Abstract
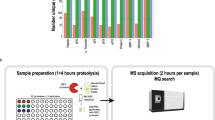
We developed a simple and rapid multiplex substrate-profiling method to reveal the substrate specificity of any endo- or exopeptidase using liquid chromatography–tandem mass spectrometry sequencing. We generated a physicochemically diverse library of peptides by incorporating all combinations of neighbor and near-neighbor amino acid pairs into decapeptide sequences that are flanked by unique dipeptides at each terminus. Addition of a panel of evolutionarily diverse peptidases to a mixture of these tetradecapeptides generated information on prime and nonprime sites as well as on substrate specificity that matched or expanded upon known substrate motifs. This method biochemically confirmed the activity of the klassevirus 3C protein responsible for polypeptide processing and allowed granzyme B substrates to be ranked by enzymatic turnover efficiency using label-free quantitation of precursor-ion abundance. Additionally, the proteolytic secretions from schistosome parasitic flatworm larvae and a pancreatic cancer cell line were deconvoluted in a subtractive strategy using class-specific peptidase inhibitors.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Puente, X.S., Gutiérrez-Fernández, A., Velasco, G. & López-Otín, C. A genomic view of the complexity of mammalian proteolytic systems. Biochem. Soc. Trans. 33, 331–334 (2005).
Van Damme, P., Vandekerckhove, J. & Gevaert, K. Disentanglement of protease substrate repertoires. Biol. Chem. 389, 371–381 (2008).
Matthews, D.J. & Wells, J.A. Substrate phage: selection of protease substrates by monovalent phage display. Science 260, 1113–1117 (1993).
Boulware, K.T. & Daugherty, P.S. Protease specificity determination by using cellular libraries of peptide substrates (CLiPS). Proc. Natl. Acad. Sci. USA 103, 7583–7588 (2006).
Turk, B.E., Huang, L.L., Piro, E.T. & Cantley, L.C. Determination of protease cleavage site motifs using mixture-based oriented peptide libraries. Nat. Biotechnol. 19, 661–667 (2001).
Harris, J.L. et al. Rapid and general profiling of protease specificity by using combinatorial fluorogenic substrate libraries. Proc. Natl. Acad. Sci. USA 97, 7754–7759 (2000).
Choe, Y. et al. Substrate profiling of cysteine proteases using a combinatorial peptide library identifies functionally unique specificities. J. Biol. Chem. 281, 12824–12832 (2006).
auf dem Keller, U. & Schilling, O. Proteomic techniques and activity-based probes for the system-wide study of proteolysis. Biochimie 92, 1705–1714 (2010).
Mahrus, S. et al. Global sequencing of proteolytic cleavage sites in apoptosis by specific labeling of protein N termini. Cell 134, 866–876 (2008).
Staes, A. et al. Improved recovery of proteome-informative, protein N-terminal peptides by combined fractional diagonal chromatography (COFRADIC). Proteomics 8, 1362–1370 (2008).
Impens, F. et al. A quantitative proteomics design for systematic identification of protease cleavage events. Mol. Cell. Proteomics 9, 2327–2333 (2010).
Colaert, N., Helsens, K., Martens, L., Vandekerckhove, J. & Gevaert, K. Improved visualization of protein consensus sequences by iceLogo. Nat. Methods 6, 786–787 (2009).
Chajkowski, S.M. et al. Highly selective hydrolysis of kinins by recombinant prolylcarboxypeptidase. Biochem. Biophys. Res. Commun. 405, 338–343 (2011).
Sun, J. et al. Importance of the P4′ residue in human granzyme B inhibitors and substrates revealed by scanning mutagenesis of the proteinase inhibitor 9 reactive center loop. J. Biol. Chem. 276, 15177–15184 (2001).
Cordingley, M.G., Callahan, P.L., Sardana, V.V., Garsky, V.M. & Colonno, R.J. Substrate requirements of human rhinovirus 3C protease for peptide cleavage in vitro. J. Biol. Chem. 265, 9062–9065 (1990).
