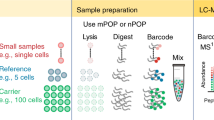
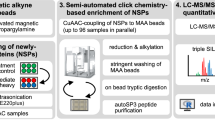
Abstract
We report a technique to selectively and continuously label the proteomes of individual cell types in coculture, named cell type–specific labeling using amino acid precursors (CTAP). Through transgenic expression of exogenous amino acid biosynthesis enzymes, vertebrate cells overcome their dependence on supplemented essential amino acids and can be selectively labeled through metabolic incorporation of amino acids produced from heavy isotope–labeled precursors. When testing CTAP in several human and mouse cell lines, we could differentially label the proteomes of distinct cell populations in coculture and determine the relative expression of proteins by quantitative mass spectrometry. In addition, using CTAP we identified the cell of origin of extracellular proteins secreted from cells in coculture. We believe that this method, which allows linking of proteins to their cell source, will be useful in studies of cell-cell communication and potentially for discovery of biomarkers.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Joyce, J.A. & Pollard, J.W. Microenvironmental regulation of metastasis. Nat. Rev. Cancer 9, 239–252 (2009).
Mcmillin, D.W. et al. Tumor cell-specific bioluminescence platform to identify stroma-induced changes to anticancer drug activity. Nat. Med. 16, 483–489 (2010).
Olumi, A.F. et al. Carcinoma-associated fibroblasts direct tumor progression of initiated human prostatic epithelium. Cancer Res. 59, 5002–5011 (1999).
Gygi, S.P. et al. Quantitative analysis of complex protein mixtures using isotope-coded affinity tags. Nat. Biotechnol. 17, 994–999 (1999).
Ong, S.-E. et al. Stable isotope labeling by amino acids in cell culture, SILAC, as a simple and accurate approach to expression proteomics. Mol. Cell Proteomics 1, 376–386 (2002).
Ross, P.L. et al. Multiplexed protein quantitation in Saccharomyces cerevisiae using amine-reactive isobaric tagging reagents. Mol. Cell Proteomics 3, 1154–1169 (2004).
Jørgensen, C. et al. Cell-specific information processing in segregating populations of Eph receptor ephrin-expressing cells. Science 326, 1502–1509 (2009).
Rechavi, O. et al. Trans-SILAC: sorting out the non-cell-autonomous proteome. Nat. Methods 7, 923–927 (2010).
Naba, A. et al. The matrisome: in silico definition and in vivo characterization by proteomics of normal and tumor extracellular matrices. Mol. Cell Proteomics 11, M111.014647 (2012).
van den Bemd, G.-J.C.M. et al. Mass spectrometric identification of human prostate cancer-derived proteins in serum of xenograft-bearing mice. Mol. Cell Proteomics 5, 1830–1839 (2006).
Ngo, J.T. et al. Cell-selective metabolic labeling of proteins. Nat. Chem. Biol. 5, 715–717 (2009).
Truong, F., Yoo, T.H., Lampo, T.J. & Tirrell, D.A. Two-strain, cell-selective protein labeling in mixed bacterial cultures. J. Am. Chem. Soc. 134, 8551–8556 (2012).
Ngo, J.T., Schuman, E.M. & Tirrell, D.A. Mutant methionyl-tRNA synthetase from bacteria enables site-selective N-terminal labeling of proteins expressed in mammalian cells. Proc. Natl. Acad. Sci. USA 110, 4992–4997 (2013).
Liu, C.C. & Schultz, P.G. Adding new chemistries to the genetic code. Annu. Rev. Biochem. 79, 413–444 (2010).
Xu, H., Andi, B., Qian, J., West, A.H. & Cook, P.F. The alpha-aminoadipate pathway for lysine biosynthesis in fungi. Cell Biochem. Biophys. 46, 43–64 (2006).
Ong, S.-E. & Mann, M. A practical recipe for stable isotope labeling by amino acids in cell culture (SILAC). Nat. Protoc. 1, 2650–2660 (2006).
Saqib, K.M., Hay, S.M. & Rees, W.D. The expression of Escherichia coli diaminopimelate decarboxylase in mouse 3T3 cells. Biochim. Biophys. Acta 1219, 398–404 (1994).
Jouanneau, J., Stragier, P., Bouvier, J., Patte, J.C. & Yaniv, M. Expression in mammalian cells of the diaminopimelic acid decarboxylase of Escherichia coli permits cell growth in lysine-free medium. Eur. J. Biochem. 146, 173–178 (1985).
Cox, J. & Mann, M. MaxQuant enables high peptide identification rates, individualized p.p.b.-range mass accuracies and proteome-wide protein quantification. Nat. Biotechnol. 26, 1367–1372 (2008).
Sury, M.D., Chen, J.-X. & Selbach, M. The SILAC fly allows for accurate protein quantification in vivo. Mol. Cell Proteomics 9, 2173–2183 (2010).
Spellman, D.S., Deinhardt, K., Darie, C.C., Chao, M.V. & Neubert, T.A. Stable isotopic labeling by amino acids in cultured primary neurons: application to brain-derived neurotrophic factor-dependent phosphotyrosine-associated signaling. Mol. Cell Proteomics 7, 1067–1076 (2008).
