Abstract
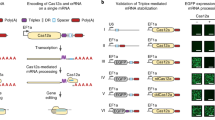
Study of the nematode Caenorhabditis elegans has provided important insights in a wide range of fields in biology. The ability to precisely modify genomes is critical to fully realize the utility of model organisms. Here we report a method to edit the C. elegans genome using the clustered, regularly interspersed, short palindromic repeats (CRISPR) RNA-guided Cas9 nuclease and homologous recombination. We demonstrate that Cas9 is able to induce DNA double-strand breaks with specificity for targeted sites and that these breaks can be repaired efficiently by homologous recombination. By supplying engineered homologous repair templates, we generated gfp knock-ins and targeted mutations. Together our results outline a flexible methodology to produce essentially any desired modification in the C. elegans genome quickly and at low cost. This technology is an important addition to the array of genetic techniques already available in this experimentally tractable model organism.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Change history
16 September 2013
In the version of this article initially published online, the phrase "insertion of gfp into the nmy-2 gene and a 23-bp loxP site" should have read "insertion of gfp into the nmy-2 gene and a 34-bp loxP site." In the Online Methods, under the heading "Single-copy transgene insertion with MosSCI," "pGH8 (Prab-8::mCherry neuronal co-injection marker)" should have read "pGH8 (Prab-3::mCherry neuronal co-injection marker)." The errors have been corrected for the print, PDF and HTML versions of this article.
References
Jinek, M. et al. A programmable dual-RNA-guided DNA endonuclease in adaptive bacterial immunity. Science 337, 816–821 (2012).
Hwang, W.Y. et al. Efficient genome editing in zebrafish using a CRISPR-Cas system. Nat. Biotechnol. 31, 227–229 (2013).
DiCarlo, J.E. et al. Genome engineering in Saccharomyces cerevisiae using CRISPR-Cas systems. Nucleic Acids Res. 41, 4336–4343 (2013).
Gratz, S.J. et al. Genome engineering of Drosophila with the CRISPR RNA-guided Cas9 nuclease. Genetics 194, 1029–1035 (2013).
Cong, L. et al. Multiplex genome engineering using CRISPR/Cas systems. Science 339, 819–823 (2013).
Wang, H. et al. One-step generation of mice carrying mutations in multiple genes by CRISPR/Cas-mediated genome engineering. Cell 153, 910–918 (2013).
Jiang, W., Bikard, D., Cox, D., Zhang, F. & Marraffini, L.A. RNA-guided editing of bacterial genomes using CRISPR-Cas systems. Nat. Biotechnol. 31, 233–239 (2013).
Friedland, A.E. et al. Heritable genome editing in C. elegans via a CRISPR-Cas9 system. Nat. Methods 10, 741–743 (2013).
Bassett, A.R., Tibbit, C., Ponting, C.P. & Liu, J.-L. Highly efficient targeted mutagenesis of Drosophila with the CRISPR/Cas9 system. Cell Rep. 4, 220–228 (2013).
Chang, N. et al. Genome editing with RNA-guided Cas9 nuclease in zebrafish embryos. Cell Res. 23, 465–472 (2013).
Mali, P. et al. RNA-guided human genome engineering via Cas9. Science 339, 823–826 (2013).
Cho, S.W., Kim, S., Kim, J.M. & Kim, J.-S. Targeted genome engineering in human cells with the Cas9 RNA-guided endonuclease. Nat. Biotechnol. 31, 230–232 (2013).
Shen, B. et al. Generation of gene-modified mice via Cas9/RNA-mediated gene targeting. Cell Res. 23, 720–723 (2013).
Wood, A.J. et al. Targeted genome editing across species using ZFNs and TALENs. Science 333, 307 (2011).
Lo, T.-W. et al. Heritable genome editing using TALENs and CRISPR/Cas9 to engineer precise insertions and deletions in evolutionarily diverse nematode species. Genetics doi:10.1534/genetics.113.155382 (9 August 2013).
Gerstein, M.B. et al. Integrative analysis of the Caenorhabditis elegans genome by the modENCODE project. Science 330, 1775–1787 (2010).
Frøkjær-Jensen, C., Davis, M.W., Ailion, M. & Jorgensen, E.M. Improved Mos1-mediated transgenesis in C. elegans. Nat. Methods 9, 117–118 (2012).
