (1998).
Vapnik, V.N. Statistical Learning Theory (Wiley, 1998).
Ng, A.Y. & Jordan, M.I. On discriminative vs. generative classifiers: a comparison of logistic regression and naive Bayes. Adv. Neural Inf. Process. Syst. 14, 841–848 (2002).
Ning, J., Zhang, L., Zhang, D. & Wu, C. Interactive image segmentation by maximal similarity based region merging. Pattern Recognit. 43, 445–456 (2010).
Breiman, L. Random Forests. Mach. Learn. 45, 5–32 (2001).
Otsu, N. A threshold selection method from gray-level histograms. IEEE Trans. Syst. Man Cybern. 9, 62–66 (1979).
Klein, S., Staring, M., Murphy, K., Viergever, M.A. & Pluim, J.P.W. Elastix: a toolbox for intensity-based medical image registration. IEEE Trans. Med. Imaging 29, 196–205 (2010).
Haralick, R.M., Shanmugam, K. & Dinstein, I. Textural features for image classification. IEEE Trans. Syst. Man Cybern. SMC-3, 610–621 (1973).
Hall, M. et al. The WEKA data mining software: an update. SIGKDD Explor. 11, 10–18 (2009).
Acknowledgements
S.F. was supported by a research fellowship (FL 820/1-1) from the DFG (Deutsche Forschungsgemeinschaft). This project was supported by US National Institutes of Health grants R01-CA164448, S10RR0266360 and PO1-CA139980. We thank P. Choi for helpful discussions. We are grateful for the support of The Nikon Imaging Center at Harvard Medical School. We thank K. Krukenberg (Harvard Medical School) for the MCF7 and T47D cell lines stably expressing H2B-GFP; P. Keller (Howard Hughes Medical Institute, Janelia Research Campus) for Drosophila, mouse and zebrafish data sets; M. Sebas for surgical implantation of the DSCs; and J. Moore for assistance with flow cytometry. The FUCCI viral particles were a kind gift from P. Jorgensen, A. Tzur and M. Chung (Harvard Medical School) generated with vectors kindly provided by the laboratory of A. Miyawaki (RIKEN Brain Science Institute). The H2B-CFP construct was obtained from A. Loewer (Max Delbrueck Center).
Author information
Authors and Affiliations
Contributions
D.R.C. designed and developed the automated image analysis framework. S.F. initiated and coordinated the project and designed the experiments. S.F. and R.H.K. performed the mouse experiments. S.F. performed the spheroid experiments. D.R.C. and S.F. analyzed all experiments and wrote the manuscript, R.H.K. developed the grid-based imaging setup for longitudinal observations. J.D.O. generated the stable HT-1080 cell line. Y.I. performed the immunohistochemistry staining. R.W., G.D. and T.J.M. initiated the collaboration and helped edit the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
Supplementary Figure 1 Segmentation accuracy in undiluted xenograft tumors
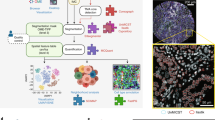
Original images and segmentation results for a full density, undiluted xenograft HT-1080 tumor expressing H2B CFP and FUCCI are shown. The local signal to background thresholding algorithm provided in the software was used to obtain the initial foreground-background segmentation for these datasets.
Supplementary Figure 2 Segmentation accuracy for varying levels of cell density
Cell density is estimated as the percentage of voxels belonging to the foreground/cell-nuclei in the histone channel wherein the foreground mask is computed using the locally adaptive Poisson-based minimum error thresholding method used in our nuclei segmentation algorithm. Shown below the plot are 3D volume visualizations of a few volumes in our test dataset along with their cell densities.
Supplementary Figure 3 Segmentation accuracy in different biological systems acquired with multiple microscopy setups
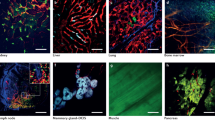
The datasets in (a)-(c) were kindly provided by Philipp Keller and were also analyzed by his group in Fig. 3 of Amat et al., 201420. (a) Segmentation analysis of a stage 6 Drosophila embryo expressing His2Av-mRFP1 acquired on a Zeiss Lightsheet Z.1 microscope (timepoint 500 of the dataset), (b) a zebrafish embryo during early gastrulation expressing H2B-mCherry acquired on a Zeiss LSM 710 laser-scanning microscope (timepoint 0 of the dataset) and (c) a mouse embryo expressing H2B GFP acquired on a SimView light sheet microscope (timepoint 0 of the dataset) (d)-(f) Segmentation results for cancer cell line spheroids grown from (d) HT-1080, (e) T47D and (f) MCF7 cells and expressing H2B GFP. The LoG response thresholding and the local signal to background ratio thresholding algorithms provided in the software were chosen to obtain the initial foreground-background segmentation for the datasets in panels (a) and (b-f), respectively.
