Abstract
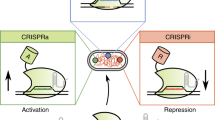
A key obstacle to creating sophisticated genetic circuits has been the lack of scalable device libraries. Here we present a modular transcriptional repression architecture based on clustered regularly interspaced palindromic repeats (CRISPR) system and examine approaches for regulated expression of guide RNAs in human cells. Subsequently we demonstrate that CRISPR regulatory devices can be layered to create functional cascaded circuits, which provide a valuable toolbox for engineering purposes.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
References
E. Andrianantoandro, S. Basu, D. K. Karig & Weiss, R. Mol. Systems Biol. 2, 0028 (2006).
Ruder, W.C., Lu, T. & Collins, J.J. Science 333, 1248–1252 (2011).
Slusarczyk, A.L., Lin, A. & Weiss, R. Nat. Rev. Genet. 13, 406–420 (2012).
Bird, J. Engineering Mathematics 532 (Elsevier Science, 2007).
Peirce, C.S. Collected Papers of Charles Sanders Peirce vol. 4, 12–20 (Harvard University Press, 1933).
Farzadfard, F., Perli, S.D. & Lu, T.K. ACS Synth. Biol. 2, 604–613 (2013).
Khalil, A.S. et al. Cell 150, 647–658 (2012).
Garg, A. et al. Nucleic Acids Res. 40, 7584–7595 (2012).
Kramer, B.P., Fischer, C. & Fussenegger, M. Biotechnol. Bioeng. 87, 478–484 (2004).
Stanton, B.C. et al. Nat. Chem. Biol. 10, 99–105 (2014).
Fu, Y. et al. Nat. Biotechnol. 31, 822–826 (2013).
Mali, P. et al. Science 339, 823–826 (2013).
Qi, L.S. et al. Cell 152, 1173–1183 (2013).
Nielsen, A.A., Segall-Shapiro, T.H. & Voigt, C.A. Curr. Opin. Chem. Biol. 17, 878–892 (2013).
Henriksen, J.R. et al. Nucleic Acids Res. 35, e67 (2007).
Ko, J.K. et al., FASEB J. 25, 2638–2649 (2011).
Xia, H. et al. Nat. Biotechnol. 20, 1006–1010 (2002).
Giering, J.C., Grimm, D., Storm, T.A. & Kay, M.A. Mol. Ther. 16, 1630–1636 (2008).
Lin, S.L. et al. Biochem. Biophys. Res. Commun. 310, 754–760 (2003).
Cardinale, S. & Arkin, A.P. Biotechnol. J. 7, 856–866 (2012).
Gyorgy, A. & Del Vecchio, D. PLoS Comput. Biol. doi:10.1371/journal.pcbi.1003486 (13 March 2014).
Dennis, P.P., Ehrenberg, M. & Bremer, H. Microbiol. Mol. Biol. Rev. 68, 639–668 (2004).
Klumpp, S. & Hwa, T. Proc. Natl. Acad. Sci. USA 105, 20245–20250 (2008).
Esvelt, K.M. et al. Nat. Methods 10, 1116–1121 (2013).
Acknowledgements
This work was supported by US National Institutes of Health grants 5R01CA155320-04 and P50 GM098792. We thank L. Wrobleska and P. Guye (Massachusetts Institute of Technology) for providing the initial intronic miRNA–based plasmid and the primary Cas9 construct, and for helpful discussions, and M. Graziano (Massachusetts Institute of Technology) for providing us the HEK293 cell lines that constitutively express rtTA3. J.H. was partially supported by the Intelligent Synthetic Biology Center of Global Frontier Project (2013M3A6A8073557) funded by the Ministry of Science, Information and Communication Technology and Future Planning of Korea.
Author information
Authors and Affiliations
Contributions
R.W. and S.K. conceived the idea. S.K., R.W., M.R.E. and J.B. designed experiments. S.K. performed the majority of experiments. M.R.E. and R.N.H. helped with flow cytometry and transfections. Y.L. and Z.X. built initial versions of CRP-a and CRP-b promoters. J.H. helped with DNA constructions. J.B. developed and applied computational analysis techniques. J.B. and S.K. performed the flow cytometry and statistical analysis. S.K. wrote the manuscript. R.W., J.B. and M.R.E. edited the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–15, Supplementary Discussion and Supplementary Note (PDF 31793 kb)
Rights and permissions
About this article
Cite this article
Kiani, S., Beal, J., Ebrahimkhani, M. et al. CRISPR transcriptional repression devices and layered circuits in mammalian cells. Nat Methods 11, 723–726 (2014). https://doi.org/10.1038/nmeth.2969
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nmeth.2969
This article is cited by
-
CRISPR/Cas9 system: a reliable and facile genome editing tool in modern biology
Molecular Biology Reports (2022)
-
dCas9 regulator to neutralize competition in CRISPRi circuits
Nature Communications (2021)
-
Controlling and enhancing CRISPR systems
Nature Chemical Biology (2021)
-
Design of time-delayed safety switches for CRISPR gene therapy
Scientific Reports (2021)
-
Multistable and dynamic CRISPRi-based synthetic circuits
Nature Communications (2020)