Abstract
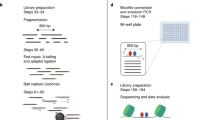
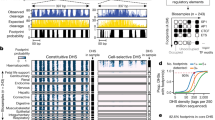
Sequencing of DNase I hypersensitive sites (DNase-seq) is a powerful technique for identifying cis-regulatory elements across the genome. We studied the key experimental parameters to optimize performance of DNase-seq. Sequencing short fragments of 50–100 base pairs (bp) that accumulate in long internucleosome linker regions was more efficient for identifying transcription factor binding sites compared to sequencing longer fragments. We also assessed the potential of DNase-seq to predict transcription factor occupancy via generation of nucleotide-resolution transcription factor footprints. In modeling the sequence-specific DNase I cutting bias, we found a strong effect that varied over more than two orders of magnitude. This indicates that the nucleotide-resolution cleavage patterns at many transcription factor binding sites are derived from intrinsic DNase I cleavage bias rather than from specific protein-DNA interactions. In contrast, quantitative comparison of DNase I hypersensitivity between states can predict transcription factor occupancy associated with particular biological perturbations.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Galas, D.J. & Schmitz, A. DNAse footprinting: a simple method for the detection of protein-DNA binding specificity. Nucleic Acids Res. 5, 3157–3170 (1978).
Song, L. et al. Open chromatin defined by DNase I and FAIRE identifies regulatory elements that shape cell-type identity. Genome Res. 21, 1757–1767 (2011).
Boyle, A.P. et al. High-resolution genome-wide in vivo footprinting of diverse transcription factors in human cells. Genome Res. 21, 456–464 (2011).
Degner, J.F. et al. DNase I sensitivity QTLs are a major determinant of human expression variation. Nature 482, 390–394 (2012).
Neph, S. et al. An expansive human regulatory lexicon encoded in transcription factor footprints. Nature 489, 83–90 (2012).
Thurman, R.E. et al. The accessible chromatin landscape of the human genome. Nature 489, 75–82 (2012).
Maurano, M.T. et al. Systematic localization of common disease-associated variation in regulatory DNA. Science 337, 1190–1195 (2012).
Voss, T.C. et al. Dynamic exchange at regulatory elements during chromatin remodeling underlies assisted loading mechanism. Cell 146, 544–554 (2011).
Ling, G., Sugathan, A., Mazor, T., Fraenkel, E. & Waxman, D.J. Unbiased, genome-wide in vivo mapping of transcriptional regulatory elements reveals sex differences in chromatin structure associated with sex-specific liver gene expression. Mol. Cell Biol. 30, 5531–5544 (2010).
John, S. et al. Chromatin accessibility pre-determines glucocorticoid receptor binding patterns. Nat. Genet. 43, 264–268 (2011).
He, H.H. et al. Differential DNase I hypersensitivity reveals factor-dependent chromatin dynamics. Genome Res. 22, 1015–1025 (2012).
Tewari, A.K. et al. Chromatin accessibility reveals insights into androgen receptor activation and transcriptional specificity. Genome Biol. 13, R88 (2012).
Zhang, Y. et al. Model-based analysis of ChIP-Seq (MACS). Genome Biol. 9, R137 (2008).
He, H.H. et al. Nucleosome dynamics define transcriptional enhancers. Nat. Genet. 42, 343–347 (2010).
Gaffney, D.J. et al. Controls of nucleosome positioning in the human genome. PLoS Genet. 8, e1003036 (2012).
Luger, K., Dechassa, M.L. & Tremethick, D.J. New insights into nucleosome and chromatin structure: an ordered state or a disordered affair? Nat. Rev. Mol. Cell Biol. 13, 436–447 (2012).
Lazarovici, A. et al. Probing DNA shape and methylation state on a genomic scale with DNase I. Proc. Natl. Acad. Sci. USA 110, 6376–6381 (2013).
Campbell, V.W. & Jackson, D.A. The effect of divalent cations on the mode of action of DNase I. The initial reaction products produced from covalently closed circular DNA. J. Biol. Chem. 255, 3726–3735 (1980).
Grontved, L. et al. Rapid genome-scale mapping of chromatin accessibility in tissue. Epigenetics Chromatin 5, 10 (2012).
Matys, V. et al. TRANSFAC and its module TRANSCompel: transcriptional gene regulation in eukaryotes. Nucleic Acids Res. 34, D108–D110 (2006).
ENCODE Project Consortium. et al. An integrated encyclopedia of DNA elements in the human genome. Nature 489, 57–74 (2012).
Zhang, Y., Shin, H., Song, J.S., Lei, Y. & Liu, X.S. Identifying positioned nucleosomes with epigenetic marks in human from ChIP-Seq. BMC Genomics 9, 537 (2008).
Acknowledgements
This work was supported by grants from the US National Institutes of Health (1R01 GM099409 to X.S.L.; 1U41 HG007000 to X.S.L. and C.A.M.; 2P50 CA090381-06 to C.A.M. and M.B.; 2R01 DK074967-06 to M.B. and X.S.L., 1K99CA172948-01 to H.H.H.); the Mazzone Award (to X.S.L.), the Department of Defense (W81XWH-10-1-0557 to H.H.H.) and the Prostate Cancer Foundation (to M.B.).
Author information
Authors and Affiliations
Contributions
H.H.H., C.A.M., H.L., X.S.L. and M.B. designed the experiments and wrote the manuscript. M.-W.C. and H.H.H. performed the experiments with the help from Y.L., P.K.R. and T.F. C.A.M., S.S.H. and H.H.H. conducted the data analysis with the help from C.Z. and H.X.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–21, Supplementary Tables 1–3 and Supplementary Protocol (PDF 5336 kb)
Rights and permissions
About this article
Cite this article
He, H., Meyer, C., Hu, S. et al. Refined DNase-seq protocol and data analysis reveals intrinsic bias in transcription factor footprint identification. Nat Methods 11, 73–78 (2014). https://doi.org/10.1038/nmeth.2762
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nmeth.2762
This article is cited by
-
TGFβ promotes widespread enhancer chromatin opening and operates on genomic regulatory domains
Nature Communications (2020)
-
Integrated epigenomic analysis stratifies chromatin remodellers into distinct functional groups
Epigenetics & Chromatin (2019)
-
XL-DNase-seq: improved footprinting of dynamic transcription factors
Epigenetics & Chromatin (2019)
-
Epigenomic signatures underpin the axonal regenerative ability of dorsal root ganglia sensory neurons
Nature Neuroscience (2019)
-
Maternal pluripotency factors initiate extensive chromatin remodelling to predefine first response to inductive signals
Nature Communications (2019)