Geiger, T., Wehner, A., Schaab, C., Cox, J. & Mann, M. Comparative proteomic analysis of eleven common cell lines reveals ubiquitous but varying expression of most proteins. Mol Cell Proteomics 11, M111.014050 (2012).
Kuan, Y.-C. et al. Biochemical characterization of a novel lysine racemase from Proteus mirabilis BCRC10725. Process Biochem. 46, 1914–1920 (2011).
Nanduri, V., Goldberg, S., Johnston, R. & Patel, R. Cloning and expression of a novel enantioselective N-carbobenzyloxy-cleaving enzyme. Enzyme Microb. Technol. 34, 304–312 (2004).
Mavrakis, K.J. et al. Tumorigenic activity and therapeutic inhibition of Rheb GTPase. Genes Dev. 22, 2178–2188 (2008).
Swift, S., Lorens, J., Achacoso, P. & Nolan, G.P. Rapid production of retroviruses for efficient gene delivery to mammalian cells using 293T cell-based systems. Curr. Protoc. Immunol. 10, 10.17C (2001).
Szymczak, A.L. et al. Correction of multi-gene deficiency in vivo using a single 'self-cleaving' 2A peptide-based retroviral vector. Nat. Biotechnol. 22, 589–594 (2004).
Smyth, G. Limma: linear models for microarray data. in Bioinformatics and Computational Biology Solutions Using R and Bioconductor (eds. Gentleman, R., Carey, V., Dudoit, S., Irizarry, R. & Huber, W.) 397–420 (Springer, New York, 2005).
Benjamini, Y. & Hochberg, Y. Controlling the false discovery rate: a practical and powerful approach to multiple testing. J. R. Stat. Soc. B 57, 289–300 (1995).
Subramanian, A. et al. Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc. Natl. Acad. Sci. USA 102, 15545–15550 (2005).
Shevchenko, A., Tomas, H., Havlis, J., Olsen, J.V. & Mann, M. In-gel digestion for mass spectrometric characterization of proteins and proteomes. Nat. Protoc. 1, 2856–2860 (2006).
Macek, B. et al. Phosphorylation of the human full-length protein kinase Ciota. J. Proteome Res. 7, 2928–2935 (2008).
Ishihama, Y., Rappsilber, J. & Mann, M. Modular stop and go extraction tips with stacked disks for parallel and multidimensional Peptide fractionation in proteomics. J. Proteome Res. 5, 988–994 (2006).
Olsen, J.V. et al. Global, in vivo, and site-specific phosphorylation dynamics in signaling networks. Cell 127, 635–648 (2006).
Cox, J. et al. Andromeda: a peptide search engine integrated into the MaxQuant environment. J. Proteome Res. 10, 1794–1805 (2011).
Acknowledgements
We acknowledge E. Larsson, Y. Gruber, D.S. Marks, A. Arvey, J. Joyce and A. Koff for helpful discussions, H. Erdjument-Bromage for pilot MS/MS investigation, A.N. Miller, J. Cross and X. Jing for technical help, and E. Larsson, J. Gauthier, J. Joyce and A.M. Miller for helpful comments on the manuscript. This work was funded in part by US National Cancer Institute grant U54 CA148967.
Author information
Authors and Affiliations
Contributions
N.P.G. and M.L.M. designed and performed experiments and analyzed data. W.E.W. generated reagents. B.S., K.J.M. and V.A.P. contributed to experiments. N.P.G. and M.L.M. wrote the manuscript. B.S., W.E.W., B.M., K.J.M., V.A.P., D.Y.G. and C.S. contributed to discussions and editing of the manuscript. N.P.G. conceived the hypothesis. N.P.G., C.S. and M.L.M. developed the concept and managed the project.
Corresponding authors
Ethics declarations
Competing interests
A provisional patent application (US 61/697,584) relating to the use of exogenous enzymes for proteomic labeling in multicellular culture has been filed by Memorial Sloan-Kettering Cancer Center with N.P.G., C.S. and M.L.M. listed as inventors.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–19, Supplementary Tables 1–8, Supplementary Notes 1–2 and Supplementary Discussion (PDF 2669 kb)
Supplementary Data 1
Supplementary Data 1 (XLSX 2650 kb)
Rights and permissions
About this article
Cite this article
Gauthier, N., Soufi, B., Walkowicz, W. et al. Cell-selective labeling using amino acid precursors for proteomic studies of multicellular environments. Nat Methods 10, 768–773 (2013). https://doi.org/10.1038/nmeth.2529
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nmeth.2529
This article is cited by
-
Proteomic characterisation of prostate cancer intercellular communication reveals cell type-selective signalling and TMSB4X-dependent fibroblast reprogramming
Cellular Oncology (2022)
-
Cell-specific bioorthogonal tagging of glycoproteins
Nature Communications (2022)
-
Nitrogen resorption and fractionation during leaf senescence in typical tree species in Japan
Journal of Forestry Research (2020)
-
Cell-type-specific metabolic labeling, detection and identification of nascent proteomes in vivo
Nature Protocols (2019)
-
Quantitative phosphoproteomic analysis reveals reciprocal activation of receptor tyrosine kinases between cancer epithelial cells and stromal fibroblasts
Clinical Proteomics (2018)