Robert, V. & Bessereau, J.-L. Targeted engineering of the Caenorhabditis elegans genome following Mos1-triggered chromosomal breaks. EMBO J. 26, 170–183 (2007).
Robert, V.J., Davis, M.W., Jorgensen, E.M. & Bessereau, J.-L. Gene conversion and end-joining-repair double-strand breaks in the Caenorhabditis elegans germline. Genetics 180, 673–679 (2008).
Frøkjær-Jensen, C. et al. Single-copy insertion of transgenes in Caenorhabditis elegans. Nat. Genet. 40, 1375–1383 (2008).
Frøkjær-Jensen, C. et al. Targeted gene deletions in C. elegans using transposon excision. Nat. Methods 7, 451–453 (2010).
Vallin, E. et al. A genome-wide collection of Mos1 transposon insertion mutants for the C. elegans research community. PLoS ONE 7, e30482 (2012).
Mello, C.C., Kramer, J.M., Stinchcomb, D. & Ambros, V. Efficient gene transfer in C.elegans: extrachromosomal maintenance and integration of transforming sequences. EMBO J. 10, 3959–3970 (1991).
Kelly, W.G., Xu, S., Montgomery, M.K. & Fire, A. Distinct requirements for somatic and germline expression of a generally expressed Caenorhabditis elegans gene. Genetics 146, 227–238 (1997).
Praitis, V., Casey, E., Collar, D. & Austin, J. Creation of low-copy integrated transgenic lines in Caenorhabditis elegans. Genetics 157, 1217–1226 (2001).
Sarov, M. et al. A genome-scale resource for in vivo tag-based protein function exploration in C. elegans. Cell 150, 855–866 (2012).
Nance, J., Munro, E.M. & Priess, J.R. C. elegans PAR-3 and PAR-6 are required for apicobasal asymmetries associated with cell adhesion and gastrulation. Development 130, 5339–5350 (2003).
Guo, S. & Kemphues, K.J. A non-muscle myosin required for embryonic polarity in Caenorhabditis elegans. Nature 382, 455–458 (1996).
Berezikov, E., Bargmann, C.I. & Plasterk, R.H.A. Homologous gene targeting in Caenorhabditis elegans by biolistic transformation. Nucleic Acids Res. 32, e40 (2004).
Ferguson, E.L. & Horvitz, H.R. Identification and characterization of 22 genes that affect the vulval cell lineages of the nematode Caenorhabditis elegans. Genetics 110, 17–72 (1985).
Tan, P.B., Lackner, M.R. & Kim, S.K. MAP kinase signaling specificity mediated by the LIN-1 Ets/LIN-31 WH transcription factor complex during C. elegans vulval induction. Cell 93, 569–580 (1998).
Fu, Y. et al. High-frequency off-target mutagenesis induced by CRISPR-Cas nucleases in human cells. Nat. Biotechnol. doi:10.1038/nbt.2623 (23 June 2013).
Granger, L., Martin, E. & Ségalat, L. Mos as a tool for genome-wide insertional mutagenesis in Caenorhabditis elegans: results of a pilot study. Nucleic Acids Res. 32, e117 (2004).
Williams, D.C., Boulin, T., Ruaud, A.-F., Jorgensen, E.M. & Bessereau, J.-L. Characterization of Mos1-mediated mutagenesis in Caenorhabditis elegans: a method for the rapid identification of mutated genes. Genetics 169, 1779–1785 (2005).
Shirayama, M. et al. piRNAs initiate an epigenetic memory of nonself RNA in the C. elegans germline. Cell 150, 65–77 (2012).
Lam, A.J. et al. Improving FRET dynamic range with bright green and red fluorescent proteins. Nat. Methods 9, 1005–1012 (2012).
Nguyen, A.W. & Daugherty, P.S. Evolutionary optimization of fluorescent proteins for intracellular FRET. Nat. Biotechnol. 23, 355–360 (2005).
Shcherbo, D. et al. Far-red fluorescent tags for protein imaging in living tissues. Biochem. J. 418, 567–574 (2009).
Stiernagle, T. in WormBook (ed. The C. elegans Research Community) doi:10.1895/wormbook.1.101.1 (2006).
