Abstract
Whole-genome bisulfite sequencing (WGBS) allows genome-wide DNA methylation profiling, but the associated high sequencing costs continue to limit its widespread application. We used several high-coverage reference data sets to experimentally determine minimal sequencing requirements. We present data-derived recommendations for minimum sequencing depth for WGBS libraries, highlight what is gained with increasing coverage and discuss the trade-off between sequencing depth and number of assayed replicates.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
References
Okano, M., Bell, D.W., Haber, D.A. & Li, E. Cell 99, 247–257 (1999).
Bird, A. Genes Dev. 16, 6–21 (2002).
Reik, W. Nature 447, 425–432 (2007).
Ziller, M.J. et al. Nature 500, 477–481 (2013).
NIH Roadmap Epigenomics Mapping Consortium. Standards and guidelines for whole genome shotgun bisulfite sequencing http://www.roadmapepigenomics.org/protocol (2011).
Mehta, T., Tanik, M. & Allison, D.B. Nat. Genet. 36, 943–947 (2004).
Gifford, C.A. et al. Cell 153, 1149–1163 (2013).
Hansen, K.D., Langmead, B. & Irizarry, R.A. Genome Biol. 13, R83 (2012).
Sun, D. et al. Genome Biol. 15, R38 (2014).
Meynert, A.M., Bicknell, L.S., Hurles, M.E., Jackson, A.P. & Taylor, M.S. BMC Bioinformatics 14, 195 (2013).
Tarazona, S., García-Alcalde, F., Dopazo, J., Ferrer, A. & Conesa, A. Genome Res. 21, 2213–2223 (2011).
Hansen, K.D. et al. Genome Res. 24, 177–184 (2014).
Gentleman, R.C. et al. Genome Biol. 5, R80 (2004).
Köster, J. & Rahmann, S. Bioinformatics 28, 2520–2522 (2012).
Xi, Y. & Li, W. BMC Bioinformatics 10, 232 (2009).
Acknowledgements
This work was supported by the NIH Common Fund (U01ES017155), NIGMS (P01GM099117) and the New York Stem Cell Foundation. A.M. receives support as a New York Stem Cell Foundation Robertson Investigator.
Author information
Authors and Affiliations
Contributions
M.J.A. and A.M. conceived of the study. M.J.Z., M.J.A. and K.D.H. performed analysis and interpreted results. M.J.Z., M.J.A., K.D.H. and A.M. wrote the paper.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
Supplementary Figure 1 Detailed analysis of WGBS coverage requirements
a. Left: Change of true positive rate (delta TPR, y-axis) as a function of coverage (x-axis) when comparing hESC and cortex using two replicates per group. True positive rate is defined as the fraction of high coverage (30×) DMRs recovered at the coverage level indicated. Line color indicates the methylation threshold used to filter DMRs by minimum methylation difference. Grey box indicates coverage range where change in TPR exhibits the largest drop. Right: Change of true positive rate (delta TPR, y-axis) as a function of coverage (x-axis) for comparing CD4 and CD8 cells using 2 replicates for each group. Grey box indicates coverage range where change in TPR exhibits the largest drop.
b. Left: True positive rate (TPR, y-axis) as a function of coverage (x-axis) for comparing hESC and cortex using 2 replicates for each group. True positive rate is defined as the fraction of high coverage (30×) DMRs recovered at the coverage level indicated. Line color indicates the methylation threshold used to filter DMRs by minimum methylation difference. Right: True positive rate (TPR, y-axis) as a function of coverage (x-axis) when comparing CD4 vs. CD8 using 2 replicates per group. True positive rate is defined as the fraction of high coverage (30×) DMRs recovered at the coverage level indicated. Line color indicates the methylation threshold used to filter DMRs by minimum methylation difference.
c. Left: Percentage of 30× DMRs within distinct size ranges recovered as a function of coverage based on a two replicate comparison between hESC and cortex. Middle: Percentage of 30× DMRs within distinct CpG density ranges recovered as a function of coverage based on a two replicate comparison between hESC and cortex. Right: Distribution of DMR sizes (x-axis) and average methylation difference (y-axis) for DMRs discovered at 1× (grey) and additional DMRs discovered when increasing the coverage from 1× to 5× (dark red), 5× to 10× (light red) and 10× to 30× (orange) in the CD4s vs CD8 comparison using 2 replicates each. Black dots indicate median and ellipsoids span from the 25th to the 75th percentile in each dimension.