Sojka, D. et al. Characterization of gut-associated cathepsin D hemoglobinase from tick Ixodes ricinus (IrCD1). J. Biol. Chem. 287, 21152–21163 (2012).
Ingram, J.R. et al. Investigation of the proteolytic functions of an expanded cercarial elastase gene family in Schistosoma mansoni. PLoS Negl. Trop. Dis. 6, e1589 (2012).
Knudsen, G.M., Medzihradszky, K.F., Lim, K.-C., Hansell, E. & McKerrow, J.H. Proteomic analysis of Schistosoma mansoni cercarial secretions. Mol. Cell. Proteomics 4, 1862–1875 (2005).
Curwen, R.S., Ashton, P.D., Sundaralingam, S. & Wilson, R.A. Identification of novel proteases and immunomodulators in the secretions of schistosome cercariae that facilitate host entry. Mol. Cell. Proteomics 5, 835–844 (2006).
Nolan-Stevaux, O. et al. GLI1 is regulated through Smoothened-independent mechanisms in neoplastic pancreatic ducts and mediates PDAC cell survival and transformation. Genes Dev. 23, 24–36 (2009).
Fricker, L. Carboxypeptidase E. in Handbook of Proteolytic Enzymes 2nd edn. (eds. Barrett, A.J., Rawlings, N.D. & Woessner, J.F.) 840–844 (Academic, 2004).
Caglic, D. et al. Murine and human cathepsin B exhibit similar properties: possible implications for drug discovery. Biol. Chem. 390, 175–179 (2009).
Mason, R.W., Johnson, D.A., Barrett, A.J. & Chapman, H.A. Elastinolytic activity of human cathepsin L. Biochem. J. 233, 925–927 (1986).
Zaidi, N., Herrmann, T., Voelter, W. & Kalbacher, H. Recombinant cathepsin E has no proteolytic activity at neutral pH. Biochem. Biophys. Res. Commun. 360, 51–55 (2007).
Impens, F. et al. MS-driven protease substrate degradomics. Proteomics 10, 1284–1296 (2010).
Rawlings, N.D., Barrett, A.J. & Bateman, A. MEROPS: the database of proteolytic enzymes, their substrates and inhibitors. Nucleic Acids Res. 40, D343–D350 (2012).
Ruggles, S.W., Fletterick, R.J. & Craik, C.S. Characterization of structural determinants of granzyme B reveals potent mediators of extended substrate specificity. J. Biol. Chem. 279, 30751–30759 (2004).
Prudova, A., auf dem Keller, U., Butler, G.S. & Overall, C.M. Multiplex N-terminome analysis of MMP-2 and MMP-9 substrate degradomes by iTRAQ-TAILS quantitative proteomics. Mol. Cell. Proteomics 9, 894–911 (2010).
Zhou, C. et al. Design and synthesis of prolylcarboxypeptidase (PrCP) inhibitors to validate PrCP as a potential target for obesity. J. Med. Chem. 53, 7251–7263 (2010).
Cohen, F.E. et al. Arresting tissue invasion of a parasite by protease inhibitors chosen with the aid of computer modeling. Biochemistry 30, 11221–11229 (1991).
Cruz-Monserrate, Z. et al. Detection of pancreatic cancer tumours and precursor lesions by cathepsin E activity in mouse models. Gut 61, 1315–1322 (2012).
Barkan, D.T. et al. Prediction of protease substrates using sequence and structure features. Bioinformatics 26, 1714–1722 (2010).
Harris, J.L., Peterson, E.P., Hudig, D., Thornberry, N.A. & Craik, C.S. Definition and redesign of the extended substrate specificity of granzyme B. J. Biol. Chem. 273, 27364–27373 (1998).
Salter, J.P. et al. Cercarial elastase is encoded by a functionally conserved gene family across multiple species of schistosomes. J. Biol. Chem. 277, 24618–24624 (2002).
Chalkley, R.J., Baker, P.R., Medzihradszky, K.F., Lynn, A.J. & Burlingame, A.L. In-depth analysis of tandem mass spectrometry data from disparate instrument types. Mol. Cell. Proteomics 7, 2386–2398 (2008).
