Abstract
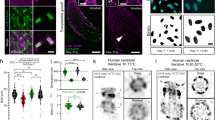
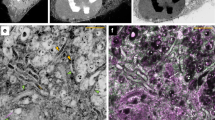
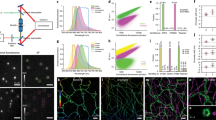
We combine super-resolution localization fluorescence microscopy with transmission electron microscopy of metal replicas to locate proteins on the landscape of the cellular plasma membrane at the nanoscale. We validate robust correlation on the scale of 20 nm by imaging endogenous clathrin (in two and three dimensions) and apply the method to find the previously unknown three-dimensional position of the endocytic protein epsin on clathrin-coated structures at the plasma membrane.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Betzig, E. et al. Science 313, 1642–1645 (2006).
Heilemann, M. et al. Angew. Chem. Int. Ed. Engl. 47, 6172–6176 (2008).
Brown, T.A. et al. Mol. Cell. Biol. 31, 4994–5010 (2011).
Bates, M., Huang, B., Dempsey, G.T. & Zhuang, X. Science 317, 1749–1753 (2007).
Kopek, B.G., Shtengel, G., Grimm, J.B., Clayton, D.A. & Hess, H.F. PLoS ONE 8, e77209 (2013).
Shtengel, G. et al. Proc. Natl. Acad. Sci. USA 106, 3125–3130 (2009).
Collins, A., Warrington, A., Taylor, K.A. & Svitkina, T. Curr. Biol. 21, 1167–1175 (2011).
Engqvist-Goldstein, Å.E.Y. et al. J. Cell Biol. 154, 1209–1223 (2001).
Suleiman, H. et al. eLife 2, e01149 (2013).
Watanabe, S. et al. Nat. Methods 8, 80–84 (2011).
Heuser, J. J. Cell Biol. 84, 560–583 (1980).
Ford, M.G. et al. Nature 419, 361–366 (2002).
Boucrot, E. et al. Cell 149, 124–136 (2012).
Hawryluk, M.J. et al. Traffic 7, 262–281 (2006).
Horvath, C.A., Vanden Broeck, D., Boulet, G.A., Bogers, J. & De Wolf, M.J. Int. J. Biochem. Cell Biol. 39, 1765–1770 (2007).
Saffarian, S., Cocucci, E. & Kirchhausen, T. PLoS Biol. 7, e1000191 (2009).
Mastronarde, D.N. J. Struct. Biol. 152, 36–51 (2005).
Kremer, J.R., Mastronarde, D.N. & McIntosh, J.R. J. Struct. Biol. 116, 71–76 (1996).
Zhang, M. et al. Nature Methods 9, 727–729 (2012).
Gaidarov, I., Santini, F., Warren, R.A. & Keen, J.H. Nat. Cell Biol. 1, 1–7 (1999).
Kanchanawong, P. et al. Nature 468, 580–584 (2010).
Sochacki, K.A. et al. Nature Commun. 3, 1154 (2012).
Schneider, C.A., Rasband, W.S. & Eliceiri, K.W. Nat. Methods 9, 671–675 (2012).
Acknowledgements
We thank M. Daniels and the US National Heart, Lung, and Blood Institute (NHLBI) electron microscopy core for help with EM; H. Shroff, J. Shaw and K. Neuman for critical reading of the manuscript; W. Li and Y. Wang (Janelia Farm) for EM grid preparation; L. Greene (NHLBI) for antibodies; S. Yu (NHLBI) for plasmid preparation; A. Nagy for acquiring atomic force microscopy (AFM) images; and E. Tyler and A. Hoofring of NIH Medical Arts for design work on Figure 1. J.W.T. is supported by the Intramural Research Program of the NHLBI, NIH. G.S. and H.F.H. are supported by the Howard Hughes Medical Institute.