Redemann, S. et al. Codon adaptation-based control of protein expression in C. elegans. Nat. Methods 8, 250–252 (2011).
Macosko, E.Z. et al. A hub-and-spoke circuit drives pheromone attraction and social behaviour in C. elegans. Nature 458, 1171–1175 (2009).
Merritt, C., Rasoloson, D., Ko, D. & Seydoux, G. 3′ UTRs are the primary regulators of gene expression in the C. elegans germline. Curr. Biol. 18, 1476–1482 (2008).
Maduro, M. & Pilgrim, D. Identification and cloning of unc-119, a gene expressed in the Caenorhabditis elegans nervous system. Genetics 141, 977–988 (1995).
Zhang, Y., Chen, D., Smith, M.A., Zhang, B. & Pan, X. Selection of reliable reference genes in Caenorhabditis elegans for analysis of nanotoxicity. PLoS ONE 7, e31849 (2012).
Hoogewijs, D., Houthoofd, K., Matthijssens, F., Vandesompele, J. & Vanfleteren, J.R. Selection and validation of a set of reliable reference genes for quantitative sod gene expression analysis in C. elegans. BMC Mol. Biol. 9, 9 (2008).
Altschul, S.F. et al. Gapped BLAST and PSI-BLAST: a new generation of protein database search programs. Nucleic Acids Res. 25, 3389–3402 (1997).
Acknowledgements
We thank C. Frøkjær-Jensen (University of Utah) for sharing strains, plasmids and protocols; K. Kemphues (Cornell University) for antibodies; and K. Bloom, A. Maddox, G. Monsalve, N. Pujol, K. Slep, S. Taubert, K. Yamamoto and members of the Goldstein lab for helpful suggestions and comments on the manuscript. Some strains were provided by the Caenorhabditis Genetics Center, which is funded by the US National Institutes of Health (NIH) Office of Research Infrastructure Programs (P40 OD010440). This work was supported by NIH T32 CA009156 and a Howard Hughes postdoctoral fellowship from the Helen Hay Whitney Foundation (D.J.D.); a postdoctoral fellowship from the Canadian Institutes of Health Research (award #234765) (J.D.W.); NIH R01 GM085309 (D.J.R.); NIH CA20535 and US National Science Foundation (NSF) MCB 1157767 (K. Yamamoto); and NIH R01 GM083071 and NSF IOS 0917726 (B.G.).
Author information
Authors and Affiliations
Contributions
D.J.D. and J.D.W. jointly conceived of the project, and all authors discussed and contributed to the experimental design. D.J.D. performed the experiments. D.J.D. and D.J.R. analyzed the data. D.J.D. prepared the manuscript, and all authors discussed and contributed to the final version.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1 and 2, Supplementary Tables 1–6 and Supplementary Protocol (PDF 4030 kb)
NMY-2–GFP dynamics in zuIs45 and knock-in embryos
1-cell embryos homozygous for either zuIs45 or the nmy-2::gfp knock-in were placed side by side on the same coverslip and filmed simultaneously. Still images from this movie are shown in Fig. 3e. (AVI 53807 kb)
Rights and permissions
About this article
Cite this article
Dickinson, D., Ward, J., Reiner, D. et al. Engineering the Caenorhabditis elegans genome using Cas9-triggered homologous recombination. Nat Methods 10, 1028–1034 (2013). https://doi.org/10.1038/nmeth.2641
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nmeth.2641
This article is cited by
-
Design Principles of a Novel Construct for HBB Gene-Editing and Investigation of Its Gene-Targeting Efficiency in HEK293 Cells
Molecular Biotechnology (2024)
-
Molecular mechanisms of neurite regeneration and repair: insights from C. elegans and Drosophila
Cell Regeneration (2023)
-
Ribo-On and Ribo-Off tools using a self-cleaving ribozyme allow manipulation of endogenous gene expression in C. elegans
Communications Biology (2023)
-
UNC-43/CaMKII-triggered anterograde signals recruit GABAARs to mediate inhibitory synaptic transmission and plasticity at C. elegans NMJs
Nature Communications (2023)
-
ATP sulfurylase atypical leucine zipper interacts with Cys3 and calcineurin A in the regulation of sulfur amino acid biosynthesis in Cryptococcus neoformans
Scientific Reports (2023)