d. True positive rate (TPR, y-axis) as a function of coverage for a two replicate based comparison of hESC and cortex as well as CD4 and CD8. DMRs were identified using a single CpG and not a smoothing based method. A separate, single CpG method derived high coverage (30×) set was used as a reference.
e. False positive discorey rate (FDR, y-axis) as a function of coverage for a two replicate based comparison of hESC and cortex as well as CD4 and CD8. DMRs were identified using a single CpG and not a smoothing based method. A separate, single CpG method derived high coverage (30×) set was used as a reference.
f. Number of CpGs covered with at least one read (y-axis) as a function of average genomic coverage (x-axis) for 3 replicates of hESC and 3 replicates of human cortex.
g. Left: Percentage of CpGs covered with different numbers of reads (y-axis) as a function of total genomic coverage (x-axis). Right: Number of CpGs covered with different numbers of reads (y-axis) as a function of total genomic coverage (x-axis).
h. Median CpG coverage (y-axis) as a function of total genomic coverage (x-axis). False discovery rate (FDR, y-axis) as function of coverage (x-axis) for DMRs arising in the hESC vs. cortex comparison using two replicates for each group. Line color indicates the methylation threshold used to filter DMRs by minimum methylation difference.
i. True positive rate (TPR, y-axis) as a function of coverage (x-axis) when comparing hESC and cortex using a high stringency DMR set employing 3 replicates for each group (see Online Methods for details). True positive rate is defined as the fraction of high stringency DMRs recovered at the coverage level indicated. Line color indicates the methylation threshold used to filter DMRs by minimum methylation difference.
j. False discovery rate (FDR, y-axis) as function of coverage (x-axis) for DMRs arising in the hESC vs. cortex comparison using two replicates for each group. Line color indicates the methylation threshold used to filter DMRs by minimum methylation difference.
k. False discovery rate (FDR, y-axis) as function of coverage (x-axis) for DMRs arising in the CD4 vs. CD8 comparison using two replicates per group. Line color indicates the methylation threshold used to filter DMRs by minimum methylation difference.
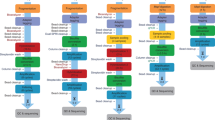
Supplementary Figure 2 Experimental design recommendations for WGBS replicate number and sequencing depth
a. True positive rate (TPR, y-axis) as a function of coverage (x-axis) comparing for DMRs identified between CD4 and CD8 or CD184 and liver using one or two replicates in each comparison for DMRs with a methylation difference greater than 20%. High coverage (30×) DMR set based on two replicates was used as reference true positive set.
b. False discovery rate (FDR, y-axis) as a function of coverage (x-axis) comparing for DMRs identified between CD4 and CD8 or CD184 and liver using one or two replicates in each comparison for DMRs with a methylation difference greater than 20%. High coverage (30×) DMR set based on two replicates was used as reference true positive set.
c. True positive rate (left) and false discovery rate (right) as a function of coverage using 1, 2, or 3 replicates of hESC and cortex samples, requiring a minimum methylation difference of 20%. DMRs were identified using a single CpG and not a smoothing based method. High coverage (30×), 3 replicate set was used as a reference.
d. Schematic summarizing study results with y-axis indicating number of replicates and x-axis indicating coverage level. Recommended number of replicates and sequencing depth for the hESC vs cortex and CD4 vs CD8 comparisons are indicated by red crosses. Red striped area indicates coverage below recommended levels, with the exception of situations where only very large DMRs are of interest.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1 and 2 (PDF 1273 kb)
Source data
Rights and permissions
About this article
Cite this article
Ziller, M., Hansen, K., Meissner, A. et al. Coverage recommendations for methylation analysis by whole-genome bisulfite sequencing. Nat Methods 12, 230–232 (2015). https://doi.org/10.1038/nmeth.3152
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nmeth.3152