Author information
Authors and Affiliations
Contributions
K.A.S., G.S. and J.W.T. designed the experiments. K.A.S. and G.S. performed the experiments. G.S. and K.A.S. processed data. K.A.S. analyzed the results. K.A.S. and J.W.T. wrote the manuscript. S.B.v.E. designed plasmids. J.W.T. and H.F.H. oversaw the project. All authors discussed the results and commented on the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–8 (PDF 1489 kb)
Correlated clathrin-AF647 tomograms viewed in XY
Tomograms of six clathrin structures correlated with clathrin-AF647 (clathrin-AF647 is magenta, myristoylated psCFP2 is cyan). The tomograms are viewed in the XY plane and scan the Z dimension over time. Each voxel is 2.3 nm in all dimensions but a frame is shown every 6.9 nm in the Z-dimension. Each tomogram is 459 nm wide and tall. (MOV 14766 kb)
Correlated clathrin-AF647 tomograms viewed in YZ
The tomograms from Supplementary Video 1 are viewed in the YZ plane and scan the X dimension over time (clathrin-AF647 is magenta, myristoylated psCFP2 is cyan). Each voxel is 2.3 nm in all dimensions but a frame is shown every 6.9 nm in the X-dimension. Each tomogram is 459 nm wide in the Y dimension. (MOV 24262 kb)
Membrane iPALM/EM alignment validation
A tomogram of clathrin data is viewed in the YZ plane and scans the X dimension over time (clathrin-AF647 is magenta, myristoylated psCFP2 is cyan). An example of membrane misalignment between fluorescence and EM is shown in frames 19-106 on the right hand side. Regions with membrane misalignment were not used for analysis. Each voxel is 2.3 nm in all dimensions but a frame is shown every 6.9 nm in the X-dimension. The tomogram is 215.7 nm high in the Z dimension. (MOV 25780 kb)
Correlated epsin-AF647 tomograms viewed in XY
Tomograms of six clathrin structures correlated with epsin-AF647 (epsin-AF647 is magenta, myristoylated psCFP2 is cyan). The tomograms are viewed in the XY plane and scan the Z dimension over time. Each voxel is 2.3 nm in all dimensions but a frame is shown every 6.9 nm in the Z-dimension. Each tomogram is 459 nm wide and tall. (MOV 13360 kb)
Correlated epsin-AF647 tomograms viewed in YZ
The tomograms from Video 4 are viewed in the YZ plane and scan the X dimension over time (epsin-AF647 is magenta, myristoylated psCFP2 is cyan). Each voxel is 2.3 nm in all dimensions but a frame is shown every 6.9 nm in the X-dimension. Each tomogram is 459 nm wide in the Y dimension. (MOV 20493 kb)
Isosurface models of epsin-AF647 tomograms
TEM tomograms (grey) and their corresponding epsin-AF647 fluorescence (magenta) are represented as isosurface models (Fig. 3f–h). Two models are shown, including three CCSs. Each model is 459 nm in the X and Y dimension and is rotated around the X-axis from 0-90°. (MOV 17873 kb)
Rights and permissions
About this article
Cite this article
Sochacki, K., Shtengel, G., van Engelenburg, S. et al. Correlative super-resolution fluorescence and metal-replica transmission electron microscopy. Nat Methods 11, 305–308 (2014). https://doi.org/10.1038/nmeth.2816
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nmeth.2816
This article is cited by
-
Adhesion energy controls lipid binding-mediated endocytosis
Nature Communications (2024)
-
A conformational switch in clathrin light chain regulates lattice structure and endocytosis at the plasma membrane of mammalian cells
Nature Communications (2023)
-
Actin polymerization promotes invagination of flat clathrin-coated lattices in mammalian cells by pushing at lattice edges
Nature Communications (2022)
-
The molecular organization of differentially curved caveolae indicates bendable structural units at the plasma membrane
Nature Communications (2022)
-
The nanoscale molecular morphology of docked exocytic dense-core vesicles in neuroendocrine cells
Nature Communications (2021)