Supplementary Figure 4 Validation of cell-cycle quantification in vivo: histology
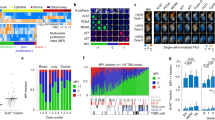
Twelve mice were injected with 2 million HT-1080 cells into the flank and subcutaneous tumors were allowed to grow for 3 weeks. Then, 3 mice each were either left untreated, or injected with a single dose of 40 mg/kg paclitaxel i.v. and sacrificed after 1, 4 or 7 days, respectively. Tumors were fixed and sections were stained with an antibody against the mitotic marker phospho-histone H3. The mitotic index (% mitotic cells) was quantified at 6 positions in each tumor. Note that the increase in mitotic index is relatively low compared to a 2D tissue culture environment and similar to, though slightly lower, than the data in Fig. 5b. The slightly lower mitotic index obtained through histology is probably due to the fact that the immunohistochemistry staining counts host cells as well, while the FUCCI imaging system is more accurate in this regard, labeling only tumor cells.
Supplementary Figure 5 Validation of cell-cycle quantification in 3D culture: flow cytometry
(a) a comparison of the FUCCI system and the DNA content quantification approach for determination of the cell cycle distribution. Note that they are not equivalent: the FUCCI/H2B combination cannot discriminate between S and G2, while the DNA content method does not separate G2 and M (one 4N peak). (b) HT-1080 spheroids were treated with 2 µM paclitaxel or the corresponding concentration of DMSO as control and the cell cycle distribution was assessed by spinning disk confocal microscopy of three spheroids from each group followed by software based automated analysis using our computational framework. (c) Then, all spheroids were dispersed into a single-cell suspension by a 1 min treatment with trypsin, stained with DyeCyle Violet dye and the DNA content was analyzed. Note that the changes in the G1 and S/G2/M compartments are almost identical with both methods. ~1500 cells per condition were analyzed in (b) and 5000 cells per condition were analyzed in (c).
Supplementary Figure 6 Montage of all cells analyzed from mice treated with paclitaxel
Cells from the mice treated with paclitaxel, as analyzed in Fig. 5, were arranged as in Fig. 4e to allow visual inspection of segmentation/classification accuracy and cell morphology by timepoint. A gap of two or more days between the imaged timepoints is highlighted by two oblique lines intersecting the axis. To see the cell thumbnails at full resolution, we recommend looking at the electronic version.
Supplementary Figure 7 Montage of all cells analyzed from mice treated with eribulin
Cells from the second mouse treated with eribulin, as analyzed in Fig. 5, were arranged as in Fig. 4e to allow visual inspection of segmentation/classification accuracy and cell morphology by timepoint. A gap of two or more days between the imaged timepoints is highlighted by two oblique lines intersecting the axis. To see the cell thumbnails at full resolution, we recommend looking at the electronic version.
Supplementary Figure 8 Montage of all cells analyzed from mice treated with ispinesib
Cells from the mice treated with ispinesib, as analyzed in Fig. 5, were arranged as in Fig. 4e to allow visual inspection of segmentation/classification accuracy and cell morphology by timepoint. A gap of two or more days between the imaged timepoints is highlighted by two oblique lines intersecting the axis. To see the cell thumbnails at full resolution, we recommend looking at the electronic version.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–8 and Supplementary Tables 1–13 (PDF 2133 kb)
Supplementary Software
InvivoCytometer 2.0 The software implementation of our proposed segmentation and cell cycle classification algorithms. Included: MATLAB source code, documentation, sample data (ZIP 147667 kb)
Source data
Rights and permissions
About this article
Cite this article
Chittajallu, D., Florian, S., Kohler, R. et al. In vivo cell-cycle profiling in xenograft tumors by quantitative intravital microscopy. Nat Methods 12, 577–585 (2015). https://doi.org/10.1038/nmeth.3363
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nmeth.3363
This article is cited by
-
Temporal and spatial topography of cell proliferation in cancer
Nature Cell Biology (2022)
-
In vivo microscopy reveals macrophage polarization locally promotes coherent microtubule dynamics in migrating cancer cells
Nature Communications (2020)
-
Cell cycle dynamics during diapause entry and exit in an annual killifish revealed by FUCCI technology
EvoDevo (2019)
-
Predicting How to Distribute Work Between Algorithms and Humans to Segment an Image Batch
International Journal of Computer Vision (2019)
-
Heterogeneity in tumor chromatin-doxorubicin binding revealed by in vivo fluorescence lifetime imaging confocal endomicroscopy
Nature Communications (2018)