Elias, J.E. & Gygi, S.P. Target-decoy search strategy for increased confidence in large-scale protein identifications by mass spectrometry. Nat. Methods 4, 207–214 (2007).
Liu, H., Sadygov, R.G. & Yates, J.R. III. A model for random sampling and estimation of relative protein abundance in shotgun proteomics. Anal. Chem. 76, 4193–4201 (2004).
Choi, H., Fermin, D. & Nesvizhskii, A.I. Significance analysis of spectral count data in label-free shotgun proteomics. Mol. Cell. Proteomics 7, 2373–2385 (2008).
O'Donoghue, A.J. et al. Inhibition of a secreted glutamic peptidase prevents growth of the fungus Talaromyces emersonii. J. Biol. Chem. 283, 29186–29195 (2008).
Acknowledgements
We thank F.A. Lewis at the US National Institutes of Health–National Institute of Allergy and Infectious Disease Schistosomiasis Resource Center for providing S. mansoni–infected Biomphalaria glabrata and K.C. Lim of the Sandler Center for Drug Discovery at UCSF for maintenance of the S. mansoni life cycle. We thank C. Tajon, C. Brown, G. Lee and S. Clarke from UCSF, M. Tuohy from National University of Ireland, Galway, and W. Geissler from Merck & Co. for providing purified peptidases and D. Hanahan from UCSF for the mouse PDAC cell line. This project was supported by grants from the US National Institutes of Health: P50 GM082250 (C.S.C.), RO1 CA128765 (C.S.C.), P41 RR001614 (A.L.B.), P41 GM103481 (A.L.B.) and K12 GM081266 (A.A.E.-R.) and the Sandler Center (C.S.C.).
Author information
Authors and Affiliations
Contributions
A.J.O., M.Z. and C.S.C. conceptualized the MSP-MS library and assay. C.S.C. directed and coordinated the project. M.O.A. and G.Q. wrote the pair-fitting script and J.B.S. wrote the MSP-MS extractor script. A.J.O., A.A.E.-R. and M.Z. synthesized and purified the peptides. A.J.O., A.A.E.-R. and G.M.K. performed the MSP-MS assays and analyzed the data. G.M.K., D.A.M. and A.L.B. developed the mass spectrometry protocol. J.I. and J.H.M. generated S. mansoni–infected snail samples. A.J.O. and D.R.H. generated conditioned PDAC medium. A.L.G. and J.L.D. provided viral peptidases. A.J.O., G.M.K., A.A.E.-R. and C.S.C. wrote the manuscript, and all authors participated in editing it.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–5 and Supplementary Tables 1–5 (PDF 785 kb)
Supplementary Data
P4–P4' cleavage sequences generated by multiplex substrate profiling by mass spectrometry for cathepsin E, prolylcarboxypeptidase, cruzain, MMP2, matriptase, DPP-IV, eqolisin, aspergillopepsin, HIV-1 and HIV-2 proteases, granzyme B, pancreatic ductal adenocarcinoma–conditioned medium and Shistosoma mansoni–infected snail sheds. (XLSX 130 kb)
Rights and permissions
About this article
Cite this article
O'Donoghue, A., Eroy-Reveles, A., Knudsen, G. et al. Global identification of peptidase specificity by multiplex substrate profiling. Nat Methods 9, 1095–1100 (2012). https://doi.org/10.1038/nmeth.2182
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nmeth.2182
This article is cited by
-
Mutations in α-synuclein, TDP-43 and tau prolong protein half-life through diminished degradation by lysosomal proteases
Molecular Neurodegeneration (2023)
-
Fluorescence Determination of Peptidase Activity Based on the Quenching of a Fluorophore-Labelled Peptide by Graphene Oxide
The Protein Journal (2021)
-
High throughput protease profiling comprehensively defines active site specificity for thrombin and ADAMTS13
Scientific Reports (2018)
-
Block-based characterization of protease specificity from substrate sequence profile
BMC Bioinformatics (2017)
-
The Rational Design of Therapeutic Peptides for Aminopeptidase N using a Substrate-Based Approach
Scientific Reports (2017)